Abstract
Phosphorylation of the spliceosome is essential for RNA splicing, yet how and to what extent kinase signaling affects splicing have not been defined on a genome-wide basis. Using a chemical genetic approach, we show in Schizosaccharomyces pombe that the SR protein kinase Dsk1 is required for efficient splicing of introns with suboptimal splice sites. Systematic substrate mapping in fission yeast and human cells revealed that SRPKs target evolutionarily conserved spliceosomal proteins, including the branchpoint-binding protein Bpb1 (SF1 in humans), by using an RXXSP consensus motif for substrate recognition. Phosphorylation of SF1 increases SF1 binding to introns with nonconsensus splice sites in vitro, and mutation of such sites to consensus relieves the requirement for Dsk1 and phosphorylated Bpb1 in vivo. Modulation of splicing efficiency through kinase signaling pathways may allow tuning of gene expression in response to environmental and developmental cues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Wahl, M.C., Will, C.L. & Lührmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009).
Mermoud, J.E., Cohen, P. & Lamond, A.I. Ser/Thr-specific protein phosphatases are required for both catalytic steps of pre-mRNA splicing. Nucleic Acids Res. 20, 5263–5269 (1992).
Mermoud, J.E., Cohen, P.T. & Lamond, A.I. Regulation of mammalian spliceosome assembly by a protein phosphorylation mechanism. EMBO J. 13, 5679–5688 (1994).
Tazi, J. et al. Thiophosphorylation of U1-70K protein inhibits pre-mRNA splicing. Nature 363, 283–286 (1993).
Gui, J.F., Lane, W.S. & Fu, X.D. A serine kinase regulates intracellular localization of splicing factors in the cell cycle. Nature 369, 678–682 (1994).
Ghosh, G. & Adams, J.A. Phosphorylation mechanism and structure of serine-arginine protein kinases. FEBS J. 278, 587–597 (2011).
Giannakouros, T., Nikolakaki, E., Mylonis, I. & Georgatsou, E. Serine-arginine protein kinases: a small protein kinase family with a large cellular presence. FEBS J. 278, 570–586 (2011).
Colwill, K. et al. The Clk/Sty protein kinase phosphorylates SR splicing factors and regulates their intranuclear distribution. EMBO J. 15, 265–275 (1996).
Duncan, P.I., Stojdl, D.F., Marius, R.M., Scheit, K.H. & Bell, J.C. The Clk2 and Clk3 dual-specificity protein kinases regulate the intranuclear distribution of SR proteins and influence pre-mRNA splicing. Exp. Cell Res. 241, 300–308 (1998).
Gross, T. et al. Functional analysis of the fission yeast Prp4 protein kinase involved in pre-mRNA splicing and isolation of a putative mammalian homologue. Nucleic Acids Res. 25, 1028–1035 (1997).
Roth, M.B., Zahler, A.M. & Stolk, J.A. A conserved family of nuclear phosphoproteins localized to sites of polymerase II transcription. J. Cell Biol. 115, 587–596 (1991).
Graveley, B.R. Sorting out the complexity of SR protein functions. RNA 6, 1197–1211 (2000).
Zhou, Z. & Fu, X.-D. Regulation of splicing by SR proteins and SR protein-specific kinases. Chromosoma 122, 191–207 (2013).
Cao, D.-S. et al. Large-scale prediction of human kinase-inhibitor interactions using protein sequences and molecular topological structures. Anal. Chim. Acta 792, 10–18 (2013).
Xiao, S.H. & Manley, J.L. Phosphorylation of the ASF/SF2 RS domain affects both protein-protein and protein-RNA interactions and is necessary for splicing. Genes Dev. 11, 334–344 (1997).
Xiao, S.H. & Manley, J.L. Phosphorylation-dephosphorylation differentially affects activities of splicing factor ASF/SF2. EMBO J. 17, 6359–6367 (1998).
Cao, W., Jamison, S.F. & Garcia-Blanco, M.A. Both phosphorylation and dephosphorylation of ASF/SF2 are required for pre-mRNA splicing in vitro. RNA 3, 1456–1467 (1997).
Tacke, R., Chen, Y. & Manley, J.L. Sequence-specific RNA binding by an SR protein requires RS domain phosphorylation: creation of an SRp40-specific splicing enhancer. Proc. Natl. Acad. Sci. USA 94, 1148–1153 (1997).
