Abstract
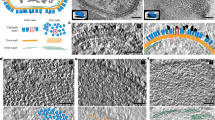
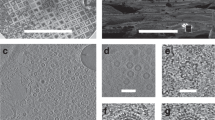
The 20S particle, which is composed of the N-ethylmaleimide–sensitive factor (NSF), soluble NSF attachment proteins (SNAPs) and the SNAP receptor (SNARE) complex, has an essential role in intracellular vesicle fusion events. Using single-particle cryo-EM and negative stain EM, we reconstructed four related three-dimensional structures: Chinese hamster NSF hexamer in the ATPγS, ADP-AlFx and ADP states, and the 20S particle. These structures reveal a parallel arrangement between the D1 and D2 domains of the hexameric NSF and characterize the nucleotide-dependent conformational changes in NSF. The structure of the 20S particle shows that it holds the SNARE complex at two interaction interfaces around the C terminus and N-terminal half of the SNARE complex, respectively. These findings provide insight into the molecular mechanism underlying disassembly of the SNARE complex by NSF.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Rothman, J.E. Mechanisms of intracellular protein transport. Nature 372, 55–63 (1994).
Jahn, R. & Scheller, R.H. SNAREs—engines for membrane fusion. Nat. Rev. Mol. Cell Biol. 7, 631–643 (2006).
Söllner, T. et al. SNAP receptors implicated in vesicle targeting and fusion. Nature 362, 318–324 (1993).
Sutton, R.B., Fasshauer, D., Jahn, R. & Brunger, A.T. Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 Å resolution. Nature 395, 347–353 (1998).
Stein, A., Weber, G., Wahl, M.C. & Jahn, R. Helical extension of the neuronal SNARE complex into the membrane. Nature 460, 525–528 (2009).
Antonin, W., Fasshauer, D., Becker, S., Jahn, R. & Schneider, T.R. Crystal structure of the endosomal SNARE complex reveals common structural principles of all SNAREs. Nat. Struct. Biol. 9, 107–111 (2002).
Weber, T. et al. SNAREpins: minimal machinery for membrane fusion. Cell 92, 759–772 (1998).
Söllner, T., Bennett, M.K., Whiteheart, S.W., Scheller, R.H. & Rothman, J.E. A protein assembly-disassembly pathway in vitro that may correspond to sequential steps of synaptic vesicle docking, activation, and fusion. Cell 75, 409–418 (1993).
Mayer, A., Wickner, W. & Haas, A. Sec18p (NSF)-driven release of Sec17p (alpha-SNAP) can precede docking and fusion of yeast vacuoles. Cell 85, 83–94 (1996).
Wilson, D.W., Whiteheart, S.W., Wiedmann, M., Brunner, M. & Rothman, J.E. A multisubunit particle implicated in membrane fusion. J. Cell Biol. 117, 531–538 (1992).
Hanson, P.I. & Whiteheart, S.W. AAA+ proteins: have engine, will work. Nat. Rev. Mol. Cell Biol. 6, 519–529 (2005).
Ogura, T. & Wilkinson, A.J. AAA+ superfamily ATPases: common structure–diverse function. Genes Cells 6, 575–597 (2001).
Fleming, K.G. et al. A revised model for the oligomeric state of the N-ethylmaleimide-sensitive fusion protein, NSF. J. Biol. Chem. 273, 15675–15681 (1998).
Tagaya, M., Wilson, D.W., Brunner, M., Arango, N. & Rothman, J.E. Domain structure of an N-ethylmaleimide-sensitive fusion protein involved in vesicular transport. J. Biol. Chem. 268, 2662–2666 (1993).
Zhao, C., Smith, E.C. & Whiteheart, S.W. Requirements for the catalytic cycle of the N-ethylmaleimide-sensitive factor (NSF). Biochim. Biophys. Acta 1823, 159–171 (2012).
Nagiec, E.E., Bernstein, A. & Whiteheart, S.W. Each domain of the N-ethylmaleimide-sensitive fusion protein contributes to its transport activity. J. Biol. Chem. 270, 29182–29188 (1995).
Lenzen, C.U., Steinmann, D., Whiteheart, S.W. & Weis, W.I. Crystal structure of the hexamerization domain of N-ethylmaleimide-sensitive fusion protein. Cell 94, 525–536 (1998).
Yu, R.C., Hanson, P.I., Jahn, R. & Brunger, A.T. Structure of the ATP-dependent oligomerization domain of N-ethylmaleimide sensitive factor complexed with ATP. Nat. Struct. Biol. 5, 803–811 (1998).
May, A.P., Misura, K.M., Whiteheart, S.W. & Weis, W.I. Crystal structure of the amino-terminal domain of N-ethylmaleimide-sensitive fusion protein. Nat. Cell Biol. 1, 175–182 (1999).
