Abstract
The voltage-gated sodium channels are fundamental units that evoke the action potential in excitable cells such as neurons. These channels are integral membrane proteins typically consisting of one α-subunit, which forms the larger central pore of the channel, and two smaller auxiliary β-subunits, which modulate the channel functions. Genetic alterations in the SCN1A gene coding for the α-subunit of the neuronal voltage-gated sodium ion channel, type 1 (NaV 1.1), is associated with a spectrum of seizure-related disorders in human, ranging from a relatively milder form of febrile seizures to a more severe epileptic condition known as the Dravet syndrome. Among the epilepsy genes, the SCN1A gene perhaps known to have the largest number of disease-associated alleles. Here we present a meta-analysis on the SCN1A gene variants and provide comprehensive information on epilepsy-associated gene variants, their frequency, the predicted effect on the protein, the ethnicity of the affected along with the inheritance pattern and the associated epileptic phenotype. We also summarize our current understanding on the pathophysiology of the SCN1A gene defects, disease mechanism, genetic modifiers and their clinical and diagnostic relevance.
Similar content being viewed by others
Introduction
Among the genes known to be involved in epilepsy, the SCN1A gene represents one of the most commonly mutated human epilepsy genes, referred to be as super culprit gene,1 Approximately 900 distinct mutations reported in close to as many families. The SCN1A gene codes for the α-subunit of the neuronal voltage-gated sodium ion channel, type1 (NaV 1.1),2 and is expressed in the central and peripheral nervous systems and in cardiac myocytes.1, 3, 4 Mutations in the SCN1A gene were first reported in two families with generalized epilepsy with febrile seizures plus (GEFS+),5 although subsequent studies have shown that a majority (>80%) of the mutations are associated with the Dravet syndrome (DS) or its variants called severe myoclonic epilepsy, borderline (SMEB)6, 7 and intractable childhood epilepsy with generalized tonic-clonic seizures (ICEGTC).8 Both of these latter cases represent a phenotype lacking one or more features of the classical DS.9 Therefore, the SCN1A gene mutations are thought to represent a genetic form of early-onset epileptic encephalopathies10 with a common designation as DS11 (MIM#607208). Besides DS and its variants, SCN1A mutations are also associated with other rarer forms of epileptic encephalopathies as listed in the Table 1. As DS spectrum disorders account for the majority of the reported cases with the SCN1A gene mutations, we restricted our analyses on DS and tried to offer as much information as possible to the readers on DS.
DS, also known as severe myoclonic epilepsy in infancy (SMEI) (MIM# 607208), is an intractable childhood-onset epileptic encephalopathy, first reported by Charlotte Dravet, a French epileptologist.12 DS is characterized by the prolonged febrile hemiclonic or generalized tonic, clonic and tonic-clonic seizures accompanied by fever during the first year of life.13, 14, 15, 16 DS patients also show a delay in psychomotor development, dysphasia, ataxia and other neuropsychological deficits from the second year of life.17 The other seizure types seen in DS patients include myoclonus, absence and partial seizures,18 and almost all seizure types are remarkably resistant to anti-epileptic medications.19 The incidence rate of DS is nearly one in around 30 000 live births20, 21 and more than 85% of the DS patients survive up to adulthood,22, 23, 24 although early death in infancy is also known in several cases.11, 25 Curiously, males are more often affected with DS than females, with the ratio of 2:1, however, a reason for this skewed ratio is unknown.20, 21
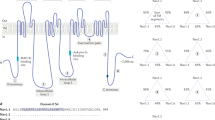
DS is caused by de novo heterozygous mutations in the SCN1A gene in about 95% of the patients26, 27 (see Supplementary Table 1). The SCN1A gene is located on the chromosome 2q24, composed of 26 exons spanning 6030 bp and encodes the large α-subunit of the voltage-gated sodium ion channel, type 1 (NaV 1.1)28 (see Figure 1). Additional alternate exons harboring the 5′-untranslated region of the gene have been reported, but the exact combination of these with the coding exons is yet to be resolved.29 Alternate splicing of exon 5 of the SCN1A gene is known to give rise to two alternate exons (exon 5N and 5A, expressed in neonates and in adults, respectively), and it has been suggested that duplication events could have lead to the evolution of such mutually exclusive exons in independent lineages.30 As expected of its function, in rodents, the expression of the Scn1a gene becomes noticeable post partum and it increases till the adulthood, predominantly in the caudal region and the spinal cord.31, 32 Within the CNS, Scn1a shows higher expression in dendrites, cell bodies31, 33 and also at the parvalbumin positive inter-neuronal axon initial segments.32, 34 The SCN1A-coded α-subunit is a 260 kDa protein that harbors four homologous domains, each spanning six transmembrane segments.35 The NaV 1.1 channel is heteromeric complex consisting of the α-subunit, which forms the larger central pore of the channel, and two smaller auxiliary β-subunits.36 The α-subunit regulates the sodium ion selectivity and can on its own function as a channel,35, 37, 38 whereas β-subunits regulate the gating and kinetics of the channel and also help in the interaction with the cytoskeleton and other proteins for the trafficking of the channel protein to the cell membrane.2, 35, 39, 40
Schematic diagram of the SCN1A gene showing coding exons in gray boxes and introns as black lines (not in scale). The position (arrows) and frequency of various SCN1A mutations (present in two or more patients, shown as bar diagram) are shown. Larger deletions and duplications are listed on the top left. A complete list of mutations and their source are given in the Supplementary Table 1. The mutation nomenclature used in the schematic is based on GenBank protein sequence reference number NP_001159435.1, and as suggested by the Human Genome Variation Society.106
Approximately 700 distinct sequence variations in the SCN1A gene are known to be associated with the DS (Figure 1; Supplementary Table 1) and all of them were found in heterozygous state, suggesting that their homozygous state could be embryonic lethal. These variations were found all along the coding region of the gene and also include large deletions and duplications (Supplementary Figure 2; Supplementary Table 1. Therefore, it can be proposed that these disease-associated variations are the loss-of-function mutations and the DS results from haploinsufficiency.38, 41, 42, 43, 44 A support for this notion comes from animal models wherein both homozygous and heterozygous Scn1a knockout mice developed epileptic seizures soon after birth.45 More importantly, a knock-in mouse model having a common DS-associated SCN1A mutation developed seizure phenotype both in homozygous and heterozygous states.32 Thus, a significant reduction in the sodium channel activity is likely to underlie the seizures phenotype42 (also see below). These animal models have shown that sodium channels are essential for the survival, as Scn1a null mutants exhibited premature death.46 Functional studies, especially the electrophysiological approaches, have suggested that the possible ‘gain-of-function’ effect for some of the disease-associated SCN1A variants and offered an alternative hypothesis on the etiology of the SCN1A-associated diseases.47, 48, 49, 50, 51, 52, 53, 54, 55, 56
The spectrum of disease-associated SCN1A gene variants
