Abstract
The glucose-inhibited division gene (gid)B, which resides in the gid operon, was thought to have a role in the modulation of genes similar to that of gidA. Recent studies have indicated that GidB is a methyltransferase enzyme that is involved in the methylation of the 16S ribosomal RNA (rRNA) in Escherichia coli. In this study, we investigated the role of GidB in susceptibility to antibiotics and the overall biology of Salmonella. A gidB isogenic mutant of Salmonella was constructed and subsequently characterized under different conditions. Our data indicated that growth and invasion characteristics of the gidB mutant were similar to those of the wild type (WT). The gidB mutant was outgrown by the WT in a competitive growth assay, indicating a compromised overall bacterial fitness. Under the stress of nalidixic acid, the gidB mutant's motility was significantly reduced. Similarly, the mutant showed a filamentous morphology and smaller colony size compared with the rod-shaped and large colonies of the WT in the presence of nalidixic acid. Most importantly, deletion of gidB conferred high-level resistance to the aminoglycoside antibiotics streptomycin and neomycin. A primer extension assay determined the methylation site for the WT to be at G527 of the 16S rRNA. A lack of methylation in the mutant indicated that GidB is required for this methylation. Taken together, these data indicate that the GidB enzyme has a significant role in the alteration of antibiotic susceptibility and the modulation of growth and morphology under stress conditions in Salmonella.
Similar content being viewed by others
Introduction
Salmonella is one of the most common food-borne pathogens in the United States.1 Salmonella is transmitted by the fecal-oral route through the consumption of contaminated food or water.2, 3 In the United States, the number of salmonellosis cases has been estimated to be 1.39 million per year, with 415 deaths, costing about $2.65 billion annually due to medical costs and loss of productivity.4
The glucose-inhibited division (gid)AB operon, which includes the gidA and gidB genes, was originally related to the glucose-mediated inhibition of cell division in Escherichia coli (E. coli).5 GidA is a highly conserved gene, first described in E. coli and was thought to have a role in chromosomal division and replication due to its location near the origin of replication.6, 7 The role of gidA as a global virulence gene regulator has been established in several bacterial pathogens including Pseudomonas syringae, Aeromonas hydrophila, Pseudomonas aeruginosa and Streptococcus pyogenes.8, 9, 10, 11 Most recently, a study from our laboratory has shown that gidA affects Salmonella virulence.12 Deletion of gidA resulted in attenuation of Salmonella in vitro and in a murine model of infection, indicating a role in modulation of virulence factors.12 Studies in E. coli indicate that GidA uses a posttranscriptional-based mechanism to modify transfer RNA.8, 9, 10, 11, 13, 14, 15, 16 On the other hand, the gidB gene encodes for a methyltransferase enzyme17, 18 and therefore, proposed to be redesignated to rsmG (ribosomal small subunit methyltransferase G).17 The gidB gene is highly conserved in both Gram-negative and Gram-positive bacteria, and binds the 30S subunit of the ribosome and methylates the 16S ribosomal (rRNA).17, 18, 19 Mutation in the gidB homolog (rsmG) in E. coli, Mycobacterium tuberculosis, Thermus thermophilus and Bacillus subtilis showed a low level of streptomycin resistance due to lack of methylation of the 16S rRNA.17, 18, 19, 20 Additionally, the gidB mutant of T. thermophilus was outgrown by the wild type (WT) in a competitive growth assay, indicating a physiological cost of the methylation deficiency.19 These data indicate that GidB affects antimicrobial susceptibility in bacteria by a mechanism that involves a posttranscriptional modification.
