Abstract
Sexual development in Schizosaccharomyces pombe culminates in meiosis and sporulation. We used ribosome profiling to investigate the translational landscape of this process. We show that the translation efficiency of hundreds of genes is regulated in complex patterns, often correlating with changes in RNA levels. Ribosome-protected fragments show a three-nucleotide periodicity that identifies translated sequences and their reading frame. Using this property, we identified 46 new translated genes and found that 24% of noncoding RNAs are actively translated. We also detected 19 nested antisense genes, in which both DNA strands encode translated mRNAs. Finally, we identified 1,735 translated upstream open reading frames (ORFs) in leader sequences. In S. pombe, in contrast with Saccharomyces cerevisiae, sexual development is not accompanied by large increases in upstream ORF use, thus suggesting that this is an organism-specific adaptation, not a general feature of developmental processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Yamamoto, M., Imai, I. & Watanabe, Y. in The Molecular and Cellular Biology of the Yeast Saccharomyces: Life Cycle and Cell Biology (eds. Pringle, J.R., Broach, J.R. & Jones, E.W.) 1035–1106 (Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, USA, 1997).
Mata, J., Lyne, R., Burns, G. & Bähler, J. The transcriptional program of meiosis and sporulation in fission yeast. Nat. Genet. 32, 143–147 (2002).
Horie, S. et al. The Schizosaccharomyces pombe mei4+ gene encodes a meiosis-specific transcription factor containing a forkhead DNA-binding domain. Mol. Cell. Biol. 18, 2118–2129 (1998).
Mata, J., Wilbrey, A. & Bähler, J. Transcriptional regulatory network for sexual differentiation in fission yeast. Genome Biol. 8, R217 (2007).
Sugimoto, A., Iino, Y., Maeda, T., Watanabe, Y. & Yamamoto, M. Schizosaccharomyces pombe ste11+ encodes a transcription factor with an HMG motif that is a critical regulator of sexual development. Genes Dev. 5, 1990–1999 (1991).
Amorim, M.J., Cotobal, C., Duncan, C. & Mata, J. Global coordination of transcriptional control and mRNA decay during cellular differentiation. Mol. Syst. Biol. 6, 380 (2010).
Harigaya, Y. et al. Selective elimination of messenger RNA prevents an incidence of untimely meiosis. Nature 442, 45–50 (2006).
Bitton, D.A. et al. Programmed fluctuations in sense/antisense transcript ratios drive sexual differentiation in S. pombe. Mol. Syst. Biol. 7, 559 (2011).
Rhind, N. et al. Comparative functional genomics of the fission yeasts. Science 332, 930–936 (2011).
Wilhelm, B.T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453, 1239–1243 (2008).
Ingolia, N.T., Ghaemmaghami, S., Newman, J.R. & Weissman, J.S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Michel, A.M. & Baranov, P.V. Ribosome profiling: a Hi-Def monitor for protein synthesis at the genome-wide scale. Wiley Interdiscip. Rev. RNA 4, 473–490 (2013).
Brar, G.A. et al. High-resolution view of the yeast meiotic program revealed by ribosome profiling. Science 335, 552–557 (2012).
Ingolia, N.T., Lareau, L.F. & Weissman, J.S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Chew, G.L. et al. Ribosome profiling reveals resemblance between long non-coding RNAs and 5′ leaders of coding RNAs. Development 140, 2828–2834 (2013).
Guttman, M., Russell, P., Ingolia, N.T., Weissman, J.S. & Lander, E.S. Ribosome profiling provides evidence that large noncoding RNAs do not encode proteins. Cell 154, 240–251 (2013).
Michel, A.M. et al. Observation of dually decoded regions of the human genome using ribosome profiling data. Genome Res. 22, 2219–2229 (2012).
Bazzini, A.A. et al. Identification of small ORFs in vertebrates using ribosome footprinting and evolutionary conservation. EMBO J. 33, 981–993 (2014).
Iino, Y. & Yamamoto, M. Mutants of Schizosaccharomyces pombe which sporulate in the haploid state. Mol. Gen. Genet. 198, 416–421 (1985).
Nurse, P. Mutants of the fission yeast Schizosacharomyces pombe which alter the shift between cell proliferation and sporulation. Mol. Gen. Genet. 198, 497–502 (1985).