Among the epilepsy genes, the SCN1A gene perhaps is known to have the largest number of disease-associated variants so far, and therefore the SCN1A gene is also rightly named as ‘super culprit gene’.1 Two databases on the SCN1A gene mutations are reported in the literature; the SCN1A Infobase1 (http://www.scn1a.info/) and the SCN1A variant database57 (http://www.molgen.ua.ac.be/SCN1AMutations/). However, both of them have not been updated for the last couple of years, and these two databases do not provide important information such as the number of cases reported for each gene variant, the population of origin and the tested/predicted effect of the reported missense mutations. We therefore felt the need to create an up-to-date list of all the SCN1A gene variants reported to be linked with the DS-related phenotype. For this, we carried out a thorough review of the scientific literatures published in the English journals that reported a causal relationship between SCN1A gene and the epilepsy phenotype. For this, we have searched the PubMed and Medline using the query words SCN1A and SMEI, DS, SMEB or GEFS+. The search included all the papers that are listed in the PubMed until January 2013. Not all reports mention the ethnicity of the patients with the SCN1A gene defects; we therefore requested to the authors of such papers to provide us the ethnicity details of the patients in their report, and we included the details in the mutation list if such information was made available (names of authors who have kindly responded to our requests are listed in the acknowledgment section). Thus, the comprehensive list of mutations created, given in the Supplementary Table 1, was manually curated for the mutation and its frequency, its predicted effect on the protein, the ethnicity of the affected, the inheritance status of the mutation and the associated epileptic phenotype. We followed the guidelines of Human Genome Organisation (www.hgvs.org/mutnomen) for the mutation nomenclature; nucleotide 1 is the A of the ATG-translation initiation codon of the RefSeqGene (NG_011906.1) for the SCN1A gene. This RefSeqGene lists 26 exons, and therefore the exon designation given in the Supplementary Table 1 is as per this sequence. The four 5′-end untranslated exons of the SCN1A gene, reported by Long and his colleagues,29, 58 are identified in the Supplementary Table 1 with the designation as reported in the original publication (h1u, h1a, h1b and h1c). All mutations were doubly checked for accuracy. As the human SCN1A gene is known to code for at least three distinct transcripts,59 there is no uniformity in the literature with regard to which transcript should be used as the reference set for numbering the SCN1A gene mutations. Therefore, the reported mutations cannot be directly compared. In this report, we reanalyzed the mutations, and if the original report used the sequence of a shorter transcript of the SCN1A gene for numbering then the same has been corrected on the basis of the RefSeqGene (NG_011906.1;which represent the longest transcript) and renumbered accordingly in our data set (see Supplementary Table 1). For the missense mutations, we have also looked at the phylogenetic conservation of the affected residue and listed the same (see Supplementary Table 2). For this, the individual protein sequence of the α-subunit of the NaV 1.1 channel of Homo sapiens (NP_001159436.1), Mus musculus (NP_061203.2), Rattus norvegicus (AAA79965.1), Bos taurus (NP_001180147.1), Canis lupus familiaris (XP_535941.2) and Equus caballus (XP_001916728.2) were downloaded from the NCBI and aligned using the ClustalW program (see Supplementary Figure 4). Thus, this mutation update perhaps represents one of the most comprehensive analyses on the SCN1A gene variants associated with the epilepsy.
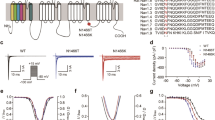
A total of 786 SCN1A gene variants are reported to be associated with DS (682), SMEB (69) or the GEFS+ (35) (see Supplementary Table 1 and Supplementary Figure 1). Therefore, DS accounts for the majority of the mutations (86%) known in the SCN1A gene. It may be noted here that the SCN1A gene variations are exclusive to the each group of epilepsy; of the 786 variants in SCN1A, only 27 of them are known to be associated with both DS and SMEB and only six with both DS and GEFS+ (see Supplementary Figure 1). Similarly, only two variants associate with both SMEB and GEFS+ and only one variant associates with all the three phenotypes (see Supplementary Figure 1), suggesting that the majority of the SCN1A gene variants are unique to the three phenotypes discussed above. At the protein level, missense mutations account for about 45% of the known variants in the SCN1A gene (Figure 2), comparing well with the data reported in the Human Gene Mutation Database (HGMD)60 (http://www.hgmd.org). Similarly, nonsense mutations (11%), small deletions (16%), splice mutations (11%) and large deletions (4.3%) in SCN1A are also in the same frequency as that reported in the HGMD. However, INDELs appear to be less frequent in the SCN1A gene (1.6%) as compared with the 15% occurrence reported in HGMD, although its significance is not obvious to us. It should be noted here that the mutations in SCN1A are de novo in origin in the majority of the patients (88%), and only 12% of the affected inherited the mutation from one of the parents (see Supplementary Table 1). Interestingly, several studies have looked at the parent of origin for the mutant allele—both for the inherited and de novo forms—and the observed patterns were strikingly opposite. Among the familial cases, 78% of the affected (39 out of 50 cases) received the mutant allele from their mother (see Supplementary Table 1), whereas in the de novo events the allele that is mutated more often appears to be paternal in origin (∼75%, in 46 out of 61 cases, see Supplementary Table 1).61, 62, 63 Although a possible cause for the skewed distribution in the familial cases is not obvious, it has been suggested that the differences in the gametogenic process between the human male and female could possibly be the reason for the increased incidence of the paternal allele undergoing mutation in de novo cases.63 For example, the meiotic entry in all egg cells is complete before birth in case of females, whereas in males spermatogenesis is a continuous process; further, the germ cells in males can more likely be exposed to mutagenic triggers, and hence more often the paternal allele could bear the mutation and appear as a de novo event.64
The schematic diagram showing the domain organization of the α-subunit of the voltage-gated sodium ion channel coded by the SCN1A gene and the positions of the missense mutations (shown as black circles). The α-subunit consists of four homologous domains (D1–D4), each having six trans-membrane segments (numbered S1–S6). The P-loop region of the channel, located between S5-S6, is identified with an asterisk. For details of individual mutation, see Supplementary Table 2.
As shown in Figure 1, genetic alterations in the SCN1A gene associating with the epileptic phenotype are seen through the gene sequence. Although genetic variants affecting the coding region of the SCN1A gene appear to be far more frequent,possibly due to the bias in the approach that normally restricts the analysis to the alterations in the coding region,sequence variations affecting the regulatory regions of the gene have also been reported. For example, detection of genomic deletion occurring at the 5′-end of the SCN1A gene led to the characterization of two promoter elements of the gene.65 Given that a majority of the mutational events in the SCN1A gene appear to be de novo in origin, it is intriguing to note that the most frequent mutational event is the large deletion removing the entire SCN1A gene (see Figure 1). Nonsense mutations p.Arg222* (15 cases) and p.Arg712* (13 cases) are the other two common mutations (Figure 1), and both of them resulted from the C-to-T transition at CpG sites, susceptible to deamination of methylated cytosine (see Supplementary Table 1). Our data set also allowed us to test whether the reported gene variants show any correlation with the ethnicity of the patients. As shown in the Supplementary Figure 3, the two groups (Asian origin and European origin) of patients did not show any difference in the frequency of the gene variants.