Yeakley, J.M. et al. Phosphorylation regulates in vivo interaction and molecular targeting of serine/arginine-rich pre-mRNA splicing factors. J. Cell Biol. 145, 447–455 (1999).
Roscigno, R.F. & Garcia-Blanco, M.A. SR proteins escort the U4/U6.U5 tri-snRNP to the spliceosome. RNA 1, 692–706 (1995).
Boutz, P.L., Bhutkar, A. & Sharp, P.A. Detained introns are a novel, widespread class of post-transcriptionally spliced introns. Genes Dev. 29, 63–80 (2015).
Zhou, Z. et al. The Akt-SRPK-SR axis constitutes a major pathway in transducing EGF Signaling to regulate alternative splicing in the nucleus. Mol. Cell 47, 422–433 (2012).
Bishop, A.C. et al. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature 407, 395–401 (2000).
Inada, M. & Pleiss, J.A. Genome-wide approaches to monitor pre-mRNA splicing. Methods Enzymol. 470, 51–75 (2010).
Hertz, N.T. et al. Chemical genetic approach for kinase-substrate mapping by covalent capture of thiophosphopeptides and analysis by mass spectrometry. Curr. Protoc. Chem. Biol. 2, 15–36 (2010).
Colwill, K. et al. SRPK1 and Clk/Sty protein kinases show distinct substrate specificities for serine/arginine-rich splicing factors. J. Biol. Chem. 271, 24569–24575 (1996).
Aubol, B.E. et al. Partitioning RS domain phosphorylation in an SR protein through the CLK and SRPK protein kinases. J. Mol. Biol. 425, 2894–2909 (2013).
Tang, Z. et al. Interacting factors and cellular localization of SR protein-specific kinase Dsk1. Exp. Cell Res. 318, 2071–2084 (2012).
Tang, Z., Yanagida, M. & Lin, R.J. Fission yeast mitotic regulator Dsk1 is an SR protein-specific kinase. J. Biol. Chem. 273, 5963–5969 (1998).
Zhang, Y. et al. Structure, phosphorylation and U2AF65 binding of the N-terminal domain of splicing factor 1 during 3′-splice site recognition. Nucleic Acids Res. 41, 1343–1354 (2013).
Krämer, A. & Utans, U. Three protein factors (SF1, SF3 and U2AF) function in pre-splicing complex formation in addition to snRNPs. EMBO J. 10, 1503–1509 (1991).
Rain, J.C., Rafi, Z., Rhani, Z., Legrain, P. & Krämer, A. Conservation of functional domains involved in RNA binding and protein-protein interactions in human and Saccharomyces cerevisiae pre-mRNA splicing factor SF1. RNA 4, 551–565 (1998).
Berglund, J.A., Abovich, N. & Rosbash, M. A cooperative interaction between U2AF65 and mBBP/SF1 facilitates branchpoint region recognition. Genes Dev. 12, 858–867 (1998).
Berglund, J.A., Chua, K., Abovich, N., Reed, R. & Rosbash, M. The splicing factor BBP interacts specifically with the pre-mRNA branchpoint sequence UACUAAC. Cell 89, 781–787 (1997).
Berglund, J.A., Fleming, M.L. & Rosbash, M. The KH domain of the branchpoint sequence binding protein determines specificity for the pre-mRNA branchpoint sequence. RNA 4, 998–1006 (1998).
Huang, T., Vilardell, J. & Query, C.C. Pre-spliceosome formation in S. pombe requires a stable complex of SF1–U2AF(59)-U2AF(23). EMBO J. 21, 5516–5526 (2002).
Liu, Z. et al. Structural basis for recognition of the intron branch site RNA by splicing factor 1. Science 294, 1098–1102 (2001).
Jacewicz, A., Chico, L., Smith, P., Schwer, B. & Shuman, S. Structural basis for recognition of intron branchpoint RNA by yeast Msl5 and selective effects of interfacial mutations on splicing of yeast pre-mRNAs. RNA 21, 401–414 (2015).
Manceau, V. et al. Major phosphorylation of SF1 on adjacent Ser-Pro motifs enhances interaction with U2AF65. FEBS J. 273, 577–587 (2006).
Wang, W. et al. Structure of phosphorylated SF1 bound to U2AF65 in an essential splicing factor complex. Structure 21, 197–208 (2013).
Stamm, S. Regulation of alternative splicing by reversible protein phosphorylation. J. Biol. Chem. 283, 1223–1227 (2008).