Antibiotics are one of the most effective methods of treatment for infectious diseases in modern medicine. However, antibiotic-resistant bacterial strains are increasingly emerging, causing a significant public health threat.21 Antimicrobial resistance in non-typhoidal Salmonella serotypes has become a serious problem worldwide.22, 23, 24, 25, 26, 27, 28 For example, Salmonella enterica serovar Typhimurium DT104 is resistant to multiple antimicrobial drugs, including ampicillin, chloramphenicol, streptomycin, sulfonamide and tetracycline.29 During the early 1990s, there was a very significant increase in the Salmonella Typhimurium multidrug-resistant isolates, making up about one-third of all human isolates in 1996.30
The aminoglycoside family of antibiotics includes several medically important drugs, such as streptomycin, neomycin and kanamycin.31 They are highly active against Gram-negative bacteria and, to some extent, Gram-positive bacteria.32 Aminoglycosides act by interfering with protein synthesis by binding to the 30S subunit of the bacterial ribosomes and inducing codon misreading, as well as inhibiting translocation.31, 33 Resistance mechanisms to aminoglycosides are mainly by alteration of ribosomal binding sites, decreased antibiotic uptake and accumulation, and production of aminoglycoside-modifying enzymes.31 Enzymatic modification is the most common type of aminoglycoside resistance. Several enzymes have been identified and classified according to the type of modification, such as acetyltransferases, adenyltransferases, and phosphotransferases.34, 35 The genes encoding for aminoglycoside-modifying enzymes are usually found on plasmids and transposons, which have an important role in their dissemination. Streptomycin binds to the 30S ribosomal subunit and interacts with the C526 and G527 residues of the 16S rRNA, resulting in inhibition of protein synthesis.36, 37 On the other hand, neomycin targets the conserved sequences in the A-site of 16S rRNA.38
In this work, we investigated the role of GidB on Salmonella phenotype and biology. Deletion of gidB conferred a high-level of resistance to aminoglycoside antibiotics in Salmonella. We further determined the precise location of the methylation target of Salmonella GidB. Additionally, analysis of the mutant indicated that deletion of gidB showed defects in morphology, motility and competitive growth characteristics of the bacteria under stress conditions, suggesting a significant role in stress response for ribosomal RNA modification in Salmonella.
Materials and methods
Bacterial strains, plasmids and cell lines
The Salmonella enterica serovar Typhimurium 14028 strain was obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA), and a spontaneous nalidixic acid-resistant strain was created in our laboratory and used as the WT.12 All Salmonella strains were grown either in lysogeny broth (LB) medium or on Salmonella–Shigella plates. Nalidixic acid (150 mg ml−1), kanamycin (50 mg ml−1), tetracycline (15 mg ml−1) or ampicillin (100 mg ml−1) was added to the media as necessary. T84 human intestinal epithelial cells were obtained from ATCC and maintained in Dulbecco's modified Eagle's medium/nutrient mixture F12 medium supplemented with 10% fetal bovine serum, and incubated at 37 °C with 5% CO2.
Construction of gidB mutant
The gidB mutant was generated using the Lambda Red system following the described methods.39 Briefly, WT Salmonella Typhimurium 14028 was transformed with the plasmid pKD46, which carries the Lambda red recombinase genes.39 Arabinose-induced WT Salmonella carrying pKD46 (WT-pKD46) was cultured and used to generate electro-competent cells. The kanamycin resistance (Km) gene was PCR amplified from the pKD439 plasmid using F4/R4 primers set (5′-TTTAACACAAGCCGCGTCGTTTTTTTTCTATTTTTTTTGATAAAAATGTGTAGGCTGGAGCTGCTTCG-3′/5′-GCCCGATAAGCGAAAGCGCATCGGGCATTTGATTATTAACAGGTACATATGAATATCCTCCTTAG-3′; primers relative locations are shown in Figure 1). These primers are homologous to the flanking sequences of the Kmr gene on pKD4. The 5′-end of the F4 primer carries an extra 47 bases homologous to the upstream sequence of Salmonella gidB gene, whereas the 5′-end of the R4 primer carries an extra 45 bases homologous to gidB downstream sequence. The PCR product was electroporated into the WT-pKD46 electro-competent cells. The transformation reaction was incubated for 5 h at 37 °C, and plated on LB agar supplemented with kanamycin. Colonies were picked and gidB deletion was confirmed by sequencing, and PCR amplification using primer sets F1/R1 (5′-CTGTGCTCGACGTTGTCACT-3′/5′-GCTCTTCGTCCAGATCATCC-3′) was used to amplify the kmr cassette, the F3/R1 (5′-GCGTGCACCAGATCACACA-3′/R1) primers were used to amplify the upstream gidB sequence along with the kmr cassette, and the F1/R3 (F1/5′-TGCTGCGTCGTAGCGCTTG-3′) was used to amplify the downstream gidB sequence along with the kmr cassette. The F2/R2 primer set (5′-AGGCATAAAAACGCCCTTTC-3′/5′-GGCTATTGTTCTCCCTGACG-3′) was used to amplify the gidB gene.