A total of 399 missense mutations are known in SCN1A, out of which 395 affect a conserved residue in the predicted protein (based on our ClustalW alignment; see Supplementary Figure 4), suggesting that each of these residues is critical for the function of the encoded protein (see Supplementary Table 2). This notion is reinforced by the observation that a significant proportion of the missense mutations (29%) are clustered at P-loop region of channel, which is responsible for the ion selectivity, and in the intracellular amino- and carboxyl-termini (12%), which modulate the activation/inactivation of channel66 (Figure 2). The remaining mutations (59%) were found in the 20 trans-membrane segments of the channel. Most of these mutations are expected to severely impair the channel function.67 Moreover, functional studies have indicated that indeed this could be the case for a few of those tested. For example, some of the mutants, when tested in cellular models, showed reduced cell surface expression, suggesting that impaired trafficking of the channel complex to the plasma membrane might underlie the disease.68 Electrophysiological properties for some of these mutant channels were also tested and were found to show mixed biophysical properties; for example, whereas some mutants were found to be ‘non functional’, a few other mutants showed persistent inward sodium current, possibly indicating a gain-of-function abnormality47, 49, 59; a few, however, did not show any difference in channel activity as compared with the wild-type channel.47, 49 Thus, although the phenotypic variations seen among the patients could be due to the difference in the electrophysiological properties of the mutant channel, the disease per se may result from more than a single molecular pathway, and the aberrant channel activity could be one of them.
The SCN1A gene variants and phenotype correlations
The general consensus has been that the SCN1A gene mutations cause a spectrum of phenotypes, ranging from the severe classic form of the DS to less severe forms in the order SMEB, GEFS+ and febrile seizure—the mildest form.67 Thus the channel functions in DS patients are expected to be more severely affected than those, for example, with the GEFS+ phenotype. Consistent with this model, we find that the mutations causing truncation of the SCN1A protein are more frequent in patients with DS rather than those that show SMEB or GEFS+ phenotype (see Supplementary Figure 5), suggesting that less severe genetic alterations could possibly be associated with GEFS+. This notion was supported by the observation that the missense mutations are more frequent in the GEFS+ group and least in the DS (see Supplementary Figure 5), and that the de novo mutations are more frequent in DS and SMEB than in GEFS+, whereas inherited mutations are more frequent in GEFS+ (see Supplementary Figure 6). Even among the missense mutations, mutations affecting the voltage and/or ion-pore regions were found to be greater in DS than in SMEB or GEFS+.67 Such predictions were also proven to be true for limited set of mutations using bioinformatics analysis69 or electrophysiological assays.70 However, no positive correlation has been found in between a specific mutation and a specific phenotype.67, 71 Also, the findings that the same gene alteration could result in either the DS or SMEB, and the same mutation could show variable phenotype among family members when inherited in a family suggest the possible involvement of modifiers of the phenotype. For example, the SCN1A gene variants are not fully penetrant as patients with the same genetic alterations are known to show a broad spectrum of seizure phenotypes.72 Guerrini et al.73 and Singh et al.74 have independently shown that an independent mutation in the SCN9A gene modulates the disease severity of patients with DS and familial mutations in the SCN1A gene. In one of these two reports,73 a large deletion in the SCN1A gene appears to confer a milder phenotype in the absence of a modifying allele in the SCN9A gene coding for another form of a sodium channel, suggesting that the penetrance of SCN1A-associated DS is dependent on other genetic factors, and that SCN1A gene screening should not be avoided in patients with milder phenotype. Similarly, allelic variants of the CACNA1A gene, which encodes a subunit of a voltage-dependent calcium channel, were also shown to modulate epileptic symptoms in DS cases with SCN1A mutation.75 Although a physiological link between these modifying alleles and the SCN1A variations is yet to be established, these observations nonetheless provide an insight into the complex interactions between ion channels in the epileptic phenotype and on the possible avenues for intervention strategies. Supporting this notion, mutations in other genes have been identified in DS patients who did not carry any alterations in the SCN1A gene. Examples for such genes include SCN1B,76, 77 SCN2A,47, 71, 118 PCDH1978, 79 and GABARG2.80, 81 Obviously, detailed functional studies are required to understand the molecular basis of genetic modifiers in the DS.
Animal models with the Scn1a gene defects
Much of our understanding on the mechanism by which SCN1A mutations could result in epilepsy has come from the animal models for the DS, created by deleting the Scn1a gene in mice or by replacing the wild-type mouse gene with a disease-associated mutant version analogous to one often seen in human DS patients.32, 45, 74, 82, 83, 84, 85, 86 Mice homozygous for the Scn1a null mutation developed severe ataxia and seizures, and died on day 15 post partum.45 Mice heterozygous for the gene mutation exhibited spontaneous seizures, and the spontaneous death occurred on or after day 21 post partum. Curiously, some of these phenotypes were dependent on the genetic background, suggesting the role of genetic modifiers in the disease severity.45 A knock-in mice model, wherein the murine Scn1a gene was altered to represent a human allele harboring a nonsense mutation in the SCN1A gene, also developed epileptic seizures within a month after birth—both in heterozygous and in homozygous conditions32— suggesting that the functional loss of the SCN1A gene could result in the epileptic phenotype as seen in the DS. In both of these models, the physiological dysfunctions in the interneurons (neurons that form local circuits between other two neurons) were thought to underlie the epileptic seizures.32, 45 This notion was supported by the observation that the conditional knockout of Scn1a gene only in the inhibitory interneuron in mouse resulted in seizure and premature death,46 suggesting that interneurons have a critical role in the pathogenesis of Scn1a-associated epilepsies. As expected, the Scn1a gene is expressed in inhibitory interneurons32, 87 and the deletion of the Scn1a gene impairs sodium ion currents and evoked action potentials specifically in the GABAergic interneurons, wherein NaV 1.1 represents the primary sodium channel.46, 86, 88, 89 Similar observations were also made in a transgenic mice model, wherein a GEFS+-specific SCN1A gene mutation was expressed in Scn1a knockout mice.85 Intriguingly, the GEFS+-specific transgenic channel was able to partially rescue the survival of Scn1a knockout mice,85 suggesting that the GEFS+-specific mutation results in the partial loss of the channel function, and therefore the GEFS+ is a milder phenotype as compared with the DS. A recent report that looked at the mechanism behind the early death in the Scn1a knockout mice demonstrates that parasympathetic hyperactivity following the tonic-clonic seizures results in the lethal bradycardia.90 One of the intriguing observations reported in the animals for DS is the behavioral deficits, such as hyperactivity, stereotyped behaviors and poor social interaction.88, 91 These autism-spectrum behaviors were also seen in patients with the DS,92 although it was not evident whether the genesis of these abnormal behaviors is secondary to the seizure-induced neuronal injury or a direct cause of the Scna1 gene loss. A recent study demonstrates that a low dose of benzodiazepine clonazepam—a positive modulator of GABAA receptors—corrected the behavioral abnormalities in mice heterozygous for the Scn1a gene mutant,88 suggesting that the autism-spectrum behaviors could be due to the defects in the SCN1A-dependent GABAergic signaling cascade. Similar to the observations made in human DS patients, the mice models for DS also showed effect of genetic modifiers; the null for the Scn1a gene in the 129/SvJ strain background showed fewer incidences of spontaneous seizures and survived longer as compared with the same allele in the C57BL/6J strain background.45 Genetic interactions between the Scn1a gene and the potassium channel gene Kcnq2 and also between the Scn1a gene and the Scn8a gene have been reported to modify the seizure phenotype in DS mice models.83, 93 Thus, these animal models could be used for the discovery of novel genetic modifiers for the DS. On similar note, one of the recent studies on the DS mice model suggests that ketogenic diet might provide protective effect on refractory patients with SCN1A mutations.87 Taken together, studies on the genetically engineered animal models for DS provide a mechanistic insight into the epileptic seizures, early death and behavioral deficits and also these models might help in discovering novel treatment strategies.
