Abstract
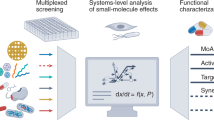
Here, we review the 'target-centric' genomic strategy to antimicrobial discovery and share our perspective on identification, validation and prioritization of potential antimicrobial drug targets in the context of emerging chemical biology, genomics and phenotypic screening strategies. We propose that coupling the dual processes of antimicrobial small-molecule screening and target identification in a whole-cell context is essential to empirically annotate 'druggable' targets and advance early stage antimicrobial discovery. We also advocate a systems-level approach to annotating synthetic-lethal genetic interactions comprehensively within yeast and bacteria models. The resulting genetic interaction networks provide a landscape to rationally predict and exploit drug synergy between cognate inhibitors. We posit that synergistic combination agents provide an important and largely unexploited strategy to 'repurpose' existing chemical space and simultaneously address issues of potency, spectrum, toxicity and drug resistance in early stages of antimicrobial drug discovery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Boucher, H.W. et al. Bad bugs, no drugs: no ESKAPE! An update from the Infectious Diseases Society of America. Clin. Infect. Dis. 48, 1–12 (2009).
Projan, S.J. Why is big Pharma getting out of antibacterial drug discovery? Curr. Opin. Microbiol. 6, 427–430 (2003).
Projan, S.J. Whither antibacterial drug discovery? Drug Discov. Today 13, 279–280 (2008).
Baltz, R.H. Marcel Faber Roundtable: is our antibiotic pipeline unproductive because of starvation, constipation or lack of inspiration? J. Ind. Microbiol. Biotechnol. 33, 507–513 (2006).
Fischbach, M.A. & Walsh, C.T. Antibiotics for emerging pathogens. Science 325, 1089–1093 (2009).
Silver, L.L. Challenges of antibacterial discovery. Clin. Microbiol. Rev. 24, 71–109 (2011).
Livermore, D.M. Discovery research: the scientific challenge of finding new antibiotics. J. Antimicrob. Chemother. 66, 1941–1944 (2011).
Payne, D.J., Gwynn, M.N., Holmes, D.J. & Pompliano, D.L. Drugs for bad bugs: confronting the challenges of antibacterial discovery. Nat. Rev. Drug Discov. 6, 29–40 (2007).
Swinney, D.C. & Anthony, J. How were new medicines discovered? Nat. Rev. Drug Discov. 10, 507–519 (2011).
Walsh, C.T. Antibiotics: Actions, Origins, Resistance (ASM Press, 2003).
Fera, M.T., La Camera, E. & De Sarro, A. New triazoles and echinocandins: mode of action, in vitro activity and mechanisms of resistance. Expert Rev. Anti Infect. Ther. 7, 981–998 (2009).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Gerdes, S.Y. et al. Experimental determination and system level analysis of essential genes in Escherichia coli MG1655. J. Bacteriol. 185, 5673–5684 (2003).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006–0008 (2006).
Akerley, B.J. et al. Systematic identification of essential genes by in vitro mariner mutagenesis. Proc. Natl. Acad. Sci. USA 95, 8927–8932 (1998).
Akerley, B.J. et al. A genome-scale analysis for identification of genes required for growth or survival of Haemophilus influenzae. Proc. Natl. Acad. Sci. USA 99, 966–971 (2002).
Jacobs, M.A. et al. Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 100, 14339–14344 (2003).
Liberati, N.T. et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc. Natl. Acad. Sci. USA 103, 2833–2838 (2006).
Langridge, G.C. et al. Simultaneous assay of every Salmonella Typhi gene using one million transposon mutants. Genome Res. 19, 2308–2316 (2009).
Forsyth, R.A. et al. A genome-wide strategy for the identification of essential genes in Staphylococcus aureus. Mol. Microbiol. 43, 1387–1400 (2002).
Chaudhuri, R.R. et al. Comprehensive identification of essential Staphylococcus aureus genes using Transposon-Mediated Differential Hybridisation (TMDH). BMC Genomics 10, 291 (2009).
