Abstract
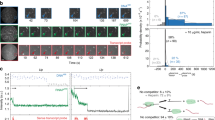
During gene expression, RNA polymerase (RNAP) encounters a major barrier at a nucleosome and yet must access the nucleosomal DNA. Previous in vivo evidence has suggested that multiple RNAPs might increase transcription efficiency through nucleosomes. Here we have quantitatively investigated this hypothesis using Escherichia coli RNAP as a model system by directly monitoring its location on the DNA via a single-molecule DNA-unzipping technique. When an RNAP encountered a nucleosome, it paused with a distinctive 10–base pair periodicity and backtracked by ∼10–15 base pairs. When two RNAPs elongate in close proximity, the trailing RNAP apparently assists in the leading RNAP's elongation, reducing its backtracking and enhancing its transcription through a nucleosome by a factor of 5. Taken together, our data indicate that histone-DNA interactions dictate RNAP pausing behavior, and alleviation of nucleosome-induced backtracking by multiple polymerases may prove to be a mechanism for overcoming the nucleosomal barrier in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Izban, M.G. & Luse, D.S. Transcription on nucleosomal templates by RNA polymerase II in vitro: inhibition of elongation with enhancement of sequence-specific pausing. Genes Dev. 5, 683–696 (1991).
Studitsky, V.M., Clark, D.J. & Felsenfeld, G. Overcoming a nucleosomal barrier to transcription. Cell 83, 19–27 (1995).
Studitsky, V.M., Kassavetis, G.A., Geiduschek, E.P. & Felsenfeld, G. Mechanism of transcription through the nucleosome by eukaryotic RNA polymerase. Science 278, 1960–1963 (1997).
Kireeva, M.L. et al. Nucleosome remodeling induced by RNA polymerase II: loss of the H2A/H2B dimer during transcription. Mol. Cell 9, 541–552 (2002).
Walter, W., Kireeva, M.L., Studitsky, V.M. & Kashlev, M. Bacterial polymerase and yeast polymerase II use similar mechanisms for transcription through nucleosomes. J. Biol. Chem. 278, 36148–36156 (2003).
Kireeva, M.L. et al. Nature of the nucleosomal barrier to RNA polymerase II. Mol. Cell 18, 97–108 (2005).
Bondarenko, V.A. et al. Nucleosomes can form a polar barrier to transcript elongation by RNA polymerase II. Mol. Cell 24, 469–479 (2006).
Ujvari, A., Hsieh, F.K., Luse, S.W., Studitsky, V.M. & Luse, D.S. Histone N-terminal tails interfere with nucleosome traversal by RNA polymerase II. J. Biol. Chem. 283, 32236–32243 (2008).
Hodges, C., Bintu, L., Lubkowska, L., Kashlev, M. & Bustamante, C. Nucleosomal fluctuations govern the transcription dynamics of RNA polymerase II. Science 325, 626–628 (2009).
Kulaeva, O.I. et al. Mechanism of chromatin remodeling and recovery during passage of RNA polymerase II. Nat. Struct. Mol. Biol. 16, 1272–1278 (2009).
O'Brien, T. & Lis, J.T. Rapid changes in Drosophila transcription after an instantaneous heat shock. Mol. Cell. Biol. 13, 3456–3463 (1993).
Tennyson, C.N., Klamut, H.J. & Worton, R.G. The human dystrophin gene requires 16 hours to be transcribed and is cotranscriptionally spliced. Nat. Genet. 9, 184–190 (1995).
Darzacq, X. et al. In vivo dynamics of RNA polymerase II transcription. Nat. Struct. Mol. Biol. 14, 796–806 (2007).
Awrey, D.E. et al. Yeast transcript elongation factor (TFIIS), structure and function. II: RNA polymerase binding, transcript cleavage, and read-through. J. Biol. Chem. 273, 22595–22605 (1998).
Weilbaecher, R.G., Awrey, D.E., Edwards, A.M. & Kane, C.M. Intrinsic transcript cleavage in yeast RNA polymerase II elongation complexes. J. Biol. Chem. 278, 24189–24199 (2003).
Galburt, E.A. et al. Backtracking determines the force sensitivity of RNAP II in a factor-dependent manner. Nature 446, 820–823 (2007).
Brown, C.E., Lechner, T., Howe, L. & Workman, J.L. The many HATs of transcription coactivators. Trends Biochem. Sci. 25, 15–19 (2000).
Anderson, J.D., Lowary, P.T. & Widom, J. Effects of histone acetylation on the equilibrium accessibility of nucleosomal DNA target sites. J. Mol. Biol. 307, 977–985 (2001).