Complementation of the gidB mutant
The gidB gene insert was PCR amplified from WT chromosomal DNA using the F5/R5 primers (5′-AGAAGCGACGTTGTCATGTT-3′/5′-AAGGTATGCTGCGTCGTA-3′). The PCR product was blunt-ended using a PCR polishing kit (Stratagene, Santa Clara, CA, USA), ligated to a blunt-ended pBR322 vector (lab stock) and digested with ScaI restriction enzyme. The recombinant vector was transformed into the gidB mutant strain by electroporation. To confirm the complementation, recombinant pBR322 was isolated from the gidB-complemented strain and digested with BamHI restriction enzyme. The product was separated by gel electrophoresis to analyze bands sizes. Additionally, the recombinant plasmid was PCR amplified, using primers F5/R5 flanking the gidB upstream and downstream sequences, and analyzed by gel electrophoresis.
Morphology
To observe colony morphology, LB agar plates with and without nalidixic acid were plated with the WT, gidB mutant and gidB complement. Bacterial colony morphology was examined following overnight incubation at 37 °C.
Bacterial cell morphology for gidB, WT and complemented strains was examined by growing bacteria in LB broth medium with and without nalidixic acid. Bacterial cells were Gram-stained and examined under AxioCam HRm light microscopy and analyzed by Zeiss AxioVision Rel. 4.8.1 software (Carl Zeiss Microscopy, Thornwood, NY, USA).
Growth assays
The growth characteristic of the gidB mutant was determined using a conventional growth assay. Overnight cultures of various Salmonella strains were quantified and standardized to the same OD600. An equal number of cells were inoculated into a fresh LB broth media with and without nalidixic acid and incubated at 37 °C for 8 h. Culture samples were taken every hour to determine the OD600.
A competitive growth assay was used to determine the ability of the gidB mutant to compete with the WT under stressful growth conditions. The assay was performed following the described methods.40 In brief, overnight cultures of the WT and the gidB mutant were sub-cultured in a fresh LB broth media. At an OD600 of 0.6, both cultures were mixed together in the same culture flask in a ratio of 1:1; then, the mixture was inoculated in 100 ml of fresh LB broth media in a dilution of 1:100 and incubated at 37 °C shaking at 160 r.p.m. for 2 weeks. A sample of culture was taken every second day, serially diluted and plated on LB agar plates with the appropriate antibiotics. The CFU was determined for each strain, and a growth curve was plotted.
Antibiotic susceptibility
The gidB mutant's susceptibility to antibiotics was tested by determining the MIC using conventional tube method and an automated Sensititre Susceptibility System (Trek Diagnostic Systems, Cleveland, OH, USA). Cultures of the WT, gidB mutant and the gidB complement were streaked on LB agar plates containing streptomycin, incubated overnight at 37 °C, and growth was examined. For the conventional MIC test, streptomycin, gentamicin and chloramphenicol were serially diluted two-fold in LB medium ranging from 700 to 1.37 μg ml−1 for streptomycin, and from 20 to 0.625 μg ml−1 for gentamicin and chloramphenicol. An equal number of cells from overnight cultures of the WT, gidB mutant and the complemented strain were inoculated into each antibiotic dilution and incubated overnight at 37 °C with shaking. The next day, growth versus no growth was scored. The automated Sensititre Susceptibility test was performed at the Wisconsin Veterinary Diagnostic Laboratory to determine the MIC of a wide range of the most commonly used antibiotics in veterinary medicine (in the treatment of bovine and avian species, as well as in companion animals) for gidB mutant and the WT. Briefly, colonies of the gidB mutant and the WT were suspended in demineralized water to 0.5 McFarland standard (1 × 105 CFU ml−1). A volume of 10 μl of the standardized cells were transferred to 11 ml of Mueller Hinton Broth, and the Sensititre plates (microtiter plates containing increasing doses of lyophilized antimicrobials) were inoculated with 50 μl of the cell suspension. The plates were then incubated on the Sensititre machine at 37 °C overnight.
Motility assay
Cultures of various Salmonella strains used in this study were grown in LB broth. An equal number of bacteria (1 × 106) of each strain was stabbed into a 0.35% agar (semi-solid) plates, with or without nalidixic acid, and incubated overnight at 37 °C. The motility was determined by measuring the distance migrated from the center of inoculation to the periphery of the plate.
