Abstract
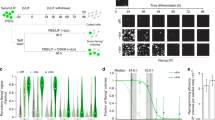
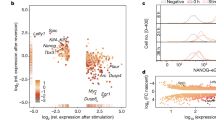
Substantial scientific interest has been dedicated recently to the crucial factors that control the pluripotent state of stem cells. To gain a comprehensive understanding of the molecular mechanisms regulating mouse embryonic stem cell (mESC) self-renewal and lineage differentiation, we have developed a robust method for studying the role of a particular gene in these processes. This protocol describes detailed procedures for the design and generation of the complementation rescue system and its application in dissecting the network of pluripotency-associated factors, using mESCs as a model. Specifically, three main procedures are described: (i) screening pluripotency-associated factors by competition assay; (ii) setting up an inducible complementation rescue system; and (iii) dynamically studying the pluripotency network response to target depletion. Completion of the competition assay and complementation rescue system takes 35 and 30 d, respectively, and an additional 16 d to study the dynamic molecular effects of a gene of interest in the pluripotency network.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Zhang, X. et al. FOXO1 is an essential regulator of pluripotency in human embryonic stem cells. Nat. Cell. Biol. 13, 1092–1099 (2011).
Ivanova, N. et al. Dissecting self-renewal in stem cells with RNA interference. Nature 442, 533–538 (2006).
Jiang, H. et al. Role for Dpy-30 in ES cell-fate specification by regulation of H3K4 methylation within bivalent domains. Cell 144, 513–525 (2011).
Ang, Y.S. et al. Wdr5 mediates self-renewal and reprogramming via the embryonic stem cell core transcriptional network. Cell 145, 183–197 (2011).
Shen, X. et al. EZH1 mediates methylation on histone H3 lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol. Cell 32, 491–502 (2008).
Schaniel, C. et al. Smarcc1/Baf155 couples self-renewal gene repression with changes in chromatin structure in mouse embryonic stem cells. Stem Cells 27, 2979–2991 (2009).
Fazzio, T.G., Huff, J.T. & Panning, B. An RNAi screen of chromatin proteins identifies Tip60-p400 as a regulator of embryonic stem cell identity. Cell 134, 162–174 (2008).
Singhal, N. et al. Chromatin-remodeling components of the BAF complex facilitate reprogramming. Cell 141, 943–955 (2010).
Sato, N., Meijer, L., Skaltsounis, L., Greengard, P. & Brivanlou, A.H. Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3-specific inhibitor. Nat. Med. 10, 55–63 (2004).
Smith, A.G. et al. Inhibition of pluripotential embryonic stem cell differentiation by purified polypeptides. Nature 336, 688–690 (1988).
Williams, R.L. et al. Myeloid leukaemia inhibitory factor maintains the developmental potential of embryonic stem cells. Nature 336, 684–687 (1988).
Ying, Q.L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Marson, A. et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134, 521–533 (2008).
Loh, Y.H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nat. Genet. 38, 431–440 (2006).
Kim, J., Chu, J., Shen, X., Wang, J. & Orkin, S.H. An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132, 1049–1061 (2008).
Wang, J. et al. A protein interaction network for pluripotency of embryonic stem cells. Nature 444, 364–368 (2006).
Mathur, D. et al. Analysis of the mouse embryonic stem cell regulatory networks obtained by ChIP-chip and ChIP-PET. Genome Biol. 9, R126 (2008).
Pardo, M. et al. An expanded Oct4 interaction network: implications for stem cell biology, development, and disease. Cell Stem Cell 6, 382–395 (2010).
van den Berg, D.L. et al. An Oct4-centered protein interaction network in embryonic stem cells. Cell Stem Cell 6, 369–381 (2010).
Hu, G. et al. A genome-wide RNAi screen identifies a new transcriptional module required for self-renewal. Genes Dev. 23, 837–848 (2009).
Ho, L. et al. An embryonic stem cell chromatin remodeling complex, esBAF, is essential for embryonic stem cell self-renewal and pluripotency. Proc. Natl. Acad. Sci. USA 106, 5181–5186 (2009).
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003).
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003).
Dejosez, M. et al. Ronin is essential for embryogenesis and the pluripotency of mouse embryonic stem cells. Cell 133, 1162–1174 (2008).
Chia, N.Y. et al. A genome-wide RNAi screen reveals determinants of human embryonic stem cell identity. Nature 468, 316–320 (2010).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Meister, G. & Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343–349 (2004).
Brummelkamp, T.R., Bernards, R. & Agami, R. A system for stable expression of short interfering RNAs in mammalian cells. Science 296, 550–553 (2002).
Sharp, P.A. RNAi and double-strand RNA. Genes Dev. 13, 139–141 (1999).
Schaniel, C. et al. Delivery of short hairpin RNAs—triggers of gene silencing—into mouse embryonic stem cells. Nat. Methods 3, 397–400 (2006).
Hannon, G.J. & Rossi, J.J. Unlocking the potential of the human genome with RNA interference. Nature 431, 371–378 (2004).
McCaffrey, A.P. et al. RNA interference in adult mice. Nature 418, 38–39 (2002).
McManus, M.T., Petersen, C.P., Haines, B.B., Chen, J. & Sharp, P.A. Gene silencing using micro-RNA designed hairpins. RNA 8, 842–850 (2002).
