Abstract
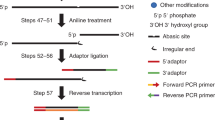
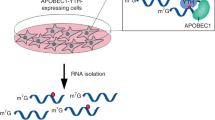
The extent and biological impact of RNA cytosine methylation are poorly understood, in part owing to limitations of current techniques for determining the targets of RNA methyltransferases. Here we describe 5-azacytidine–mediated RNA immunoprecipitation (Aza-IP), a technique that exploits the covalent bond formed between an RNA methyltransferase and the cytidine analog 5-azacytidine to recover RNA targets by immunoprecipitation. Targets are subsequently identified by high-throughput sequencing. When applied in a human cell line to the RNA methyltransferases DNMT2 and NSUN2, Aza-IP enabled >200-fold enrichment of tRNAs that are known targets of the enzymes. In addition, it revealed many tRNA and noncoding RNA targets not previously associated with NSUN2. Notably, we observed a high frequency of C→G transversions at the cytosine residues targeted by both enzymes, allowing identification of the specific methylated cytosine(s) in target RNAs. Given the mechanistic similarity of RNA cytosine methyltransferases, Aza-IP may be generally applicable for target identification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Motorin, Y., Lyko, F. & Helm, M. 5-methylcytosine in RNA: detection, enzymatic formation and biological functions. Nucleic Acids Res. 38, 1415–1430 (2010).
Squires, J.E. et al. Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res. 40, 5023–5033 (2012).
Jurkowski, T.P. et al. Human DNMT2 methylates tRNA(Asp) molecules using a DNA methyltransferase-like catalytic mechanism. RNA 14, 1663–1670 (2008).
King, M.Y. & Redman, K.L. RNA methyltransferases utilize two cysteine residues in the formation of 5-methylcytosine. Biochemistry 41, 11218–11225 (2002).
Goll, M.G. et al. Methylation of tRNAAsp by the DNA methyltransferase homolog Dnmt2. Science 311, 395–398 (2006).
Rai, K. et al. Dnmt2 functions in the cytoplasm to promote liver, brain, and retina development in zebrafish. Genes Dev. 21, 261–266 (2007).
Schaefer, M. et al. RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev. 24, 1590–1595 (2010).
Schaefer, M. & Lyko, F. Solving the Dnmt2 enigma. Chromosoma 119, 35–40 (2010).
Durdevic, Z. et al. Efficient RNA virus control in Drosophila requires the RNA methyltransferase Dnmt2. EMBO Rep. 14, 269–275 (2013).
Tuorto, F. et al. RNA cytosine methylation by Dnmt2 and NSun2 promotes tRNA stability and protein synthesis. Nat. Struct. Mol. Biol. 19, 900–905 (2012).
Brzezicha, B. et al. Identification of human tRNA:m5C methyltransferase catalysing intron-dependent m5C formation in the first position of the anticodon of the pre-tRNA Leu (CAA). Nucleic Acids Res. 34, 6034–6043 (2006).
Frye, M. & Watt, F.M. The RNA methyltransferase Misu (NSun2) mediates Myc-induced proliferation and is upregulated in tumors. Curr. Biol. 16, 971–981 (2006).
Hussain, S. et al. The nucleolar RNA methyltransferase Misu (NSun2) is required for mitotic spindle stability. J. Cell Biol. 186, 27–40 (2009).
Blanco, S. et al. The RNA-methyltransferase Misu (NSun2) poises epidermal stem cells to differentiate. PLoS Genet. 7, e1002403 (2011).
Abbasi-Moheb, L. et al. Mutations in NSUN2 cause autosomal- recessive intellectual disability. Am. J. Hum. Genet. 90, 847–855 (2012).
Khan, M.A. et al. Mutation in NSUN2, which encodes an RNA methyltransferase, causes autosomal-recessive intellectual disability. Am. J. Hum. Genet. 90, 856–863 (2012).
Martinez, F.J. et al. Whole exome sequencing identifies a splicing mutation in NSUN2 as a cause of a Dubowitz-like syndrome. J. Med. Genet. 49, 380–385 (2012).
Santi, D.V., Garrett, C.E. & Barr, P.J. On the mechanism of inhibition of DNA-cytosine methyltransferases by cytosine analogs. Cell 33, 9–10 (1983).
Lu, L.W., Chiang, G.H., Medina, D. & Randerath, K. Drug effects on nucleic acid modification. I. A specific effect of 5-azacytidine on mammalian transfer RNA methylation in vivo. Biochem. Biophys. Res. Commun. 68, 1094–1101 (1976).
Lu, L.J. & Randerath, K. Effects of 5-azacytidine on transfer RNA methyltransferases. Cancer Res. 39, 940–949 (1979).
Schaefer, M., Hagemann, S., Hanna, K. & Lyko, F. Azacytidine inhibits RNA methylation at DNMT2 target sites in human cancer cell lines. Cancer Res. 69, 8127–8132 (2009).
Liu, K., Wang, Y.F., Cantemir, C. & Muller, M.T. Endogenous assays of DNA methyltransferases: evidence for differential activities of DNMT1, DNMT2, and DNMT3 in mammalian cells in vivo. Mol. Cell Biol. 23, 2709–2719 (2003).
Schermelleh, L. et al. Trapped in action: direct visualization of DNA methyltransferase activity in living cells. Nat. Methods 2, 751–756 (2005).
Yang, X., Lay, F., Han, H. & Jones, P.A. Targeting DNA methylation for epigenetic therapy. Trends Pharmacol. Sci. 31, 536–546 (2010).