The SCN1A Gene Variations: Clinical and Diagnostic Relevance
The NCBI GeneTests website (http://www.ncbi.nlm.nih.gov/sites/GeneTests/) provides a list of clinical laboratories, along with the contact details, that carryout the genetic screening for the DS. Considering the allelic heterogeneity in the SCN1A gene variations, sequencing the entire gene sequence (including the regulatory regions) seems to be the first choice for the genetic screen. This seems to be the ideal approach as there are no clear-cut mutational hotspots for the SCN1A gene and a majority of the disease-associated genetic alterations seen in the SCN1A gene that are de novo in origin, and therefore the founder effect has negligible role in the disease occurrence. The large duplication and deletion mutations in the SCN1A gene associated with DS pose another challenge in the routine diagnosis; such mutations escape detection when conventional PCR amplification and DNA sequencing are attempted, and the copy-number variation can only be detected when multiplex ligation-dependent probe amplification assay or the comparative genome hybridization arrays were used.94 Thus, absence of ‘detectable’ mutation in the conventional sequencing approach should be interpreted with caution.80 Moreover, one could look for copy-number variation in the SCN1A gene in such patients because the frequency of deletions/duplications might be more than what is being calculated as of now. Given that the clinical spectrum of pediatric epilepsy, a few reports suggest that the genetic screen for the SCN1A gene mutations may be recommended to only those infants who meet a set of clinical characteristics.7, 95 For example, patients having the onset of febrile seizure at 7 months age or earlier, and those with frequent (>5/day) and prolonged (>10 min) seizures are more likely to have mutations in SCN1A gene as compared with those with a milder clinical form.7, 96 Another report suggests that normal developed before the onset of seizure and exacerbation with hyperthermia is a good predictor of the SCN1A mutation in DS cases.96 In each of these cases, a thorough clinical diagnosis appears to be a pre-requisite to recommend a case for DNA screening. To overcome some of these limitations, the use of next generation sequencing methods have been advocated for the genetic screens for epileptic patients. Lemke et al.97 have reported a cost-effective method wherein possible mutations in 265 genes, which includes the SCN1A gene, were analyzed for each patient in a single-step process, and this method could be used to diagnose the genetic basis even in clinically unresolved cases. This approach appears to be a better option, as even patients with deletion in the SCN1A gene may show milder forms of seizure as the severity could be modulated by genetic modifiers.73 Parental mosaicisms were also reported in the DS.63, 98 Thus, mutational screening could help expectant couples to consider prenatal diagnosis. Finally, there is no consensus with regard to the usefulness of the genetic screening in treatment modalities, as the knowledge of genetic basis of epilepsies did not seem to significantly improve the treatment modalities.99 A few reports suggest that sodium channel blockers such as carbamazepine and lamotrigine should be avoided as anti-convulsant, as they may aggravate seizure in SCN1A patients.99, 100 Similarly, SCN1A genetic diagnosis was reported to show positive influence on the choice of the medication and was shown to improve seizure control especially for the younger DS patients.101 An intronic single-nucleotide polymorphism (SNP; rs3812718) in the SCN1A gene was shown to serve as ‘predictor’ for the use of drugs (phenytoin and carbamazepine) at maximum dose with minimal side effects.81, 102, 103 The SCN1A gene is known to undergo alternate splicing, and this SNP affects the consensus sequence for the 5′-splice donor site of exon 5 that undergo alternative splicing to produce distinct SCN1A splice variants.102 As a result, the relative proportion of the splice SCN1A variants in the neonate and adult brain tissues was shown to be altered in individuals that carry the SNP.103 An association for this SNP in the drug response, however, could not be replicated in all populations.104, 105 Thus, such studies should be extended to other populations and a large prospective multi-centre cohort study is exceedingly essential. Such genetic screens could ensure accurate diagnosis of DS, to help genetic counseling and patient/family management, and eventually in improving treatment modalities.
Conclusion and future prospects
The DS spectrum disorders are considered as rarer forms of genetic encephalopathies with epileptic seizures as the major symptom. Despite its rarity, a large number of studies have looked at the genetic cause of DS and relatively a large body of data is available in the public domain with regard to the prevalence of mutations in the SCN1A gene and the phenotype correlates. These advances, however, did not significantly alter the treatment modalities or its outcome. Therefore, it is important to elucidate the physiological effects of the SCN1A gene variants and to understand the disease mechanisms. The animal models for the Scn1a mutations are invaluable resources to understand the pathomechanisms and in the development of therapeutic protocols.
References
Lossin, C. A catalog of SCN1A variants. Brain Dev. 31, 114–130 (2009).
Catterall, W. A. From ionic currents to molecular mechanisms: the structure and function of voltage-gated sodium channels. Neuron 26, 13–25 (2000).
Catterall, W. A. Structure and function of voltage-gated ion channels. Trends. Neurosci. 16, 500–506 (1993).
Isom, L. L., DeJongh, K. S. & Catterall, W. A. Auxiliary subunits of voltage-gated ion channels. Neuron 12, 1183–1194 (1994).
Escayg, A., MacDonald, B. T., Meisler, M. H., Baulac, S., Huberfeld, G., An-Gourfinkel, I. et al. Mutations of SCN1A, encoding a neuronal sodium channel, in two families with GEFS+2. Nat. Genet. 24, 343–345 (2000).
Ogino, T., Ohtsuka, Y., Amano, R., Yamatogi, Y. & Ohtahara, S. An investigation on the borderland of severe myoclonic epilepsy in infancy. Jpn. J. Psychiatry Neurol. 42, 554–555 (1988).
Hattori, J., Ouchida, M., Ono, J., Miyake, S., Maniwa, S., Mimaki, N. et al. A screening test for the prediction of Dravet syndrome before one year of age. Epilepsia 49, 626–633 (2008).
Fujiwara, T., Sugawara, T., Mazaki-Miyazaki, E., Takahashi, Y., Fukushima, K., Watanabe, M. et al. Mutations of sodium channel alpha subunit type 1 (SCN1A) in intractable childhood epilepsies with frequent generalized tonic-clonic seizures. Brain 126, 531–546 (2003).
Miller, I. O. & Sotero, de, Menezes, M. A. in GeneReviews™ (eds Pagon, R. A., Bird, T. D., Dolan, C. R.) (University of Washington, Seattle: WA, USA, 1993).
Anichkov, S. V. Position of pharmacology among the biological sciences and current trends of development. Pharmazie 32, 546–554 (1977).
Carranza, Rojo, D., Hamiwka, L., McMahon, J. M., Dibbens, L. M., Arsov, T., Suls, A. et al. De novo SCN1A mutations in migrating partial seizures of infancy. Neurology 77, 380–383 (2011).