Primer extension
This assay was performed to detect the presence or absence of methylation at G527 in the 16S rRNA of the gidB mutant and the WT of Salmonella, and therefore, determine the antibiotic resistance mechanism. Total RNA was isolated from the gidB mutant and WT using the Bacterial RiboPure RNA extraction kit (Ambion, Austin, TX, USA). N7-methylguanosines in the RNAs were cleaved by reduction with NaBH4, followed by β-elimination with acetic acid–aniline.41, 42 A yeast transfer RNA was used as an RNA carrier and was hypermodified at N7 of guanosine by dimethyl sulfate treatment, which enhanced cleavage at the N7-methylated guanosines in the rRNA.43 The rRNAs were scanned by reverse transcription using the radiolabeled primers R6 (reverse internal primer for 16S rRNA, 5′-CCGGGGATTTCACATCCGACTTGACAGAC-3′) and R7 (reverse internal primer for 16S rRNA, 5′-TTCGAATGCAGTTCCCAGGTTGAGCC-3′). The primers were 32P-labeled by mixing 12 μl of 25 ng μl−1 of the stock primers with 5 μl of [γ- 32P]ATP (Perkin-Elmer, Waltham, MA, USA), 2 μl of 10 × polynucleotide kinase buffer and 1 μl of T4 polynucleotide kinase (NEB, New England Biolabs, Ipswich, MA, USA). The mixture was incubated for 30 min at 37 °C and the free nucleotides were removed by QIAquick Nucleotide Removal Kit (Qiagen, Valencia, CA, USA). For the primer extension reaction, 19 μl (approximately 1 μM) of the treated RNA were hybridized with 1 μl of 32P-labeled oligo in the presence of 5 μl 5 × first strand buffer for 10 min at 80 °C, followed by 1 min incubation on ice. The reverse-transcription reaction was carried out by addition of 1 μl 5 × first strand buffer, 1 μl 0.1 M dithiothreitol, 2.5 μl 5 mM dNTPs and 0.5 μl of SuperScript III reverse transcriptase (Promega, Madison, WI, USA) enzyme to the RNA/oligo mix. The reaction was incubated for 30 min at 48–50 °C. To stop the reaction, one volume of formamide stop solution (95% formamide, 20 mM EDTA pH8.0, 0.05% bromophenol blue and 0.05% xylene cyanol) was added. Samples were heated to 90 °C for 1 min, then loaded on to a 9% polyacrylamide urea gel with a sequencing ladder made with the Thermo Sequenase Cycle Sequencing Kit (USB, Cleveland, OH, USA) using the pN01301 plasmid, which carries the entire rrnB rRNA operon from E. coli.44 The gel was dried, exposed to a phosphorimager screen, scanned using the Typhoon imager and analyzed using the ImageQuant software package (GE Healthcare, Piscataway, NJ, USA).
Statistical analysis
Wherever appropriate, data were analyzed using Microsoft Office Excel and a Student's t-test. P-values≤0.05 were considered to be significant. Experiments were repeated at least three times, and data were expressed as arithmetic means with s.d.
Results
Analysis of the gidB mutant and complement
The gidB mutant was generated using the Lambda Red system and confirmed by DNA sequencing and PCR analysis. The PCR amplification using chromosomal DNA from the gidB mutant revealed expected bands for the F1/R1, F3/R1 and F1/R3 primers, indicating insertion of the Kmr cassette. The amplification gave no bands with the F2/R2 primers, indicating deletion of the gidB gene. Likewise, PCR analysis using the DNA from the WT strain confirmed the presence of gidB gene and the absence of kmr cassette.
Restriction digestion analysis of the recombinant pBR322 isolated from the gidB-complemented strain showed the expected sizes of 1509 (including gidB insert) and 3733 base fragments, indicating correct orientation of the insertion. PCR amplification of the recombinant plasmid using F5/R5 primers showed the expected size (881 bp) for the gidB gene with its upstream- and downstream-flanking sequences.
GidB mutant is defective in morphology under stress condition
Examination of colony morphology indicated no significant difference in the size of colonies for the gidB mutant, compared with the WT and complemented strains (Figure 2a) when grown on LB agar without nalidixic acid. However, the gidB mutant colonies were significantly smaller than the WT and complemented strain colonies when they were grown on a nalidixic acid-containing LB agar plate (Figure 2b). Microscopic examination of Gram-stained bacterial cells indicated no significant difference in the cellular morphology of the gidB mutant compared with the WT and the complemented strains (Figure 2c) when grown in LB broth without nalidixic acid. In contrast, bacterial cultures grown in nalidixic acid-containing media showed gidB with filamentous morphology compared with the rod shape of the WT and the complemented strains (Figure 2d). These data indicated that GidB may have an important role in response of Salmonella to stress conditions such as nalidixic acid.