Roth, S.Y., Denu, J.M. & Allis, C.D. Histone acetyltransferases. Annu. Rev. Biochem. 70, 81–120 (2001).
Belotserkovskaya, R. et al. FACT facilitates transcription-dependent nucleosome alteration. Science 301, 1090–1093 (2003).
Schwabish, M.A. & Struhl, K. The Swi/Snf complex is important for histone eviction during transcriptional activation and RNA polymerase II elongation in vivo. Mol. Cell. Biol. 27, 6987–6995 (2007).
Kim, T.H. et al. A high-resolution map of active promoters in the human genome. Nature 436, 876–880 (2005).
Lee, C.K., Shibata, Y., Rao, B., Strahl, B.D. & Lieb, J.D. Evidence for nucleosome depletion at active regulatory regions genome-wide. Nat. Genet. 36, 900–905 (2004).
Varv, S., Kristjuhan, K. & Kristjuhan, A. RNA polymerase II determines the area of nucleosome loss in transcribed gene loci. Biochem. Biophys. Res. Commun. 358, 666–671 (2007).
Epshtein, V. & Nudler, E. Cooperation between RNA polymerase molecules in transcription elongation. Science 300, 801–805 (2003).
Epshtein, V., Toulme, F., Rahmouni, A.R., Borukhov, S. & Nudler, E. Transcription through the roadblocks: the role of RNA polymerase cooperation. EMBO J. 22, 4719–4727 (2003).
Wang, M.D. et al. Force and velocity measured for single molecules of RNA polymerase. Science 282, 902–907 (1998).
Shundrovsky, A., Santangelo, T.J., Roberts, J.W. & Wang, M.D. A single-molecule technique to study sequence-dependent transcription pausing. Biophys. J. 87, 3945–3953 (2004).
Ebright, R.H. RNA polymerase: structural similarities between bacterial RNA polymerase and eukaryotic RNA polymerase II. J. Mol. Biol. 304, 687–698 (2000).
Korzheva, N. & Mustaev, A. Transcription elongation complex: structure and function. Curr. Opin. Microbiol. 4, 119–125 (2001).
Shundrovsky, A., Smith, C.L., Lis, J.T., Peterson, C.L. & Wang, M.D. Probing SWI/SNF remodeling of the nucleosome by unzipping single DNA molecules. Nat. Struct. Mol. Biol. 13, 549–554 (2006).
Hall, M.A. et al. High-resolution dynamic mapping of histone-DNA interactions in a nucleosome. Nat. Struct. Mol. Biol. 16, 124–129 (2009).
Jiang, J. et al. Detection of high-affinity and sliding clamp modes for MSH2–MSH6 by single-molecule unzipping force analysis. Mol. Cell 20, 771–781 (2005).
Nudler, E., Gusarov, I., Avetissova, E., Kozlov, M. & Goldfarb, A. Spatial organization of transcription elongation complex in Escherichia coli. Science 281, 424–428 (1998).
Korzheva, N. et al. A structural model of transcription elongation. Science 289, 619–625 (2000).
Zaychikov, E., Denissova, L. & Heumann, H. Translocation of the Escherichia coli transcription complex observed in the registers 11 to 20: “jumping” of RNA polymerase and asymmetric expansion and contraction of the “transcription bubble”. Proc. Natl. Acad. Sci. USA 92, 1739–1743 (1995).
Lowary, P.T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998).
Samkurashvili, I. & Luse, D.S. Translocation and transcriptional arrest during transcript elongation by RNA polymerase II. J. Biol. Chem. 271, 23495–23505 (1996).
Luger, K., Mader, A.W., Richmond, R.K., Sargent, D.F. & Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997).
Dong, F., Hansen, J.C. & van Holde, K.E. DNA and protein determinants of nucleosome positioning on sea urchin 5S rRNA gene sequences in vitro. Proc. Natl. Acad. Sci. USA 87, 5724–5728 (1990).
Peng, H.F. & Jackson, V. Measurement of the frequency of histone displacement during the in vitro transcription of nucleosomes: RNA is a competitor for these histones. Biochemistry 36, 12371–12382 (1997).
Bai, L., Shundrovsky, A. & Wang, M.D. Sequence-dependent kinetic model for transcription elongation by RNA polymerase. J. Mol. Biol. 344, 335–349 (2004).
Saeki, H. & Svejstrup, J.Q. Stability, flexibility, and dynamic interactions of colliding RNA polymerase II elongation complexes. Mol. Cell 35, 191–205 (2009).