Yu, R.C., Jahn, R. & Brunger, A.T. NSF N-terminal domain crystal structure: models of NSF function. Mol. Cell 4, 97–107 (1999).
Babor, S.M. & Fass, D. Crystal structure of the Sec18p N-terminal domain. Proc. Natl. Acad. Sci. USA 96, 14759–14764 (1999).
Hanson, P.I., Roth, R., Morisaki, H., Jahn, R. & Heuser, J.E. Structure and conformational changes in NSF and its membrane receptor complexes visualized by quick-freeze/deep-etch electron microscopy. Cell 90, 523–535 (1997).
Hohl, T.M. et al. Arrangement of subunits in 20S particles consisting of NSF, SNAPs, and SNARE complexes. Mol. Cell 2, 539–548 (1998).
Furst, J., Sutton, R.B., Chen, J., Brunger, A.T. & Grigorieff, N. Electron cryomicroscopy structure of N-ethyl maleimide sensitive factor at 11 Å resolution. EMBO J. 22, 4365–4374 (2003).
DeLaBarre, B. & Brunger, A.T. Complete structure of p97/valosin-containing protein reveals communication between nucleotide domains. Nat. Struct. Biol. 10, 856–863 (2003).
Huyton, T. et al. The crystal structure of murine p97/VCP at 3.6 Å. J. Struct. Biol. 144, 337–348 (2003).
Rice, L.M. & Brunger, A.T. Crystal structure of the vesicular transport protein Sec17: implications for SNAP function in SNARE complex disassembly. Mol. Cell 4, 85–95 (1999).
Suno, R. et al. Structure of the whole cytosolic region of ATP-dependent protease FtsH. Mol. Cell 22, 575–585 (2006).
Ortega, A., Amoros, D. & Garcia de la Torre, J. Prediction of hydrodynamic and other solution properties of rigid proteins from atomic- and residue-level models. Biophys. J. 101, 892–898 (2011).
Zhao, C. et al. Hexahistidine-tag-specific optical probes for analyses of proteins and their interactions. Anal. Biochem. 399, 237–245 (2010).
Rossi, G., Salminen, A., Rice, L.M., Brunger, A.T. & Brennwald, P. Analysis of a yeast SNARE complex reveals remarkable similarity to the neuronal SNARE complex and a novel function for the C terminus of the SNAP-25 homolog, Sec9. J. Biol. Chem. 272, 16610–16617 (1997).
Wimmer, C. et al. Molecular mass, stoichiometry, and assembly of 20S particles. J. Biol. Chem. 276, 29091–29097 (2001).
Winter, U., Chen, X. & Fasshauer, D. A conserved membrane attachment site in alpha-SNAP facilitates N-ethylmaleimide-sensitive factor (NSF)-driven SNARE complex disassembly. J. Biol. Chem. 284, 31817–31826 (2009).
Marz, K.E., Lauer, J.M. & Hanson, P.I. Defining the SNARE complex binding surface of alpha-SNAP: implications for SNARE complex disassembly. J. Biol. Chem. 278, 27000–27008 (2003).
Davies, J.M., Brunger, A.T. & Weis, W.I. Improved structures of full-length p97, an AAA ATPase: implications for mechanisms of nucleotide-dependent conformational change. Structure 16, 715–726 (2008).
Rouiller, I. et al. Conformational changes of the multifunction p97 p97 AAA ATPase during its ATPase cycle. Nat. Struct. Biol. 9, 950–957 (2002).
Rouiller, I., Butel, V.M., Latterich, M., Milligan, R.A. & Wilson-Kubalek, E.M. A major conformational change in p97 AAA ATPase upon ATP binding. Mol. Cell 6, 1485–1490 (2000).
Rockel, B., Jakana, J., Chiu, W. & Baumeister, W. Electron cryo-microscopy of VAT, the archaeal p97/CDC48 homologue from Thermoplasma acidophilum. J. Mol. Biol. 317, 673–681 (2002).
Beuron, F. et al. Motions and negative cooperativity between p97 domains revealed by cryo-electron microscopy and quantised elastic deformational model. J. Mol. Biol. 327, 619–629 (2003).
Davies, J.M., Tsuruta, H., May, A.P. & Weis, W.I. Conformational changes of p97 during nucleotide hydrolysis determined by small-angle X-Ray scattering. Structure 13, 183–195 (2005).
Beuron, F. et al. Conformational changes in the AAA ATPase p97-p47 adaptor complex. EMBO J. 25, 1967–1976 (2006).
Pye, V.E. et al. Going through the motions: the ATPase cycle of p97. J. Struct. Biol. 156, 12–28 (2006).