Bijlsma, J.J. et al. Development of genomic array footprinting for identification of conditionally essential genes in Streptococcus pneumoniae. Appl. Environ. Microbiol. 73, 1514–1524 (2007).
Sassetti, C.M., Boyd, D.H. & Rubin, E.J. Genes required for mycobacterial growth defined by high density mutagenesis. Mol. Microbiol. 48, 77–84 (2003).
Kobayashi, K. et al. Essential Bacillus subtilis genes. Proc. Natl. Acad. Sci. USA 100, 4678–4683 (2003).
de Berardinis, V. et al. A complete collection of single-gene deletion mutants of Acinetobacter baylyi ADP1. Mol. Syst. Biol. 4, 174 (2008).
Wu, T. et al. Identification of a multicomponent complex required for outer membrane biogenesis in Escherichia coli. Cell 121, 235–245 (2005).
Thanbichler, M. & Shapiro, L. MipZ, a spatial regulator coordinating chromosome segregation with cell division in Caulobacter. Cell 126, 147–162 (2006).
Paradis-Bleau, C. et al. Lipoprotein cofactors located in the outer membrane activate bacterial cell wall polymerases. Cell 143, 1110–1120 (2010).
Roemer, T. et al. Large-scale essential gene identification in Candida albicans and applications to antifungal drug discovery. Mol. Microbiol. 50, 167–181 (2003).
Mnaimneh, S. et al. Exploration of essential gene functions via titratable promoter alleles. Cell 118, 31–44 (2004).
Park, Y.N. & Morschhauser, J. Tetracycline-inducible gene expression and gene deletion in Candida albicans. Eukaryot. Cell 4, 1328–1342 (2005).
Hu, W. et al. Essential gene identification and drug target prioritization in Aspergillus fumigatus. PLoS Pathog. 3, e24 (2007).
Becker, J.M. et al. Pathway analysis of Candida albicans survival and virulence determinants in a murine infection model. Proc. Natl. Acad. Sci. USA 107, 22044–22049 (2010).
Silhavy, T.J., Kahne, D. & Walker, S. The bacterial cell envelope. Cold Spring Harb. Perspect. Biol. 2, a000414 (2010).
Lomovskaya, O. & Bostian, K.A. Practical applications and feasibility of efflux pump inhibitors in the clinic—a vision for applied use. Biochem. Pharmacol. 71, 910–918 (2006).
Nikaido, H. & Pages, J.M. Broad-specificity efflux pumps and their role in multidrug resistance of Gram-negative bacteria. FEMS Microbiol. Rev. 36, 340–363 (2012).
Roemer, T. et al. Confronting the challenges of natural product-based antifungal discovery. Chem. Biol. 18, 148–164 (2011).
Wallace, I.M. et al. Compound prioritization methods increase rates of chemical probe discovery in model organisms. Chem. Biol. 18, 1273–1283 (2011).
Chong, C.R. & Sullivan, D.J. Jr. New uses for old drugs. Nature 448, 645–646 (2007).
Guiguemde, W.A. et al. Global phenotypic screening for antimalarials. Chem. Biol. 19, 116–129 (2012).
Ho, C.H. et al. A molecular barcoded yeast ORF library enables mode-of-action analysis of bioactive compounds. Nat. Biotechnol. 27, 369–377 (2009).
Serizawa, M. et al. Genomewide screening for novel genetic variations associated with ciprofloxacin resistance in Bacillus anthracis. Antimicrob. Agents Chemother. 54, 2787–2792 (2010).
Moffatt, J.H. et al. Colistin resistance in Acinetobacter baumannii is mediated by complete loss of lipopolysaccharide production. Antimicrob. Agents Chemother. 54, 4971–4977 (2010).
Andries, K. et al. A diarylquinoline drug active on the ATP synthase of Mycobacterium tuberculosis. Science 307, 223–227 (2005).
