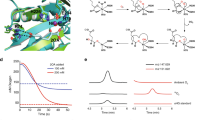
Abstract
LysW has been identified as a carrier protein in the lysine biosynthetic pathway that is active through the conversion of α-aminoadipate (AAA) to lysine. In this study, we found that the hyperthermophilic archaeon, Sulfolobus acidocaldarius, not only biosynthesizes lysine through LysW-mediated protection of AAA but also uses LysW to protect the amino group of glutamate in arginine biosynthesis. In this archaeon, after LysW modification, AAA and glutamate are converted to lysine and ornithine, respectively, by a single set of enzymes with dual functions. The crystal structure of ArgX, the enzyme responsible for modification and protection of the amino moiety of glutamate with LysW, was determined in complex with LysW. Structural comparison and enzymatic characterization using Sulfolobus LysX, Sulfolobus ArgX and Thermus LysX identify the amino acid motif responsible for substrate discrimination between AAA and glutamate. Phylogenetic analysis reveals that gene duplication events at different stages of evolution led to ArgX and LysX.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Umbarger, H.E. Amino acid biosynthesis and its regulation. Annu. Rev. Biochem. 47, 532–606 (1978).
Strassman, M. & Weinhouse, S. Biosynthetic Pathways. III. The biosynthesis of lysine by Torulopsis utilis. J. Am. Chem. Soc. 75, 1680–1684 (1953).
Vogel, H.J. Distribution of lysine pathway among fungi: evolutionary implications. Am. Nat. 98, 435–446 (1964).
Kobashi, N., Nishiyama, M. & Tanokura, M. Aspartate kinase-independent lysine synthesis in an extremely thermophilic bacterium, Thermus thermophilus: lysine is synthesized via α-aminoadipic acid, not via diaminopimeric acid. J. Bacteriol. 181, 1713–1718 (1999).
Miyazaki, J., Kobashi, N., Nishiyama, M. & Yamane, H. Functional and evolutionary relationship between arginine biosynthesis and prokaryotic lysine biosynthesis through α-aminoadipate. J. Bacteriol. 183, 5067–5073 (2001).
Nishida, H. et al. A prokaryotic gene cluster involved in synthesis of lysine through the amino adipate pathway: a key to the evolution of amino acid biosynthesis. Genome Res. 9, 1175–1183 (1999).
Brinkman, A.B., Bell, S.D., Lebbink, R.J. & de Vos, W.M. & van der Oost, J. The Sulfolobus solfataricus Lrp-like protein LysM regulates lysine biosynthesis in response to lysine availability. J. Biol. Chem. 277, 29537–29549 (2002).
Leisinger, T. & Haas, D. N-Acetylglutamate synthase of Escherichia coli regulation of synthesis and activity by arginine. J. Biol. Chem. 250, 1690–1693 (1975).
Baetens, M., Legrain, C., Boyen, A. & Glansdorff, N. Genes and enzymes of the acetyl cycle of arginine biosynthesis in the extreme thermophilic bacterium Thermus thermophilus HB27. Microbiology 144, 479–492 (1998).
Martin, P.R. & Mulks, M.H. Sequence analysis and complementation studies of the argJ gene encoding ornithine acetyltransferase from Neisseria gonorrhoeae. J. Bacteriol. 174, 2694–2701 (1992).
Horie, A. et al. Discovery of proteinaceous N-modification in lysine biosynthesis of Thermus thermophilus. Nat. Chem. Biol. 5, 673–679 (2009).
Xu, Y., Labedan, B. & Glansdorff, N. Surprising arginine biosynthesis: a reappraisal of the enzymology and evolution of the pathway in microorganisms. Microbiol. Mol. Biol. Rev. 71, 36–47 (2007).
Wagner, M. et al. Versatile genetic tool box for the Crenarchaeote Sulfolobus acidocaldarius. Front. Microbiol. 3, 214 (2012).
Hara, T., Kato, H., Katsube, Y. & Oda, J. A pseudo-Michaelis quaternary complex in the reverse reaction of a ligase: structure of Escherichia coli B glutathione synthetase complexed with ADP, glutathione, and sulfate at 2.0 Å resolution. Biochemistry 35, 11967–11974 (1996).