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Mata, J. Genome-wide mapping of polyadenylation sites in fission yeast reveals widespread alternative polyadenylation. RNA Biol. 10, 1407–1414 (2013).
Marguerat, S. et al. Quantitative analysis of fission yeast transcriptomes and proteomes in proliferating and quiescent cells. Cell 151, 671–683 (2012).
Lackner, D.H. et al. A network of multiple regulatory layers shapes gene expression in fission yeast. Mol. Cell 26, 145–155 (2007).
Matia-González, A.M., Hasan, A., Moe, G.H., Mata, J. & Rodriguez-Gabriel, M.A. Functional characterization of Upf1 targets in Schizosaccharomyces pombe. RNA Biol. 10, 1057–1065 (2013).
Isken, O. et al. Upf1 phosphorylation triggers translational repression during nonsense-mediated mRNA decay. Cell 133, 314–327 (2008).
Zhang, Z. et al. Nonsense-mediated decay targets have multiple sequence-related features that can inhibit translation. Mol. Syst. Biol. 6, 442 (2010).
Lackner, D.H., Schmidt, M.W., Wu, S., Wolf, D.A. & Bahler, J. Regulation of transcriptome, translation, and proteome in response to environmental stress in fission yeast. Genome Biol. 13, R25 (2012).
MacKay, V.L. et al. Gene expression analyzed by high-resolution state array analysis and quantitative proteomics: response of yeast to mating pheromone. Mol. Cell. Proteomics 3, 478–489 (2004).
Preiss, T., Baron-Benhamou, J., Ansorge, W. & Hentze, M.W. Homodirectional changes in transcriptome composition and mRNA translation induced by rapamycin and heat shock. Nat. Struct. Biol. 10, 1039–1047 (2003).
Wood, V. et al. PomBase: a comprehensive online resource for fission yeast. Nucleic Acids Res. 40, D695–D699 (2012).
Wood, V. et al. The genome sequence of Schizosaccharomyces pombe. Nature 415, 871–880 (2002).
Ma, J., Dobry, C.J., Krysan, D.J. & Kumar, A. Unconventional genomic architecture in the budding yeast Saccharomyces cerevisiae masks the nested antisense gene NAG1. Eukaryot. Cell 7, 1289–1298 (2008).
Galindo, M.I., Pueyo, J.I., Fouix, S., Bishop, S.A. & Couso, J.P. Peptides encoded by short ORFs control development and define a new eukaryotic gene family. PLoS Biol. 5, e106 (2007).
Kondo, T. et al. Small peptide regulators of actin-based cell morphogenesis encoded by a polycistronic mRNA. Nat. Cell Biol. 9, 660–665 (2007).
Magny, E.G. et al. Conserved regulation of cardiac calcium uptake by peptides encoded in small open reading frames. Science 341, 1116–1120 (2013).
Kelly, M., Burke, J., Smith, M., Klar, A. & Beach, D. Four mating-type genes control sexual differentiation in the fission yeast. EMBO J. 7, 1537–1547 (1988).
Hood, H.M., Neafsey, D.E., Galagan, J. & Sachs, M.S. Evolutionary roles of upstream open reading frames in mediating gene regulation in fungi. Annu. Rev. Microbiol. 63, 385–409 (2009).
Jackson, R.J., Hellen, C.U. & Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 11, 113–127 (2010).
Slavoff, S.A. et al. Peptidomic discovery of short open reading frame–encoded peptides in human cells. Nat. Chem. Biol. 9, 59–64 (2013).
Shimoda, C. et al. Cloning and analysis of transcription of the mei2 gene responsible for initiation of meiosis in the fission yeast Schizosaccharomyces pombe. J. Bacteriol. 169, 93–96 (1987).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Thorvaldsdóttir, H., Robinson, J.T. & Mesirov, J.P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
de Hoon, M.J., Imoto, S., Nolan, J. & Miyano, S. Open source clustering software. Bioinformatics 20, 1453–1454 (2004).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Saldanha, A.J. Java Treeview: extensible visualization of microarray data. Bioinformatics 20, 3246–3248 (2004).
Acknowledgements
We thank J. Bähler and R. Jackson for comments on the manuscript. This work was funded by Biotechnology and Biological Sciences Research Council (UK) research grant BB/J007153/1 (J.M.).