Dravet, C. Les _pilepsies graves de l’enfant. Vie. Med. 8, 543–548 (1978).
Dravet, C., Roger, J., Bureau, M. & Dalla, Bernardina, M. in Advances in Epileptology (eds Akimoto, H., Kazamatsuri, H., Seino, A.) 135–141 (Raven Press: New York, NY, USA, 1982).
Scheffer, I. E., Wallace, R., Mulley, J. C. & Berkovic, S. F. Clinical and molecular genetics of myoclonic-astatic epilepsy and severe myoclonic epilepsy in infancy (Dravet syndrome). Brain. Dev. 23, 732–735 (2001).
Ohmori, I., Ohtsuka, Y., Ouchida, M., Ogino, T., Maniwa, S., Shimizu, K. et al. Is phenotype difference in severe myoclonic epilepsy in infancy related to SCN1A mutations? Brain Dev. 25, 488–493 (2003).
Mulley, J. C., Scheffer, I. E., Petrou, S., Dibbens, L. M., Berkovic, S. F. & Harkin, L. A. SCN1A mutations and epilepsy. Hum. Mutat. 25, 535–542 (2005).
Harkin, L. A., McMahon, J. M., Iona, X., Dibbens, L., Pelekanos, J. T., Zuberi, S. M. et al. The spectrum of SCN1A-related infantile epileptic encephalopathies. Brain 130, 843–852 (2007).
Dravet, C., Bureau, M., Oguni, H., Fukuyama, Y. & Cokar, O. in Epileptic Syndromes in Infancy, Childhood and Adolescence (eds Roger, J., Bureau, M., Dravet, C., Genton, P., Tassinari, C. A. & Wolf, P.) 89–113 (John Libbey Eurotext: Paris, 2005).
Ceulemans, B. Overall management of patients with Dravet syndrome. Dev. Med. Child Neurol 53 (Suppl 2), 19–23 (2011).
Hurst, D. L. Epidemiology of severe myoclonic epilepsy of infancy. Epilepsia 31, 397–400 (1990).
Yakoub, M., Dulac, O., Jambaqué, I., Chiron, C. & Plouin, P. Early diagnosis of severe myoclonic epilepsy in infancy. Brain Dev. 14, 299–303 (1992).
Akiyama, M., Kobayashi, K., Yoshinaga, H. & Ohtsuka, Y. A long-term follow-up study of Dravet syndrome up to adulthood. Epilepsia 51, 1043–1052 (2010).
Genton, P., Velizarova, R. & Dravet, C. Dravet syndrome: the long-term outcome. Epilepsia 52 (Suppl 2), 44–49 (2011).
Catarino, C. B., Liu, J. Y., Liagkouras, I., Gibbons, V. S., Labrum, R. W., Ellis, R. et al. Dravet syndrome as epileptic encephalopathy: evidence from long-term course and neuropathology. Brain 134, 2982–3010 (2011).
Freilich, E. R., Jones, J. M., Gaillard, W. D., Conry, J. A., Tsuchida, T. N., Reyes, C. et al. Novel SCN1A mutation in a proband with malignant migrating partial seizures of infancy. Arch. Neurol. 68, 665–671 (2011).
Claes, L., Del-Favero, J., Ceulemans, B., Lagae, L., Van Broeckhoven, C. & De Jonghe, P. De novo mutations in the sodium-channel gene SCN1A cause severe myoclonic epilepsy of infancy. Am. J. Hum. Genet. 68, 1327–1332 (2001).
Meisler, M. H., O'Brien, J. E. & Sharkey, L. M. Sodium channel gene family: epilepsy mutations, gene interactions and modifier effects. J. Physiol. 588, 1841–1848 (2010).
Escayg, A., Heils, A., MacDonald, B. T., Haug, K., Sander, T. & Meisler, M. H. A novel SCN1A mutation associated with generalized epilepsy with febrile seizures plus and prevalence of variants in patients with epilepsy. Am. J. Hum. Genet. 68, 866–873 (2001).
Long, Y. S., Zhao, Q. H., Su, T., Cai, Y. L., Zeng, Y., Shi, Y. W. et al. Identification of the promoter region and the 5'-untranslated exons of the human voltage-gated sodium channel Nav1.1 gene (SCN1A) and enhancement of gene expression by the 5'-untranslated exons. J. Neurosci. Res 86, 3375–3381 (2008).
Copley, R. R. Evolutionary convergence of alternative splicing in ion channels. Trends. Genet. 20, 171–176 (2004).
Gong, B., Rhodes, K. J., Bekele-Arcuri, Z. & Trimmer, J. S. Type I and type II Na(+) channel alpha-subunit polypeptides exhibit distinct spatial and temporal patterning, and association with auxiliary subunits in rat brain. J. Comp. Neurol. 412, 342–352 (1999).
Ogiwara, I., Miyamoto, H., Morita, N., Atapour, N., Mazaki, E., Inoue, I. et al. Nav1.1 localizes to axons of parvalbumin-positive inhibitory interneurons: a circuit basis for epileptic seizures in mice carrying an Scn1a gene mutation. J. Neurosci. 27, 5903–5914 (2007).
Westenbroek, R. E., Merrick, D. K. & Catterall, W. A. Differential subcellular localization of the RI and RII Na+ channel subtypes in central neurons. Neuron 3, 695–704 (1989).
Duflocq, A., Le Bras, B., Bullier, E., Couraud, F. & Davenne, M. Nav1.1 is predominantly expressed in nodes of Ranvier and axon initial segments. Mol. Cell Neurosci. 39, 180–192 (2008).
Catterall, W. A., Kalume, F. & Oakley, J. C. NaV1.1 channels and epilepsy. J. Physiol. 11, 1849–1859 (2010).
Meisler, M. H. & Kearney, J. A. Sodium channel mutations in epilepsy and other neurological disorders. J. Clin. Invest. 115, 2010–2017 (2005).
Gordon, D., Merrick, D., Auld, V., Dunn, R., Goldin, A. L., Davidson, N. et al. Tissue-specific expression of the RI and RII sodium channel subtypes. Proc. Natl Acad. Sci. USA. 84, 8682–8686 (1987).
Escayg, A. & Goldin, A. L. Sodium channel SCN1A and epilepsy: mutations and mechanisms. Epilepsia 51, 1650–1658 (2010).
Moran, O., Conti, F. & Tammaro, P. Sodium channel heterologous expression in mammalian cells and the role of the endogenous beta1-subunits. Neurosci. Lett. 336, 175–179 (2003).
Aman, T. K., Grieco-Calub, T. M., Chen, C., Rusconi, R., Slat, E. A., Isom, L. L. et al. Regulation of persistent Na current by interactions between beta subunits of voltage-gated Na channels. J. Neurosci. 29, 2027–2042 (2009).
Suls, A., Claeys, K. G., Goossens, D., Harding, B., Van, Luijk, R., Scheers, S. et al. Microdeletions involving the SCN1A gene may be common in SCN1A-mutation-negative SMEI patients. Hum. Mutat. 27, 914–920 (2006).
Yamakawa, K. Molecular and cellular basis: insights from experimental models of Dravet syndrome. Epilepsia 52 (Suppl 2), 70–71 (2011).