Jackson-Grusby, L., Laird, P.W., Magge, S.N., Moeller, B.J. & Jaenisch, R. Mutagenicity of 5-aza-2′-deoxycytidine is mediated by the mammalian DNA methyltransferase. Proc. Natl. Acad. Sci. USA 94, 4681–4685 (1997).
Auxilien, S., Guerineau, V., Szweykowska-Kulinska, Z. & Golinelli-Pimpaneau, B. The human tRNA m5C methyltransferase Misu is multisite-specific. RNA Biol. 9, 1331–1338 (2012).
Becker, M. et al. Pmt1, a Dnmt2 homolog in Schizosaccharomyces pombe, mediates tRNA methylation in response to nutrient signaling. Nucleic Acids Res. 40, 11648–11658 (2012).
Motorin, Y. & Grosjean, H. Multisite-specific tRNA:m5C-methyltransferase (Trm4) in yeast Saccharomyces cerevisiae: identification of the gene and substrate specificity of the enzyme. RNA 5, 1105–1118 (1999).
Hu, S., Wu, J., Chen, L. & Shan, G. Signals from noncoding RNAs: unconventional roles for conventional pol III transcripts. Int. J. Biochem. Cell Biol. 44, 1847–1851 (2012).
Gerard, M.A. et al. The scaRNA2 is produced by an independent transcription unit and its processing is directed by the encoding region. Nucleic Acids Res. 38, 370–381 (2010).
Sakita-Suto, S. et al. Aurora-B regulates RNA methyltransferase NSUN2. Mol. Biol. Cell 18, 1107–1117 (2007).
Guelorget, A. & Golinelli-Pimpaneau, B. Mechanism-based strategies for trapping and crystallizing complexes of RNA-modifying enzymes. Structure 19, 282–291 (2011).
Nix, D.A., Courdy, S.J. & Boucher, K.M. Empirical methods for controlling false positives and estimating confidence in ChIP-Seq peaks. BMC Bioinformatics 9, 523 (2008).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Todaro, G.J. & Green, H. Quantitative studies of the growth of mouse embryo cells in culture and their development into established lines. J. Cell Biol. 17, 299–313 (1963).
Dong, A. et al. Structure of human DNMT2, an enigmatic DNA methyltransferase homolog that displays denaturant-resistant binding to DNA. Nucleic Acids Res. 29, 439–448 (2001).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Koboldt, D.C. et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics 25, 2283–2285 (2009).
Laslett, D. & Canback, B. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 32, 11–16 (2004).
Chan, P.P. & Lowe, T.M. GtRNAdb: a database of transfer RNA genes detected in genomic sequence. Nucleic Acids Res. 37, D93–D97 (2009).
Acknowledgements
We thank C. Clapier (hDNMT2 protein expression and purification), K. Rai (MTase assay set-up), V. Planelles, University of Utah (gift of the lentiviral expression construct), C. Maximiliano Rêgo Monteiro Filho and S. Dehghanizadeh (lentiviral protein expression, assays and IP experiments), J. Xu and A. Yerra (help on data not shown), and X. Cheng, Emory University (gift of pQE9 plasmid). We thank B. Dalley and N. Moss (library preparation and sequencing), T. Parnell, D. Nix, B. Milash, Y. Sun and K. Boucher (help and advice on analysis) and the Center for High Performance Computing, especially W.R. Cardoen. We thank C.J. Burrows for advice on reaction mechanisms, and D.A. Jones, C.J. Burrows and D.R. Davis for many helpful comments. This work was supported by the Howard Hughes Medical Institute, the Samuel Waxman Foundation and the National Cancer Institute CA24014 (for core facilities).
Author information
Authors and Affiliations
Contributions
V.K. contributed to experimental design and approaches, performed all experiments and analyses, and helped write the paper; B.R.C. contributed to experimental design and approaches, data interpretation and wrote (with V.K.) the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Result 1, Supplementary Methods 1–4, Supplementary Figures 1–10 and Supplementary Tables 1–6 (PDF 935 kb)
Supplementary Data set 1
DNMT2 Aza-IP (tRNAs and signature analysis reports) (XLS 2182 kb)
Supplementary Data set 2
DNMT2 Aza-IP (All Genes) (XLS 13974 kb)
Supplementary Data set 3
Differentially methylated sites in wt vs Dnmt2-null MEFs RNAs (XLS 38 kb)
Supplementary Data set 4
NSUN2 Aza-IP (All Genes) (XLS 7609 kb)
Supplementary Data set 5
NSUN2 Aza-IP (VarScan signature analysis reports) (XLS 8770 kb)
Rights and permissions
About this article
Cite this article
Khoddami, V., Cairns, B. Identification of direct targets and modified bases of RNA cytosine methyltransferases. Nat Biotechnol 31, 458–464 (2013). https://doi.org/10.1038/nbt.2566
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2566
This article is cited by
-
Ultrafast bisulfite sequencing detection of 5-methylcytosine in DNA and RNA
Nature Biotechnology (2024)
-
Recent advances in the plant epitranscriptome
Genome Biology (2023)
-
Identification and analysis of RNA-5-methylcytosine-related key genes in osteoarthritis
BMC Genomics (2023)
-
m5C-methylated lncRNA NR_033928 promotes gastric cancer proliferation by stabilizing GLS mRNA to promote glutamine metabolism reprogramming
Cell Death & Disease (2023)
-
Advances in brain epitranscriptomics research and translational opportunities
Molecular Psychiatry (2023)