Stanley, S.A. et al. Identification of novel inhibitors of M. tuberculosis growth using whole cell based high-throughput screening. ACS Chem. Biol. 7, 1377–1384 (2012).
Hartkoorn, R.C. et al. Towards a new tuberculosis drug: pyridomycin—nature's isoniazid. EMBO Mol. Med. 4, 1032–1042 (2012).
Grzegorzewicz, A.E. et al. Inhibition of mycolic acid transport across the Mycobacterium tuberculosis plasma membrane. Nat. Chem. Biol. 8, 334–341 (2012).
Nam, T.G. et al. A chemical genomic analysis of decoquinate, a Plasmodium falciparum cytochrome b inhibitor. ACS Chem. Biol. 6, 1214–1222 (2011).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nat. Genet. 21, 278–283 (1999).
Giaever, G. et al. Chemogenomic profiling: identifying the functional interactions of small molecules in yeast. Proc. Natl. Acad. Sci. USA 101, 793–798 (2004).
Lum, P.Y. et al. Discovering modes of action for therapeutic compounds using a genome-wide screen of yeast heterozygotes. Cell 116, 121–137 (2004).
Parsons, A.B. et al. Exploring the mode-of-action of bioactive compounds by chemical-genetic profiling in yeast. Cell 126, 611–625 (2006).
Xu, D. et al. Genome-wide fitness test and mechanism-of-action studies of inhibitory compounds in Candida albicans. PLoS Pathog. 3, e92 (2007).
Jiang, B. et al. PAP inhibitor with in vivo efficacy identified by Candida albicans genetic profiling of natural products. Chem. Biol. 15, 363–374 (2008).
Donald, R.G. et al. A Staphylococcus aureus fitness test platform for mechanism-based profiling of antibacterial compounds. Chem. Biol. 16, 826–836 (2009).
Xu, H.H. et al. Staphylococcus aureus TargetArray: comprehensive differential essential gene expression as a mechanistic tool to profile antibacterials. Antimicrob. Agents Chemother. 54, 3659–3670 (2010).
Shoemaker, D.D., Lashkari, D.A., Morris, D., Mittmann, M. & Davis, R.W. Quantitative phenotypic analysis of yeast deletion mutants using a highly parallel molecular bar-coding strategy. Nat. Genet. 14, 450–456 (1996).
Oh, J. et al. Gene annotation and drug target discovery in Candida albicans with a tagged transposon mutant collection. PLoS Pathog. 6, e1001140 (2010).
Yan, Z. et al. Yeast Barcoders: a chemogenomic application of a universal donor-strain collection carrying bar-code identifiers. Nat. Methods 5, 719–725 (2008).
Pierce, S.E., Davis, R.W., Nislow, C. & Giaever, G. Genome-wide analysis of barcoded Saccharomyces cerevisiae gene-deletion mutants in pooled cultures. Nat. Protoc. 2, 2958–2974 (2007).
Hoon, S., St Onge, R.P., Giaever, G. & Nislow, C. Yeast chemical genomics and drug discovery: an update. Trends Pharmacol. Sci. 29, 499–504 (2008).
Smith, A.M. et al. Quantitative phenotyping via deep barcode sequencing. Genome Res. 19, 1836–1842 (2009).
Barker, C.A., Farha, M.A. & Brown, E.D. Chemical genomic approaches to study model microbes. Chem. Biol. 17, 624–632 (2010).
Hoon, S. et al. An integrated platform of genomic assays reveals small-molecule bioactivities. Nat. Chem. Biol. 4, 498–506 (2008); erratum 4, 632 (2008).
Huber, J. et al. Chemical genetic identification of peptidoglycan inhibitors potentiating carbapenem activity against methicillin-resistant Staphylococcus aureus. Chem. Biol. 16, 837–848 (2009).
Bills, G.F. et al. Discovery of the parnafungins, antifungal metabolites that inhibit mRNA polyadenylation, from the Fusarium larvarum complex and other Hypocrealean fungi. Mycologia 101, 449–472 (2009).