Bechi, G., Scalmani, P., Schiavon, E., Rusconi, R., Franceschetti, S. & Mantegazza, M. Pure haploinsufficiency for Dravet syndrome Na(V)1.1 (SCN1A) sodium channel truncating mutations. Epilepsia 53, 87–100 (2012).
Sijben, A. E., Sithinamsuwan, P., Radhakrishnan, A., Badawy, R. A., Dibbens, L., Mazarib, A. et al. Does a SCN1A gene mutation confer earlier age of onset of febrile seizures in GEFS+? Epilepsia 50, 953–956 (2009).
Yu, F. H., Mantegazza, M., Westenbroek, R. E., Robbins, C. A., Kalume, F., Burton, K. A. et al. Reduced sodium current in GABAergic interneurons in a mouse model of severe myoclonic epilepsy in infancy. Nat. Neurosci. 9, 1142–1149 (2006).
Cheah, C. S., Yu, F. H., Westenbroek, R. E., Kalume, F. K., Oakley, J. C., Potter, G. B. et al. Specific deletion of NaV1.1 sodium channels in inhibitory interneurons causes seizures and premature death in a mouse model of Dravet syndrome. Proc. Natl Acad. Sci. USA. 109, 14646–14651 (2012).
Ogiwara, I., Ito, K., Sawaishi, Y., Osaka, H., Mazaki, E., Inoue, I. et al. De novo mutations of voltage-gated sodium channel alphaII gene SCN2A in intractable epilepsies. Neurology 13, 1046–1053 (2009).
Ohmori, I., Kahlig, K. M., Rhodes, T. H., Wang, D. W. & George, A. L. Jr Nonfunctional SCN1A is common in severe myoclonic epilepsy of infancy. Epilepsia 47, 1636–1642 (2006).
Rhodes, T. H., Vanoye, C. G., Ohmori, I., Ogiwara, I., Yamakawa, K. & George, A. L. Jr. Sodium channel dysfunction in intractable childhood epilepsy with generalized tonic-clonic seizures. J. Physiol. 569, 433–445 (2005).
Spampanato, J., Escayg, A., Meisler, M. H. & Goldin, A. L. Functional effects of two voltage-gated sodium channel mutations that cause generalized epilepsy with febrile seizures plus type 2. J. Neurosci. 21, 7481–7490 (2001).
Spampanato, J., Escayg, A., Meisler, M. H. & Goldin, A. L. Generalized epilepsy with febrile seizures plus type 2 mutation W1204R alters voltage-dependent gating of Na(v)1.1 sodium channels. Neuroscience 116, 37–48 (2003).
Spampanato, J., Kearney, J. A., de Haan, G., McEwen, D. P., Escayg, A., Aradi, I. et al. A novel epilepsy mutation in the sodium channel SCN1A identifies a cytoplasmic domain for beta subunit interaction. J. Neurosci. 24, 10022–10034 (2004).
Cossette, P., Loukas, A., Lafrenière, R. G., Rochefort, D., Harvey-Girard, E., Ragsdale, D. S. et al. Functional characterization of the D188V mutation in neuronal voltage-gated sodium channel causing generalized epilepsy with febrile seizures plus (GEFS). Epilepsy Res. 53, 107–117 (2003).
Volkers, L., Kahlig, K. M., Verbeek, N. E., Das, J. H., van, Kempen, M. J., Stroink, H. et al. Nav 1.1 dysfunction in genetic epilepsy with febrile seizures-plus or Dravet syndrome. Eur. J. Neurosci. 34, 1268–1275 (2011).
Lossin, C., Rhodes, T. H., Desai, R. R., Vanoye, C. G., Wang, D., Carniciu, S. et al. Epilepsy-associated dysfunction in the voltage-gated neuronal sodium channel SCN1A. J. Neurosci. 23, 11289–11295 (2003).
Alekov, A., Rahman, M. M., Mitrovic, N., Lehmann-Horn, F. & Lerche, H. A sodium channel mutation causing epilepsy in man exhibits subtle defects in fast inactivation and activation in vitro. J. Physiol. 529, 533–539 (2000).
Claes, L. R., Deprez, L., Suls, A., Baets, J., Smets, K., Van Dyck, T. et al. The SCN1A variant database: a novel research and diagnostic tool. Hum. Mutat. 30, 904–920 (2009).
Long, Y. S., Shi, Y. W. & Liao, W. P. Conservation-based prediction of the transcription regulatory region of the SCN1A gene. Prog. Natural Sci. 19, 1675–1681 (2009).
Lossin, C., Wang, D. W., Rhodes, T. H., Vanoye, C. G. & George, A. L. Jr. Molecular basis of an inherited epilepsy. Neuron 34, 877–884 (2003).
Stenson, P. D., Ball, E., Howells, K., Phillips, A., Mort, M. & Cooper, D. N. The Human Gene Mutation Database: 2008 update. Genome Med. 45, 124–126 (2009).
Madia, F., Striano, P., Gennaro, E., Malacarne, M., Paravidino, R., Biancheri, R. et al. Cryptic chromosome deletions involving SCN1A in severe myoclonic epilepsy of infancy. Neurology 67, 1230–1235 (2006).
Heron, S. E., Scheffer, I. E., Iona, X., Zuberi, S. M., Birch, R., McMahon, J. M. et al. De novo SCN1A mutations in Dravet syndrome and related epileptic encephalopathies are largely of paternal origin. J. Med. Genet. 47, 137–141 (2010).
Sun, H., Zhang, Y., Liu, X., Ma, X., Yang, Z., Qin, J. et al. Analysis of SCN1A mutation and parental origin in patients with Dravet syndrome. J. Hum. Genet. 55, 421–427 (2010).
Crow, T. J. A theory of the origin of cerebral asymmetry: epigenetic variation superimposed on a fixed right-shift. Laterality 15, 289–303 (2010).
Nakayama, T., Ogiwara, I., Ito, K., Kaneda, M., Mazaki, E., Osaka, H. et al. Deletions of SCN1A 5′ genomic region with promoter activity in Dravet syndrome. Hum. Mutat. 31, 820–829 (2010).
Lee, A. & Goldin, A. L. Role of the amino and carboxy termini in isoform-specific sodium channel variation. J. Physiol. 586, 3917–3926 (2008).
Zuberi, S. M., Brunklaus, A., Birch, R., Reavey, E., Duncan, J. & Forbes, G. H. Genotype-phenotype associations in SCN1A-related epilepsies. Neurology 76, 594–600 (2011).
Thompson, C. H., Porter, J. C., Kahlig, K. M., Daniels, M. A. & George, A. L. Jr Non-truncating SCN1A Mutations associated with severe myoclonic epilepsy of infancy impair cell surface expression. J. Biol. Chem. 287, 42001–42008 (2012).
Kanai, K., Yoshida, S., Hirose, S., Oguni, H., Kuwabara, S., Sawai, S. et al. Physicochemical property changes of amino acid residues that accompany missense mutations in SCN1A affect epilepsy phenotype severity. J. Med. Genet. 46, 671–679 (2009).
Sugiura, Y., Ogiwara, I., Hoshi, A., Yamakawa, K. & Ugawa, Y. Different degrees of loss of function between GEFS+ and SMEI Nav 1.1 missense mutants at the same residue induced by rescuable folding defects. Epilepsia 53, e111–e114 (2012).