Sakai, H. et al. Crystal structure of a lysine biosynthesis enzyme, LysX, from Thermus thermophilus HB8. J. Mol. Biol. 332, 729–740 (2003).
Thoden, J.B., Blanchard, C.Z., Holden, H.M. & Waldrop, G.L. Movement of the biotin carboxylase B-domain as a result of ATP binding. J. Biol. Chem. 275, 16183–16190 (2000).
Jensen, R.A. Enzyme recruitment in evolution of new function. Annu. Rev. Microbiol. 30, 409–425 (1976).
Fondi, M., Brilli, M., Emiliani, G., Paffetti, D. & Fani, R. The primordial metabolism: an ancestral interconnection between leucine, arginine, and lysine biosynthesis. BMC Evol. Biol. 7 (suppl. 2), S3 (2007).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Whelan, S. & Goldman, N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol. Biol. Evol. 18, 691–699 (2001).
Jez, J.M., Ferrer, J.L., Bowman, M.E., Dixon, R.A. & Noel, J.P. Dissection of malonyl-coenzyme A decarboxylation from polyketide formation in the reaction mechanism of a plant polyketide synthase. Biochemistry 39, 890–902 (2000).
Yasukawa, T. et al. Increase of solubility of foreign proteins in Escherichia coli by coproduction of the bacterial thioredoxin. J. Biol. Chem. 270, 25328–25331 (1995).
Ellen, A.F., Albers, S.V. & Driessen, A.J. Comparative study of the extracellular proteome of Sulfolobus species reveals limited secretion. Extremophiles 14, 87–98 (2010).
Brock, T.D., Brock, K.M., Belly, R.T. & Weiss, R.L. Sulfolobus: a new genus of sulfur-oxidizing bacteria living at low pH and high temperature. Arch. Mikrobiol. 84, 54–68 (1972).
Schägger, H. Tricine-SDS-PAGE. Nat. Protoc. 1, 16–22 (2006).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. in Macromolecular Crystallography, Part A Vol. 276 (eds. Carter C. W. Jr. & R. M. Sweet) 307–326 (Academic Press, New York, 1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Edgar, R.C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5, 113 (2004).
Acknowledgements
This work was supported in part by a grant-in-aid for scientific research from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (grant nos. 21380057 and 24228001 to M.N.), Nagase Science and Technology Foundation to M.N., and from the Asahi Glass Foundation to M.N.
Author information
Authors and Affiliations
Contributions
Research planning and supervision were by T.T., T.K. and M.N.; biochemical experiments were by T.O., K.T., A.H., A.Y. and S. Kosono; phylogenetic analysis was by H.N.; gene knockout and replacement of S. acidocaldarius was by K.L. and S.-V.A.; LC-MS/MS and MALDI-TOF MS was by H.T., R. Mineki, T.F. and C.N.; StArgX was purified by T.O., A.H., A.Y., R. Masui and S. Kuramitsu; Crystallographic analysis was by T.O., A.H. and T.T.; and the manuscript was written by T.O., T.T., H.N., S.-V.A. and M.N.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results (PDF 7660 kb)
Rights and permissions
About this article
Cite this article
Ouchi, T., Tomita, T., Horie, A. et al. Lysine and arginine biosyntheses mediated by a common carrier protein in Sulfolobus. Nat Chem Biol 9, 277–283 (2013). https://doi.org/10.1038/nchembio.1200
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1200
This article is cited by
-
Large-scale computational discovery and analysis of virus-derived microbial nanocompartments
Nature Communications (2021)
-
Molecular mechanism underlying substrate recognition of the peptide macrocyclase PsnB
Nature Chemical Biology (2021)
-
Genome-scale metabolic model analysis indicates low energy production efficiency in marine ammonia-oxidizing archaea
AMB Express (2018)
-
Large-scale computational drug repositioning to find treatments for rare diseases
npj Systems Biology and Applications (2018)
-
Amino-group carrier-protein-mediated secondary metabolite biosynthesis in Streptomyces
Nature Chemical Biology (2016)