Mani, R., St Onge, R.P., Hartman, J.L. IV, Giaever, G. & Roth, F.P. Defining genetic interaction. Proc. Natl. Acad. Sci. USA 105, 3461–3466 (2008).
Dixon, S.J., Costanzo, M., Baryshnikova, A., Andrews, B. & Boone, C. Systematic mapping of genetic interaction networks. Annu. Rev. Genet. 43, 601–625 (2009).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Tong, A.H. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Baryshnikova, A. et al. Quantitative analysis of fitness and genetic interactions in yeast on a genome scale. Nat. Methods 7, 1017–1024 (2010).
Costanzo, M. et al. The genetic landscape of a cell. Science 327, 425–431 (2010).
Parsons, A.B. et al. Integration of chemical-genetic and genetic interaction data links bioactive compounds to cellular target pathways. Nat. Biotechnol. 22, 62–69 (2004).
Kartsonis, N.A., Nielsen, J. & Douglas, C.M. Caspofungin: the first in a new class of antifungal agents. Drug Resist. Updat. 6, 197–218 (2003).
Cyert, M.S. Calcineurin signaling in Saccharomyces cerevisiae: how yeast go crazy in response to stress. Biochem. Biophys. Res. Commun. 311, 1143–1150 (2003).
Lesage, G. et al. Analysis of β-1,3-glucan assembly in Saccharomyces cerevisiae using a synthetic interaction network and altered sensitivity to caspofungin. Genetics 167, 35–49 (2004).
Lesage, G. et al. An interactional network of genes involved in chitin synthesis in Saccharomyces cerevisiae. BMC Genet. 6, 8 (2005).
Osmond, B.C., Specht, C.A. & Robbins, P.W. Chitin synthase III: synthetic lethal mutants and 'stress related' chitin synthesis that bypasses the CSD3/CHS6 localization pathway. Proc. Natl. Acad. Sci. USA 96, 11206–11210 (1999).
Chaudhary, P.M., Tupe, S.G. & Deshpande, M.V. Chitin synthase inhibitors as antifungal agents. Mini Rev. Med. Chem. 13, 222–236 (2013).
Sandovsky-Losica, H., Shwartzman, R., Lahat, Y. & Segal, E. Antifungal activity against Candida albicans of nikkomycin Z in combination with caspofungin, voriconazole or amphotericin B. J. Antimicrob. Chemother. 62, 635–637 (2008).
Verwer, P.E., van Duijn, M.L., Tavakol, M., Bakker-Woudenberg, I.A. & van de Sande, W.W. Reshuffling of Aspergillus fumigatus cell wall components chitin and β-glucan under the influence of caspofungin or nikkomycin Z alone or in combination. Antimicrob. Agents Chemother. 56, 1595–1598 (2012).
Singh, S.D. et al. Hsp90 governs echinocandin resistance in the pathogenic yeast Candida albicans via calcineurin. PLoS Pathog. 5, e1000532 (2009).
Steinbach, W.J. et al. In vitro interactions between antifungals and immunosuppressants against Aspergillus fumigatus isolates from transplant and nontransplant patients. Antimicrob. Agents Chemother. 48, 4922–4925 (2004).
Del Poeta, M., Cruz, M.C., Cardenas, M.E., Perfect, J.R. & Heitman, J. Synergistic antifungal activities of bafilomycin A(1), fluconazole, and the pneumocandin MK-0991/caspofungin acetate (L-743,873) with calcineurin inhibitors FK506 and L-685,818 against Cryptococcus neoformans. Antimicrob. Agents Chemother. 44, 739–746 (2000).
Tamae, C. et al. Determination of antibiotic hypersensitivity among 4,000 single-gene-knockout mutants of Escherichia coli. J. Bacteriol. 190, 5981–5988 (2008).
Girgis, H.S., Hottes, A.K. & Tavazoie, S. Genetic architecture of intrinsic antibiotic susceptibility. PLoS ONE 4, e5629 (2009).