Author information
Authors and Affiliations
Contributions
C.D.S.D. and J.M. conceived the study and designed the experiments. C.D.S.D. performed all the experiments, and J.M. did the bioinformatics analyses. C.D.S.D. and J.M. wrote the manuscript and have critically read and approved of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Landmarks of sexual differentiation and distribution of reads across mRNA features.
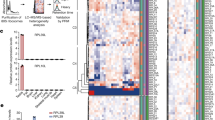
(A) Kinetics of sexual differentiation. Diploid cells are synchronized in G1 by nitrogen starvation and meiosis is induced at time 0 by heat inactivation of thermo-sensitive Pat1. S phase (S) takes place 2 hours after meiotic induction. The meiotic divisions occur at 5 hours (MI) and 5.5 hours (M II). The precursor to the spore plasma membrane (forespore membrane, F) is generated between 5.5 and 6.5 hours, and the spore cell wall appears at 10-12 hours (Spo). The progression of the meiotic divisions was followed by staining cells with DAPI followed by quantification of cells with 1 nucleus (before the meiotic divisions), 2 nuclei (after the first division) and four nuclei (after the second division). The data correspond to a single experiment. (B) Distribution of reads across RNA features. The fraction of reads mapping to coding sequences (CDS), 5' leader regions and 3'-UTRs is shown for RPFs and mRNAs. (Left) Unnormalized data. (Right) Data normalized by the length of the corresponding genomic feature.
Supplementary Figure 2 Reproducibility of RPF and mRNA measurements.
(A) Scatter plot comparing mRNA read counts (in reads per million) for every CDS for two independent biological repeats of pat1 cells 3 hours (left) or 5 hours (right) after meiotic induction. No filtering was applied to the data. Pearson correlation coefficients are indicated inside the boxes. (B) As in A, but showing RPF counts. (C) As in A, but displaying TEs, calculated by normalizing RPF counts for every CDS by the corresponding number of mRNA counts. Data are shown only for genes that had at least 10 reads in both the mRNA and the RPF data. Note that these are completely independent samples, as the two cultures were harvested on different days, and the libraries were prepared and sequenced separately.
Supplementary Figure 3 Global changes in translation and mRNA levels during meiosis.
(A) The 1,719 genes showing strongest changes in expression during a meiotic time course were clustered based on their RPF profiles. The heat map shows mRNA data based on a published microarray experiment, mRNA-seq data from this work (middle) and RPF changes (right). The colors represent log ratios of gene expression compared to the corresponding vegetative cells. (B) Correlations between the TE of uORFs and their corresponding ORFs. Boxplots displaying the log2 ratios between TEs of every translated uORF and its downstream ORF. TEs were calculated for each time point and normalized by the mean TE over the time course. Veg indicates cells growing vegetatively, and the time is the number of hours since the temperature-shift to induce meiosis. (C) Translation of the SPBPB10D8.07c mRNA. The y axis represents the TE for the uORF (red) and the corresponding ORF (blue) for each time point of the meiotic time course. SPBPB10D8.07c displays high correlation between TE changes of the uORF and ORF (Pearson coefficient = 0.99). (D) Translation of the SPBC1773.04 mRNA, which showed a negative correlation between uORF and ORF TEs (Pearson coefficient = −0.74). Labelling as in (C).
Supplementary Figure 4 Improvements to fission-yeast gene annotation.
(A) Periodicity scores separate mRNA and RPF data. Distribution of periodicity scores in frame number 1 of all CDS for mRNA fragments (blue) and RPFs (red). We chose a threshold of 0.6 to define newly discovered ORFs as translated (arrow). Only 0.1% of CDS in the mRNA dataset show a periodicity score over this threshold (in any reading frame). Genes with low periodicity scores in the RPF data are potentially mis-annotated. For example, SPAC823.02 shows a periodicity score of 0.15 in frame 1, but a clear enrichment in frame 3 (0.70). (B) Translational profile of SPAC823.02. The heat maps show mRNA and RPF levels across meiosis, starting with vegetative cells and progressing downwards. The brightness of the red color is proportional to the read number. The schemes show the position of the UTRs (lines) and the ORF (boxes) for the proposed new structures and the annotated models. Translation is predicted to occur at different frames from those annotated for both exons. (C) Translational profile of tom7, labelled as in B. (D) Translational profile of pis1, annotated as in B. In pis1 RPF data demonstrates the existence of two mRNA isoforms created by the splicing or retention of an intron. The gene contains two cleavage and polyadenylation sites (P1 and P2), one of which is located inside the intron, suggesting that intron retention may be associated with a shorter mRNA. The proposed splicing models were validated by the existence of 'trans' reads that spanned exon-exon junctions (green line, Read 1) or exon-intron junctions (green line, Read 2). All three new predicted proteins (B-D) showed high similarity to proteins from other Schizosaccharomyces species.