Paddison, P.J., Caudy, A.A., Bernstein, E., Hannon, G.J. & Conklin, D.S. Short hairpin RNAs (shRNAs) induce sequence-specific silencing in mammalian cells. Genes Dev. 16, 948–958 (2002).
Paul, C.P., Good, P.D., Winer, I. & Engelke, D.R. Effective expression of small interfering RNA in human cells. Nat. Biotechnol. 20, 505–508 (2002).
Sui, G. et al. A DNA vector-based RNAi technology to suppress gene expression in mammalian cells. Proc. Natl. Acad. Sci. USA 99, 5515–5520 (2002).
Xia, H., Mao, Q., Paulson, H.L. & Davidson, B.L. siRNA-mediated gene silencing in vitro and in vivo. Nat. Biotechnol. 20, 1006–1010 (2002).
Yu, J.Y., DeRuiter, S.L. & Turner, D.L. RNA interference by expression of short-interfering RNAs and hairpin RNAs in mammalian cells. Proc. Natl. Acad. Sci. USA 99, 6047–6052 (2002).
Schaniel, C., Lee, D.F. & Lemischka, I.R. Exploration of self-renewal and pluripotency in ES cells using RNAi. Methods Enzymol. 477, 351–365 (2010).
Lu, R. et al. Systems-level dynamic analyses of fate change in murine embryonic stem cells. Nature 462, 358–362 (2009).
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006).
Boheler, K.R. Stem cell pluripotency: a cellular trait that depends on transcription factors, chromatin state and a checkpoint deficient cell cycle. J. Cell Physiol. 221, 10–17 (2009).
Savatier, P., Huang, S., Szekely, L., Wiman, K.G. & Samarut, J. Contrasting patterns of retinoblastoma protein expression in mouse embryonic stem cells and embryonic fibroblasts. Oncogene 9, 809–818 (1994).
Ting, D.T., Kyba, M. & Daley, G.Q. Inducible transgene expression in mouse stem cells. Methods Mol. Med. 105, 23–46 (2005).
Macarthur, B.D., Ma′ayan, A. & Lemischka, I.R. Systems biology of stem cell fate and cellular reprogramming. Nat. Rev. Mol. Cell Biol. 10, 672–681 (2009).
Lewis, D.L., Hagstrom, J.E., Loomis, A.G., Wolff, J.A. & Herweijer, H. Efficient delivery of siRNA for inhibition of gene expression in postnatal mice. Nat. Genet. 32, 107–108 (2002).
Tam, P.P. & Rossant, J. Mouse embryonic chimeras: tools for studying mammalian development. Development 130, 6155–6163 (2003).
Conner, D.A. Mouse embryo fibroblast (MEF) feeder cell preparation. Curr. Protoc. Mol. Biol. Chapter 23, Unit 23 22 (2001).
Lachmann, A. et al. ChEA: transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics 26, 2438–2444 (2010).
Schmittgen, T.D. & Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Kutner, R.H., Zhang, X.Y. & Reiser, J. Production, concentration and titration of pseudotyped HIV-1-based lentiviral vectors. Nat. Protoc. 4, 495–505 (2009).
Acknowledgements
We sincerely acknowledge N. Ivanova for her original development of the rescue strategy. We thank S. Ghaffari, X. Zhang and Y.S. Ang for sharing their results for use in this protocol. We gratefully acknowledge the members of the Lemischka and Moore laboratories for advice and discussions. We thank G. Daley (Children's Hospital Boston) for Ainv15 ESCs. This work was supported by the US National Institutes of Health grant no. 5R01GM078465 (to I.R.L.), and the Empire State Stem Cell Fund through the New York State Department of Health (NYSTEM) grant nos. C024176 (to I.R.L.) and C024410 (to C.S. and I.R.L.). D.-F.L. is a New York Stem Cell Foundation Stanley and Fiona Druckenmiller Fellow.
Author information
Authors and Affiliations
Contributions
D.-F.L., J.S. and A.S. designed the experiments, carried out the work and wrote the protocol. J.G. edited and modified the protocol design. C.S. helped direct and coordinate research activity, edited and modified the protocol; I.R.L. directed the study and coordinated research activity.
Corresponding authors
Rights and permissions
About this article
Cite this article
Lee, DF., Su, J., Sevilla, A. et al. Combining competition assays with genetic complementation strategies to dissect mouse embryonic stem cell self-renewal and pluripotency. Nat Protoc 7, 729–748 (2012). https://doi.org/10.1038/nprot.2012.018
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.018
This article is cited by
-
Rewired m6A epitranscriptomic networks link mutant p53 to neoplastic transformation
Nature Communications (2023)
-
Prolonged breastfeeding protects from obesity by hypothalamic action of hepatic FGF21
Nature Metabolism (2022)
-
PBX homeobox 1 enhances hair follicle mesenchymal stem cell proliferation and reprogramming through activation of the AKT/glycogen synthase kinase signaling pathway and suppression of apoptosis
Stem Cell Research & Therapy (2019)
-
Arrayed functional genetic screenings in pluripotency reprogramming and differentiation
Stem Cell Research & Therapy (2019)
-
The long non-coding RNA Snhg3 is essential for mouse embryonic stem cell self-renewal and pluripotency
Stem Cell Research & Therapy (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.