Tang, W.K. et al. A novel ATP-dependent conformation in p97 N-D1 fragment revealed by crystal structures of disease-related mutants. EMBO J. 29, 2217–2229 (2010).
Brunger, A.T. & DeLaBarre, B. NSF and p97/VCP: similar at first, different at last. FEBS Lett. 555, 126–133 (2003).
Baker, T.A. & Sauer, R.T. ClpXP, an ATP-powered unfolding and protein-degradation machine. Biochim. Biophys. Acta 1823, 15–28 (2012).
Effantin, G., Ishikawa, T., De Donatis, G.M., Maurizi, M.R. & Steven, A.C. Local and global mobility in the ClpA AAA+ chaperone detected by cryo-electron microscopy: functional connotations. Structure 18, 553–562 (2010).
Lee, S., Sielaff, B., Lee, J. & Tsai, F.T. CryoEM structure of Hsp104 and its mechanistic implication for protein disaggregation. Proc. Natl. Acad. Sci. USA 107, 8135–8140 (2010).
Bohn, S. et al. Structure of the 26S proteasome from Schizosaccharomyces pombe at subnanometer resolution. Proc. Natl. Acad. Sci. USA 107, 20992–20997 (2010).
May, A.P., Whiteheart, S.W. & Weis, W.I. Unraveling the mechanism of the vesicle transport ATPase NSF, the N-ethylmaleimide-sensitive factor. J. Biol. Chem. 276, 21991–21994 (2001).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Eswar, N. et al. Comparative protein structure modeling using Modeller. Curr. Protoc. Bioinformatics Chapter 5 Unit 5.6 (2006).
Wriggers, W. & Birmanns, S. Using situs for flexible and rigid-body fitting of multiresolution single-molecule data. J. Struct. Biol. 133, 193–202 (2001).
Trabuco, L.G., Villa, E., Schreiner, E., Harrison, C.B. & Schulten, K. Molecular dynamics flexible fitting: a practical guide to combine cryo-electron microscopy and X-ray crystallography. Methods 49, 174–180 (2009).
Pettersen, E.F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We are grateful to E.R. Chapman (University of Wisconsin, Madison), J. Rizo (University of Texas Southwestern Medical Center) for providing plasmids. We thank F. Zhang, N. Gao, J.-W. Wu, N. Yan, J. Wang and L. Cheng for helpful discussions. We also thank J. Lei for setting up a semiautomated data collection program and Y. Wu for providing electron microscope for our use at the early stage of this work. This work was supported by the National Basic Research Program of China (2011CB910500/2010CB833706/2010CB912400) and the National Natural Science Foundation of China (30830028).
Author information
Authors and Affiliations
Contributions
L.-F.C., S.C. and C.-C.L. conducted the experiments and analyzed the data with the help of X.P., J.J., X.-C.B. and X.X.; L.-F.C. prepared the figures; H.-W.W. commented on the experiments; J.J. and H.-W.W edited the manuscript; L.-F.C., S.C. and S.-F.S. planned the experiments and prepared the manuscript; S.-F.S. coordinated the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1 and 2, and Supplementary Methods (PDF 1827 kb)
Supplementary Movie 1
This movie shows the cryo-EM map of a NSF protomer in ADP-AlFx state. Crystal structures of D1 and D2 were flexibly fitted as in Figure 2a. (MOV 3514 kb)
Supplementary Movie 2
This movie shows the cryo-EM map of a NSF protomer in ADP state. Crystal structures of D1 and D2 were flexibly fitted as in Figure 3a. (MOV 3198 kb)
Supplementary Movie 3
This movie shows the linear morphing between the D1 hexamer (ribbon diagram of flexibly fitted structures) in the ADP-AlFx state and in the ADP state as shown in Figure 4d, which was generated by “morph conformations” program in Chimera. (MOV 1804 kb)
Rights and permissions
About this article
Cite this article
Chang, LF., Chen, S., Liu, CC. et al. Structural characterization of full-length NSF and 20S particles. Nat Struct Mol Biol 19, 268–275 (2012). https://doi.org/10.1038/nsmb.2237
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2237
This article is cited by
-
NSF/αSNAP2-mediated cis-SNARE complex disassembly precedes vesicle fusion in Arabidopsis cytokinesis
Nature Plants (2023)
-
Extreme parsimony in ATP consumption by 20S complexes in the global disassembly of single SNARE complexes
Nature Communications (2021)
-
Chaperoning SNARE assembly and disassembly
Nature Reviews Molecular Cell Biology (2016)
-
Cryo-EM structure of SNAP-SNARE assembly in 20S particle
Cell Research (2015)
-
Molecular snapshots of the Pex1/6 AAA+ complex in action
Nature Communications (2015)