Shi, X., Yasumoto, S., Kurahashi, H., Nakagawa, E., Fukasawa, T., Uchiya, S. et al. Clinical spectrum of SCN2A mutations. Brain Dev. 34, 541–545 (2012).
Osaka, H., Ogiwara, I., Mazaki, E., Okamura, N., Yamashita, S., Iai, M. et al. Patients with a sodium channel alpha 1 gene mutation show wide phenotypic variation. Epilepsy Res. 75, 46–51 (2007).
Guerrini, R., Cellini, E., Mei, D., Metitieri, T., Petrelli, C., Pucatti, D. et al. Variable epilepsy phenotypes associated with a familial intragenic deletion of the SCN1A gene. Epilepsia 51, 2474–2477 (2010).
Singh, N. A., Pappas, C., Dahle, E. J., Claes, L. R., Pruess, T. H., De Jonghe, P. et al. A role of SCN9A in human epilepsies, as a cause of febrile seizures and as a potential modifier of Dravet syndrome. PLoS Genet. 5, e1000649 (2009).
Ohmori, I., Ouchida, M., Kobayashi, K., Jitsumori, Y., Mori, A., Michiue, H. et al. CACNA1A variants may modify the epileptic phenotype of Dravet syndrome. Neurobiol. Dis. 50, 209–217 (2013).
Patino, G. A., Claes, L. R., Lopez-Santiago, L. F., Slat, E. A., Dondeti, R. S., Chen, C. et al. A functional null mutation of SCN1B in a patient with Dravet syndrome. J. Neurosci. 29, 10764–10778 (2009).
Ogiwara, I., Nakayama, T., Yamagata, T., Ohtani, H., Mazaki, E., Tsuchiya, S. et al. A homozygous mutation of voltage-gated sodium channel β(I) gene SCN1B in a patient with Dravet syndrome. Epilepsia 53, e200–e203 (2012).
Depienne, C., Trouillard, O., Saint-Martin, C., Gourfinkel-An, I., Bouteiller, D., Carpentier, W. et al. Spectrum of SCN1A gene mutations associated with Dravet syndrome: analysis of 333 patients. J. Med. Genet. 46, 183–191 (2009).
Kwong, A. K., Fung, C. W., Chan, S. Y. & Wong, V. C. Identification of SCN1A and PCDH19 mutations in Chinese children with Dravet syndrome. PLoS ONE 7, e41802 (2012).
Shi, X., Huang, M. C., Ishii, A., Yoshida, S., Okada, M., Morita, K. et al. Mutational analysis of GABRG2 in a Japanese cohort with childhood epilepsies. J. Hum. Genet. 55, 375–378 (2010).
Huang, X., Tian, M., Hernandez, C. C., Hu, N. & Macdonald, R. L. The GABRG2 nonsense mutation, Q40X, associated with Dravet syndrome activated NMD and generated a truncated subunit that was partially rescued by aminoglycoside-induced stop codon read-through. Neurobiol. Dis. 48, 115–123 (2012).
Kearney, J. A., Wiste, A. K., Stephani, U., Trudeau, M. M., Siegel, A., RamachandranNair, R. et al. Recurrent de novo mutations of SCN1A in severe myoclonic epilepsy of infancy. Pediatr. Neurol. 34, 116–120 (2006).
Martin, M. S., Tang, B., Papale, L. A., Yu, F. H., Catterall, W. A. & Escayg, A. The voltage-gated sodium channel Scn8a is a genetic modifier of severe myoclonic epilepsy of infancy. Hum. Mol. Genet. 16, 2892–2899 (2007).
Papale, L. A., Beyer, B., Jones, J. M., Sharkey, L. M., Tufik, S., Epstein, M. et al. Heterozygous mutations of the voltage-gated sodium channel SCN8A are associated with spikewave discharges and absence epilepsy in mice. Hum. Mol. Genet. 18, 1633–1641 (2009).
Tang, B., Dutt, K., Papale, L., Rusconi, R., Shankar, A., Hunter, J. et al. A BAC transgenic mouse model reveals neuron subtype-specific effects of a generalized epilepsy with febrile seizures plus (GEFS+) mutation. Neurobiol. Dis. 35, 91–102 (2009).
Martin, M. S., Dutt, K., Papale, L. A., Dubé, C. M., Dutton, S. B., de Haan, G. et al. Altered function of the SCN1A voltage-gated sodium channel leads to gamma-aminobutyric acid-ergic (GABAergic) interneuron abnormalities. J. Biol. Chem. 285, 9823–9834 (2010).
Dutton, S. B., Makinson, C. D., Papale, L. A., Shankar, A., Balakrishnan, B., Nakazawa, K. et al. Preferential inactivation of Scn1a in parvalbumin interneurons increases seizure susceptibility. Neurobiol. Dis. 49, 211–220 (2012).
Han, S., Tai, C., Westenbroek, R. E., Yu, F. H., Cheah, C. S., Potter, G. B. et al. Autistic-like behaviour in Scn1a+/− mice and rescue by enhanced GABA-mediated neurotransmission. Nature 489, 385–390 (2012).
Han, S., Yu, F. H., Schwartz, M. D., Linton, J. D., Bosma, M. M., Hurley, J. B. et al. Na(V)1.1 channels are critical for intercellular communication in the suprachiasmatic nucleus and for normal circadian rhythms. Proc. Natl Acad. Sci. USA. 109, E368–E377 (2012).
Kalume, F., Westenbroek, R. E., Cheah, C. S., Yu, F. H., Oakley, J. C., Scheuer, T. et al. Sudden unexpected death in a mouse model of Dravet syndrome. J. Clin. Invest. 123, 1798–1808 (2013).
Ito, S., Ogiwara, I., Yamada, K., Miyamoto, H., Hensch, T. K., Osawa, M. et al. Mouse with Na(v)1.1 haploinsufficiency, a model for Dravet syndrome, exhibits lowered sociability and learning impairment. Neurobiol. Dis. 49, 29–40 (2012).
Li, B. M., Liu, X. R., Yi, Y. H., Deng, Y. H., Su, T., Zou, X. et al. Autism in Dravet syndrome: prevalence, features, and relationship to the clinical characteristics of epilepsy and mental retardation. Epilepsy Behav. 21, 291–295 (2011).
Hawkins, N. A., Martin, M. S., Frankel, W. N., Kearney, J. A. & Escayg, A. Neuronal voltage-gated ion channels are genetic modifiers of generalized epilepsy with febrile seizures plus. Neurobiol. Dis. 41, 655–660 (2011).
Marini, C., Scheffer, I. E., Nabbout, R., Mei, D., Cox, K., Dibbens, L. M. et al. SCN1A duplications and deletions detected in Dravet syndrome: implications for molecular diagnosis. Epilepsia 50, 1670–1678 (2009).
Fountain-Capal, J. K., Holland, K. D., Gilbert, D. L. & Hallinan, B. E. When should clinicians order genetic testing for Dravet syndrome? Pediatr Neurol. 45, 319–323 (2011).