Shaevitz, J.W., Abbondanzieri, E.A., Landick, R. & Block, S.M. Backtracking by single RNA polymerase molecules observed at near-base-pair resolution. Nature 426, 684–687 (2003).
Bai, L., Fulbright, R.M. & Wang, M.D. Mechanochemical kinetics of transcription elongation. Phys. Rev. Lett. 98, 068103 (2007).
Yankulov, K., Blau, J., Purton, T., Roberts, S. & Bentley, D.L. Transcriptional elongation by RNA polymerase II is stimulated by transactivators. Cell 77, 749–759 (1994).
Mason, P.B. & Struhl, K. Distinction and relationship between elongation rate and processivity of RNA polymerase II in vivo. Mol. Cell 17, 831–840 (2005).
Archambault, J., Lacroute, F., Ruet, A. & Friesen, J.D. Genetic interaction between transcription elongation factor TFIIS and RNA polymerase II. Mol. Cell. Biol. 12, 4142–4152 (1992).
Orlova, M., Newlands, J., Das, A., Goldfarb, A. & Borukhov, S. Intrinsic transcript cleavage activity of RNA polymerase. Proc. Natl. Acad. Sci. USA 92, 4596–4600 (1995).
Strobl, L.J. & Eick, D. Hold back of RNA polymerase II at the transcription start site mediates down-regulation of c-myc in vivo. EMBO J. 11, 3307–3314 (1992).
Benjamin, L.R. & Gilmour, D.S. Nucleosomes are not necessary for promoter-proximal pausing in vitro on the Drosophila hsp70 promoter. Nucleic Acids Res. 26, 1051–1055 (1998).
Conaway, J.W., Shilatifard, A., Dvir, A. & Conaway, R.C. Control of elongation by RNA polymerase II. Trends Biochem. Sci. 25, 375–380 (2000).
Cheng, C. & Sharp, P.A. RNA polymerase II accumulation in the promoter-proximal region of the dihydrofolate reductase and γ-actin genes. Mol. Cell. Biol. 23, 1961–1967 (2003).
Saunders, A., Core, L.J. & Lis, J.T. Breaking barriers to transcription elongation. Nat. Rev. Mol. Cell Biol. 7, 557–567 (2006).
Core, L.J. & Lis, J.T. Transcription regulation through promoter-proximal pausing of RNA polymerase II. Science 319, 1791–1792 (2008).
Orphanides, G. & Reinberg, D. RNA polymerase II elongation through chromatin. Nature 407, 471–475 (2000).
Koch, S.J., Shundrovsky, A., Jantzen, B.C. & Wang, M.D. Probing protein-DNA interactions by unzipping a single DNA double helix. Biophys. J. 83, 1098–1105 (2002).
Acknowledgements
We thank members of the Wang laboratory for critical reading of the manuscript, J. Widom (Northwestern University) for the plasmid containing the 601 NPE and the J. Roberts laboratory (Cornell University) for help with the phosphorescence gel scanner. M.D.W. wishes to acknowledge support from the US National Institutes of Health (GM059849), the US National Science Foundation (MCB-0820293), the Keck Foundation Distinguished Young Scholar in Medical Research Award and the Cornell Nanobiotechnology Center.
Author information
Authors and Affiliations
Contributions
J.J. designed and constructed transcription templates, designed and performed bulk and single-molecule transcription experiments, analyzed the data, participated in all discussions and proposal of the model and also wrote the manuscript; L.B., D.S.J. and M.L.K. offered suggestions and technical advice and helped troubleshoot the experiments; R.M.F. purified E. coli RNAP and histones and revised the manuscript; M.K. helped with the design of the project, offered advice on biochemical studies and contributed significantly to the manuscript revision; M.D.W. supervised the study throughout all stages of the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Discussion (PDF 3717 kb)
Rights and permissions
About this article
Cite this article
Jin, J., Bai, L., Johnson, D. et al. Synergistic action of RNA polymerases in overcoming the nucleosomal barrier. Nat Struct Mol Biol 17, 745–752 (2010). https://doi.org/10.1038/nsmb.1798
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1798
This article is cited by
-
When push comes to shove - RNA polymerase and DNA-bound protein roadblocks
Biophysical Reviews (2023)
-
Polarity of the CRISPR roadblock to transcription
Nature Structural & Molecular Biology (2022)
-
Optical tweezers in single-molecule biophysics
Nature Reviews Methods Primers (2021)
-
Helicase promotes replication re-initiation from an RNA transcript
Nature Communications (2018)
-
Single molecule high-throughput footprinting of small and large DNA ligands
Nature Communications (2017)