Supplementary Figure 5 Polymorphism of the nup184 gene and translation of new genes.
A. Alignment of the 3' end sequence of the nup184 gene of S. pombe, S. cryophilus and S. octosporus. The S. pombe reference sequence (from the 972 h- strain) contains a single nucleotide deletion (yellow) that creates an in-frame stop codon (green) that should prevent production of the C-terminal part of the Nup184 protein. B. Polymorphisms in the nup184 locus. Our wild type strain (968 h90) contains an insertion of a C at this position (yellow) that reverts the effect of the deletion. In the pat1 diploid, one chromosome appeared to have the insertion, while the second one contained the deletion associated with two upstream single nucleotide polymorphisms (green). Translation downstream of the predicted stop codon takes place in a frame consistent with the presence of the insertion. (C) Length distribution of novel translated regions (in nucleotides).
Supplementary Figure 6 Examples of nested antisense genes and overlapping uORFs.
(A) Translation of the spt16 locus. The heat maps show mRNA (top) and RPF levels (bottom) across the meiotic time course, starting with vegetative cells at the top and progressing downwards. The top graphs correspond to the sense strand, and the bottom ones to the antisense. The brightness of the red color is proportional to the number of reads. The scheme shows the position of the UTRs (arrows) and the ORF (boxes). LS corresponds to the long annotated sense gene and AS to the novel antisense translated coding sequence. (B) Triplet periodicity for the LS and AS forms of the spt16 locus. The graphs show the fraction of codons in which a majority of the reads map to nucleotides 1, 2 or 3 for the AS and LS ORFs. (C) Translation of the SPCC1235.01 locus. The top graphs shows the triplet periodicity across the gene calculated for a running window of 35 codons, and represented as the fraction of codons with a majority of reads in nucleotide 1, 2 or 3 within a codon. The heat maps are labelled as in (A). L corresponds to the annotated gene and S to the predicted overlapping uORFs. Note the change in reading frame as the window moves from one ORF to the next one. S overlaps with L and is more heavily translated, and is thus the dominant frame. P1 and P2 represent cleavage and polyadenylation sites. The fact that the 3' region of the gene contains a lower number of mRNA reads and the presence of an internal cleavage site (P1) suggests the existence of two different isoforms of the gene. (B) Translation of the SPAC6G9.05 locus. Annotations as in (B).
Supplementary Figure 8 Triplet periodicity of mRNA and RPF reads.
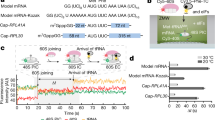
(A) Bar charts displaying the fraction of reads in which nucleotide 13 corresponded to the first, second or third nucleotide within a codon, calculated for all reads that mapped to annotated coding sequences. Only reads of exactly 28 nucleotides are shown. Veg indicates cells growing vegetatively, and the time indicates the number of hours since the temperature-shift to induce meiosis. Data for mRNA reads are displayed on the left, and those for RPFs on the right. (B) As in (A), but the reads were split according to their length (from 25 to 32 nucleotides) and plotted separately. Only reads of the 1 hour meiotic time point are shown, but similar patterns were obtained for every sample. Note the high periodicity of reads of 28 to 30 nucleotides, which were selected for further analysis.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1568 kb)
Supplementary Tables
Supplementary Tables 1–10 (XLSX 496 kb)
Rights and permissions
About this article
Cite this article
Duncan, C., Mata, J. The translational landscape of fission-yeast meiosis and sporulation. Nat Struct Mol Biol 21, 641–647 (2014). https://doi.org/10.1038/nsmb.2843
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2843