Aubol, B.E. et al. Processive phosphorylation of alternative splicing factor/splicing factor 2. Proc. Natl. Acad. Sci. USA 100, 12601–12606 (2003).
Ngo, J.C.K. et al. A sliding docking interaction is essential for sequential and processive phosphorylation of an SR protein by SRPK1. Mol. Cell 29, 563–576 (2008).
Cho, S. et al. Interaction between the RNA binding domains of Ser-Arg splicing factor 1 and U1–70K snRNP protein determines early spliceosome assembly. Proc. Natl. Acad. Sci. USA 108, 8233–8238 (2011).
Shin, C., Feng, Y. & Manley, J.L. Dephosphorylated SRp38 acts as a splicing repressor in response to heat shock. Nature 427, 553–558 (2004).
Huang, Y., Gattoni, R., Stévenin, J. & Steitz, J.A. SR splicing factors serve as adapter proteins for TAP-dependent mRNA export. Mol. Cell 11, 837–843 (2003).
Gao, Q. et al. Evaluation of cancer dependence and druggability of PRP4 kinase using cellular, biochemical, and structural approaches. J. Biol. Chem. 288, 30125–30138 (2013).
Rutz, B. & Séraphin, B. Transient interaction of BBP/ScSF1 and Mud2 with the splicing machinery affects the kinetics of spliceosome assembly. RNA 5, 819–831 (1999).
Rutz, B. & Séraphin, B. A dual role for BBP/ScSF1 in nuclear pre-mRNA retention and splicing. EMBO J. 19, 1873–1886 (2000).
Jang, S.-W. et al. Interaction of Akt-phosphorylated SRPK2 with 14–3–3 mediates cell cycle and cell death in neurons. J. Biol. Chem. 284, 24512–24525 (2009).
Potashkin, J., Naik, K. & Wentz-Hunter, K. U2AF homolog required for splicing in vivo. Science 262, 573–575 (1993).
Smyth, G.K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. App. Genet. Mol. Biol. 3, Article3 (2004).
Durinck, S. et al. BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21, 3439–3440 (2005).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Plass, M., Agirre, E., Reyes, D., Camara, F. & Eyras, E. Co-evolution of the branch site and SR proteins in eukaryotes. Trends Genet. 24, 590–594 (2008).
Acknowledgements
We thank J. Pleiss for designing the S. pombe splicing microarrays; N. Hertz, R. Levin and A. Burlingame for assistance with mass spectrometry; R. Freilich for assistance with strain construction; T. Tani (Kumamoto University) for providing us with the prp2-1 S. pombe strain; and M. Sattler (Technical University Munich) for the SF1 expression plasmid. We are grateful to D. Ruggero and members of the Guthrie and Shokat laboratories for helpful discussions and critical reading of the manuscript. Mass spectrometry was provided by the Bio-Organic Biomedical Mass Spectrometry Resource at University of California, San Francisco (A. Burlingame) supported by the Biomedical Technology Research Centers program of the US National Institutes of Health (NIH) National Institute of General Medical Sciences, 8P41GM103481. J.J.L. received a postdoctoral fellowship from the Jane Coffin Childs Memorial Fund for Medical Research. This work was supported by the Samuel Waxman Cancer Research Foundation (CA-0052023; K.M.S.) and NIH grants R01GM021119 (C.G.), F32GM101764 (M.C.M.) and R01AI094098 (K.M.S.). C.G. is supported as an American Cancer Society Research Professor of Molecular Genetics.
Author information
Authors and Affiliations
Contributions
J.J.L. and M.C.M. designed, performed and analyzed the experiments. J.J.L., M.C.M., K.M.S. and C.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of S. pombe splicing microarrays.
(A) Western blot of in vitro kinase assay analyzing the phosphorylation of myelin basic protein (MPB) by recombinant Lkh1-wild type (wt) (left) and Lkh1-analog-sensitive (as) (right) in the presence of various concentrations of the inhibitor 3-MB-PP1.
(B) Western blot of in vitro kinase assay analyzing the auto-phosphorylation of recombinant Prp4-wt (left) and Prp4-as (right) in the presence of various concentrations of the inhibitor 3-MB-PP1.
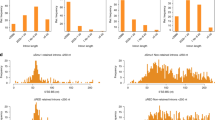
(C) Bar graphs showing the growth rate of dsk1-as, lkh1-as, and prp4-as strains in the presence of either DMSO, 10 μM 3-BrB-PP1 (3BrB), or 30 μM 3BrB relative to DMSO-treated control strains.