Liu, A. et al. Antibiotic sensitivity profiles determined with an Escherichia coli gene knockout collection: generating an antibiotic bar code. Antimicrob. Agents Chemother. 54, 1393–1403 (2010).
Nichols, R.J. et al. Phenotypic landscape of a bacterial cell. Cell 144, 143–156 (2011).
Flores, A.R., Parsons, L.M. & Pavelka, M.S. Jr. Characterization of novel Mycobacterium tuberculosis and Mycobacterium smegmatis mutants hypersusceptible to β-lactam antibiotics. J. Bacteriol. 187, 1892–1900 (2005).
Fajardo, A. et al. The neglected intrinsic resistome of bacterial pathogens. PLoS One 3, e1619 (2008).
Gomez, M.J. & Neyfakh, A.A. Genes involved in intrinsic antibiotic resistance of Acinetobacter baylyi. Antimicrob. Agents Chemother. 50, 3562–3567 (2006).
Lee, S.H. et al. Antagonism of chemical genetic interaction networks resensitize MRSA to β-lactam antibiotics. Chem. Biol. 18, 1379–1389 (2011).
Münch, D. et al. Identification and in vitro analysis of the GatD/MurT enzyme-complex catalyzing lipid II amidation in Staphylococcus aureus. PLoS Pathog. 8, e1002509 (2012).
Meredith, T.C., Wang, H., Beaulieu, P., Grundling, A. & Roemer, T. Harnessing the power of transposon mutagenesis for antibacterial target identification and evaluation. Mob. Genet. Elements 2, 171–178 (2012).
Therien, A.G. et al. Broadening the spectrum of β-lactam antibiotics through inhibition of signal peptidase type I. Antimicrob. Agents Chemother. 56, 4662–4670 (2012).
Tan, C.M. et al. Restoring methicillin-resistant Staphylococcus aureus susceptibility to β-lactam antibiotics. Sci. Transl. Med. 4, 126ra35 (2012).
Haydon, D.J. et al. An inhibitor of FtsZ with potent and selective anti-staphylococcal activity. Science 321, 1673–1675 (2008).
Fukumoto, A. et al. Cyslabdan, a new potentiator of imipenem activity against methicillin-resistant Staphylococcus aureus, produced by Streptomyces sp. K04–0144. II. Biological activities. J. Antibiot. (Tokyo) 61, 7–10 (2008).
Farha, M.A. et al. Inhibition of WTA synthesis blocks the cooperative action of PBPs and sensitizes MRSA to β-lactams. ACS Chem. Biol. 8, 226–233 (2013).
Borisy, A.A. et al. Systematic discovery of multicomponent therapeutics. Proc. Natl. Acad. Sci. USA 100, 7977–7982 (2003).
Kristiansen, J.E. et al. Reversal of resistance in microorganisms by help of non-antibiotics. J. Antimicrob. Chemother. 59, 1271–1279 (2007).
Ejim, L. et al. Combinations of antibiotics and nonantibiotic drugs enhance antimicrobial efficacy. Nat. Chem. Biol. 7, 348–350 (2011).
Kalan, L. & Wright, G.D. Antibiotic adjuvants: multicomponent anti-infective strategies. Expert Rev. Mol. Med. 13, e5 (2011).
Cowen, L.E. et al. Harnessing Hsp90 function as a powerful, broadly effective therapeutic strategy for fungal infectious disease. Proc. Natl. Acad. Sci. USA 106, 2818–2823 (2009).
Shapiro, R.S., Robbins, N. & Cowen, L.E. Regulatory circuitry governing fungal development, drug resistance, and disease. Microbiol. Mol. Biol. Rev. 75, 213–267 (2011).
Cokol, M. et al. Systematic exploration of synergistic drug pairs. Mol. Syst. Biol. 7, 544 (2011).
Severyn, B. et al. Parsimonious discovery of synergistic drug combinations. ACS Chem. Biol. 6, 1391–1398 (2011).