Petrelli, C., Passamonti, C., Cesaroni, E., Mei, D., Guerrini, R., Zamponi, N. et al. Early clinical features in Dravet syndrome patients with and without SCN1A mutations. Epilepsy Res. 99, 21–27 (2012).
Lemke, J. R., Riesch, E., Scheurenbrand, T., Schubach, M., Wilhelm, C., Steiner, I. et al. Targeted next generation sequencing as a diagnostic tool in epileptic disorders. Epilepsia 53, 1387–1398 (2012).
Depienne, C., Trouillard, O., Gourfinkel-An, I., Saint-Martin, C., Bouteiller, D., Graber, D. et al. Mechanisms for variable expressivity of inherited SCN1A mutations causing Dravet syndrome. J. Med. Genet. 47, 404–410 (2010).
Delgado-Escueta, A. V. & Bourgeois, B. F. Debate: Does genetic information in humans help us treat patients? PRO—genetic information in humans helps us treat patients. CON—genetic information does not help at all. Epilepsia 49 (Suppl 9), 13–24 (2008).
Brunklaus, A., Dorris, L., Ellis, R., Reavey, E., Lee, E., Forbes, G. et al. The clinical utility of an SCN1A genetic diagnosis in infantile-onset epilepsy. Dev. Med. Child Neurol. 55, 154–161 (2012).
Brunklaus, A., Ellis, R., Reavey, E., Forbes, G. H. & Zuberi, S. M. Prognostic, clinical and demographic features in SCN1A mutation-positive Dravet syndrome. Brain 135, 2329–2336 (2012).
Tate, S. K., Depondt, C., Sisodiya, S. M., Cavalleri, G. L., Schorge, S., Soranzo, N. et al. Genetic predictors of the maximum doses patients receive during clinical use of the anti-epileptic drugs carbamazepine and phenytoin. Proc. Natl Acad. Sci. USA 102, 5507–5512 (2005).
Heinzen, E. L., Yoon, W., Tate, S. K., Sen, A., Wood, N. W., Sisodiya, S. M. et al. Nova2 interacts with a cis-acting polymorphism to influence the proportions of drug-responsive splice variants of SCN1A. Am. J. Hum. Genet. 80, 876–883 (2007).
Manna, I., Gambardella, A., Bianchi, A., Striano, P., Tozzi, R. & Aguglia, U. A functional polymorphism in the SCN1A gene does not influence antiepileptic drug responsiveness in Italian patients with focal epilepsy. Epilepsia 52, e40–e44 (2011).
Haerian, B. S., Baum, L., Tan, H. J., Kwan, P., Raymond, A. A., Saruwatari, J. et al. SCN1A IVS5N+5 polymorphism and response to sodium valproate: a multicenter study. Pharmacogenomics 13, 1477–1485 (2012).
den, Dunnen, J. T. & Antonarakis, S. E. Mutation nomenclature extensions and suggestions to describe complex mutations: a discussion. Hum. Mut. 15, 7–12 (2000).
Fukuma, G., Oguni, H., Shirasaka, Y., Watanabe, K., Miyajima, T., Yasumoto, S. et al. Mutations of neuronal voltage-gated Na+ channel alpha 1 subunit gene SCN1A in core severe myoclonic epilepsy in infancy (SMEI) and in borderline SMEI (SMEB). Epilepsia 45, 140–148 (2004).
Scheffer, I. E. Severe infantile epilepsies: molecular genetics challenge clinical classification. Brain 126, 513–514 (2003).
Guerrini, R., Parmeggiani, L., Bonanni, P., Kaminska, A. & Dulac, O. in Epilepsy Syndromes in Infancy, Childhood and Adolescence. (eds Roger, J., Bureau, M., Dravet, C., Genton, P., Tassinari, C. A. & Wolf, P.) 115–124 (John Libbey Eurotext Ltd: Montrouge, 2005).
Ebach, K., Joos, H., Doose, H., Stephani, U., Kurlemann, G., Fiedler, B. et al. SCN1A mutation analysis in myoclonic astatic epilepsy and severe idiopathic generalized epilepsy of infancy with generalized tonic-clonic seizures. Neuropediatrics 36, 210–213 (2005).
Rasmussen, T., Olszewski, J. & Lloydsmith, D. Focal seizures due to chronic localized encephalitis. Neurology 8, 435–445 (1958).
Bien, C. G., Widman, G., Urbach, H., Sassen, R., Kuczaty, S., Wiestler, O. D. et al. The natural history of Rasmussen's encephalitis. Brain 125, 1751–1759 (2002).
Ohmori, I., Ouchida, M., Kobayashi, K., Jitsumori, Y., Inoue, T., Shimizu, K. et al. Rasmussen encephalitis associated with SCN 1 A mutation. Epilepsia 49, 521–526 (2008).
Cone, T. E. Jr. On a peculiar form of infantile convulsions (hypsarrhythmia) as described in his own infant son by Dr. W.J. West in 1841. Pediatrics 46, 603 (1970).
Wallace, R. H., Hodgson, B. L., Grinton, B. E., Gardiner, R. M., Robinson, R., Rodriguez-Casero, V. et al. Sodium channel alpha1-subunit mutations in severe myoclonic epilepsy of infancy and infantile spasms. Neurology 61, 765–769 (2003).
Blume, W. T. Pathogenesis of Lennox-Gastaut syndrome: considerations and hypotheses. Epileptic Disord. 3, 183–196 (2001).
Markand, O. N. Lennox-Gastaut syndrome (childhood epileptic encephalopathy). J. Clin. Neurophysiol. 20, 426–441 (2003).
Shi, X., Yasumoto, S., Nakagawa, E., Fukasawa, T., Uchiya, S. & Hirose, S. Missense mutation of the sodium channel gene SCN2A causes Dravet syndrome. Brain Dev. 31, 758–762 (2009).
Acknowledgements
We thank Dr Ingrid E Scheffer (University of Melbourne, Australia), Dr Christel Depienne (INSERM, France), Dr Daria Riva (IRCCS Istituto Neurologico C. Besta, Italy) and Dr Iori Ohmori (Okayama University, Japan) for providing ethnicity details of the DS patients reported in their publications. Epilepsy research in the author’s laboratory was supported by sponsored research grants from the Department of Science and Technology, Department of Biotechnology and the Department of Atomic Energy, Government of India (to SG); the Council of Scientific and Industrial Research (New Delhi) in the form of research fellowship (to RP).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Parihar, R., Ganesh, S. The SCN1A gene variants and epileptic encephalopathies. J Hum Genet 58, 573–580 (2013). https://doi.org/10.1038/jhg.2013.77
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.77
Keywords
This article is cited by
-
SCN1A Gene Mutations in Indian Children With Epilepsy: Single Center Experience
Indian Pediatrics (2023)
-
Deletion of a non-canonical regulatory sequence causes loss of Scn1a expression and epileptic phenotypes in mice
Genome Medicine (2021)
-
Re-annotation of 191 developmental and epileptic encephalopathy-associated genes unmasks de novo variants in SCN1A
npj Genomic Medicine (2019)
-
The sorting receptor Rer1 controls Purkinje cell function via voltage gated sodium channels
Scientific Reports (2017)
-
Unexpected Efficacy of a Novel Sodium Channel Modulator in Dravet Syndrome
Scientific Reports (2017)