(D) Schematic representation of probe types in the splicing microarray: Exon (E), Intron (I), and Junction (J). Each splicing event is measured a combination score (S).
(E) Heatmap of Pearson correlation coefficients (ρ) between pair-wise combinations of the three probe types (E, I, J). Control represents microarray data comparing two wt strains.
(F) Bar graphs analyzing the average log2 value of exon probes (light grey) and score (dark grey) for introns for which no significant change of splicing was observed (not significant) or which were spliced significantly worse upon kinase inhibition. Results for dsk1 (left) and prp4 (right) are shown.
(G) Bar graphs of the fold enrichment of fkh1 intron 1 pre-mRNA upon inhibition of S. pombe splicing kinases Dsk1, Lkh1, and Prp4 as analyzed by (RT-qPCR). Error bars represent s.e.m. (n = 3 cell cultures). “n.s.” P > 0.05; “*” P < 0.05; “**” P < 0.01; “***” P < 0.001 by two-sided Student’s t-test.
(H) Heatmap of clustered scores for lkh1Δ. See Fig. 1B for detailed description. The “control” column is based on the same data used in Fig. 1B.
Supplementary Figure 2 Splicing defects of Dsk1 substrates Bpb1, Srp1, and Srp2, analyzed by RT-qPCR.
(A) Bar graphs of the fold enrichment of rpl25a intron 1 pre-mRNA in phosphomutant strains of S. pombe SR proteins Srp1 and Srp2 as analyzed by RT-qPCR. Values were normalized to their respective controls. Error bars represent s.e.m. (n = 3 cell cultures). “*” P < 0.05; “**” P < 0.01; “***” P < 0.001 by two-sided Student’s t-test.
(B) Same as in (A) but analyzing the fkh1 intron 3 in bpb1-wt and bpb1-2A strains.
Supplementary Figure 3 Splice sites of introns affected in dsk1-as and bpb1-2A are predicted to bind less well to U1 and U2 snRNAs.
(A) Bar graphs showing the predicted free energy upon heterodimer formation between the 5’ splice site (5’ SS) and the U1-snRNA (left panel) and the branchpoint sequence (BPS) and the U2-snRNA (right panel) for significantly affected introns accumulated with dsk1-as, bpb1-2A, prp2-1. The graph displays the differences of the means between unchanged and significantly retained introns for the three genotypes. Error bars represent s.e.m.
(B) Heatmap of hierarchically clustered scores for prp2-1 grown at restrictive (37 ˚C) versus permissive (25 ˚C) temperature. See Fig 1B for detailed description. The “control” column is based on the same data used in Fig 1B.
(C) Bar graphs of the fold enrichment of kes1 intron 5 pre-mRNA using RT-qPCR. Values were normalized to their respective controls. Error bars represent s.e.m. (n = 3 cell cultures). “*” P < 0.05; “**” P < 0.01; “***” P < 0.001 by two-sided Student’s t-test.
Supplementary Figure 4 Binding of SF1 to a different suboptimal branchpoint sequence is also promoted by phosphorylation.
(A) Native gel-shift assays using unphosphorylated or phosphorylated versions of recombinant SF1 (1-260) and U6 RNA with suboptimal (UUCUAAC) branchpoint sequence. The gel-shifts with consensus RNA from Fig. 5C are shown again for comparison.
(B) Quantification of fraction of bound over total RNA at various concentrations of SF1. Error bars represent s.e.m. (n = 4, independent experiments).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 7 and 8 (PDF 2382 kb)
Supplementary Data Set 1
Raw western blot exposures (PDF 6467 kb)
Supplementary Table 1
Dsk1 substrates (XLSX 79 kb)
Supplementary Table 2
Lkh1 substrates (XLSX 67 kb)
Supplementary Table 3
Srpk2 substrates (XLSX 99 kb)
Supplementary Table 4
Clk1 substrates (XLSX 80 kb)
Supplementary Table 5
Dsk1 substrates GO term enrichment (XLSX 39 kb)
Supplementary Table 6
Features for logistic regression modeling (XLSX 52 kb)
Rights and permissions
About this article
Cite this article
Lipp, J., Marvin, M., Shokat, K. et al. SR protein kinases promote splicing of nonconsensus introns. Nat Struct Mol Biol 22, 611–617 (2015). https://doi.org/10.1038/nsmb.3057
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3057