Drawz, S.M. & Bonomo, R.A. Three decades of β-lactamase inhibitors. Clin. Microbiol. Rev. 23, 160–201 (2010).
Ostrosky-Zeichner, L., Casadevall, A., Galgiani, J.N., Odds, F.C. & Rex, J.H. An insight into the antifungal pipeline: selected new molecules and beyond. Nat. Rev. Drug Discov. 9, 719–727 (2010).
Wright, G.D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 5, 175–186 (2007).
Wright, G.D. The antibiotic resistome. Expert Opin. Drug. Discov. 5, 779–788 (2010).
Feng, Y. et al. Evolution and pathogenesis of Staphylococcus aureus: lessons learned from genotyping and comparative genomics. FEMS Microbiol. Rev. 32, 23–37 (2008).
Pace, N.R., Sapp, J. & Goldenfeld, N. Phylogeny and beyond: scientific, historical, and conceptual significance of the first tree of life. Proc. Natl. Acad. Sci. USA 109, 1011–1018 (2012).
Gelperin, D.M. et al. Biochemical and genetic analysis of the yeast proteome with a movable ORF collection. Genes Dev. 19, 2816–2826 (2005).
Nakayama, H. et al. Tetracycline-regulatable system to tightly control gene expression in the pathogenic fungus Candida albicans. Infect. Immun. 68, 6712–6719 (2000).
Nakayama, H., Nakayama, N., Arisawa, M. & Aoki, Y. In vitro and in vivo effects of 14α-demethylase (ERG11) depletion in Candida glabrata. Antimicrob. Agents Chemother. 45, 3037–3045 (2001).
Nakayama, H., Izuta, M., Nakayama, N., Arisawa, M. & Aoki, Y. Depletion of the squalene synthase (ERG9) gene does not impair growth of Candida glabrata in mice. Antimicrob. Agents Chemother. 44, 2411–2418 (2000).
Balemans, W. et al. Essentiality of FASII pathway for Staphylococcus aureus. Nature 463, E3, discussion E4 (2010).
Brinster, S. et al. Type II fatty acid synthesis is not a suitable antibiotic target for Gram-positive pathogens. Nature 458, 83–86 (2009).
Sanglard, D. Resistance of human fungal pathogens to antifungal drugs. Curr. Opin. Microbiol. 5, 379–385 (2002).
Amsterdam, D. Susceptibility testing of antimicrobials in liquid media. in Antibiotics in Laboratory Medicine (ed. Lorian, V.) 89–93 (Lippincott Williams and Wilkins, Philadelphia, 2005).
Odds, F.C. Synergy, antagonism, and what the chequerboard puts between them. J. Antimicrob. Chemother. 52, 1 (2003).
Lehár, J. et al. Synergistic drug combinations tend to improve therapeutically relevant selectivity. Nat. Biotechnol. 27, 659–666 (2009).
Acknowledgements
We thank M. Costanzo, J. Piotrowski and S. Li for contributions to this manuscript. This work was supported by a grant from the Canadian Institutes of Health Research (MOP-57830) to C.B.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Terry Roemer is an employee of Merck, as stated in the affiliations, and potentially owns stock and/or holds stock in the company.
Rights and permissions
About this article
Cite this article
Roemer, T., Boone, C. Systems-level antimicrobial drug and drug synergy discovery. Nat Chem Biol 9, 222–231 (2013). https://doi.org/10.1038/nchembio.1205
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1205
This article is cited by
-
Evolutionary based drug synergy prediction using adaptive Lévy based neural network structure
Multimedia Tools and Applications (2023)
-
BIONIC: biological network integration using convolutions
Nature Methods (2022)
-
Leveraging machine learning essentiality predictions and chemogenomic interactions to identify antifungal targets
Nature Communications (2021)
-
Crystal structures of human NSDHL and development of its novel inhibitor with the potential to suppress EGFR activity
Cellular and Molecular Life Sciences (2021)
-
Drug combinations: a strategy to extend the life of antibiotics in the 21st century
Nature Reviews Microbiology (2019)