Colony and cellular morphology of the WT, gidB mutant and complemented strains. In panel (a), cells were grown in LB agar without nalidixic acid, and displayed no difference in colony morphology in the Salmonella strains. Panel (b) showed the gidB mutant with tiny colonies compared with the WT and complemented strains when grown on nalidixic acid-containing medium. Panel (c) showed Gram-stains for WT, gidB mutant and complemented strains having the normal rod-shaped cells when grown in LB broth without nalidixic acid. Panel (d) indicated filamentous morphology in the gidB mutant when grown in nalidixic acid-containing media compared with the normal rod-shaped WT and complemented strains. A full color version of this figure is available at The Journal of Antibiotics journal online.
Deletion of gidB affects growth and survival under stress condition
Growth curves indicated no significant difference in the growth pattern of the gidB mutant compared with the WT and the complemented strains grown with or without nalidixic acid (Figure 3a), indicating that deletion of gidB did not affect Salmonella growth. However, the competitive growth assay showed that during the first 7 days of the assay, both gidB mutant and WT strains showed similar growth rates and equally competed for the available nutrients. However, after day 8, the mutant strain showed progressive decrease in growth compared with the WT, indicating decreased overall fitness and lower survival rate (Figure 3b).
Growth curves for gidB mutant and WT strains. No significant difference in the growth pattern was observed for the two strains when grown with or without nalidixic acid. Panel (a) showed the gidB mutant and WT strains grown in LB broth without nalidixic acid (growth curve in LB broth with nalidixic acid is not shown). Panel (b) showed the competitive growth assay for gidB mutant and WT strains. The gidB mutant was unable to compete for limited nutrient sources. A full color version of this figure is available at The Journal of Antibiotics journal online.
Deletion of gidB confers antibiotic resistance
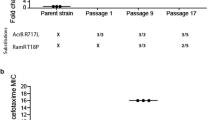
Although the gidB mutant was able to grow on LB plates containing 250 μg ml−1 of streptomycin, the WT and the complemented strains were not (data not shown). Table 1 shows the antibiotic resistance pattern of the gidB mutant compared with the WT for streptomycin, gentamicin and chloramphenicol. Although the data showed no significant difference in the gentamicin and chloramphenicol MICs between the WT and the gidB mutant, the streptomycin MIC was 700 μg ml−1 for the gidB mutant, 87.5 μg ml−1 for the WT and 175 μg ml−1 for the complemented strain. These data indicated that deletion of gidB conferred streptomycin resistance to Salmonella.
The automated Sensititre Susceptibility test showed little difference in the MIC pattern for florfenicol (WT: intermediate, gidB mutant: sensitive) and spectinomycin (WT: resistant, gidB mutant: intermediate) between the WT and the gidB mutant. However, there was a significant difference for neomycin (WT: sensitive (⩽4 μg ml−1), gidB mutant: resistant (⩾32 μg ml−1)) and streptomycin (WT: sensitive (16 μg ml−1), gidB: resistant (128 μg ml−1)) MICs in the gidB mutant compared with the WT, indicating a significant role for GidB in altering antibiotic sensitivity in Salmonella. Interestingly, streptomycin and neomycin are from the same antibiotics family, aminoglycosides, which work by binding to the small ribosomal subunit and interfering with protein synthesis.
GidB mutant is defective in bacterial motility under stress condition
Motility assay indicated that the distance migrated from the point of inoculation to the periphery was 45.3 mm for the gidB mutant and 43.33 mm for the WT when inoculated in media with no nalidixic acid (Figure 4a). However, motility plates supplemented with nalidixic acid showed that gidB mutant migrated 30.5 mm from the point of inoculation to the periphery, whereas the WT and the complemented strains migrated 63.5 mm and 51.5 mm, respectively. These data indicate that the presence of nalidixic acid in the medium significantly reduced gidB mutant motility (Figure 4b) and not the WT.
Motility assay for various Salmonella strains. Panel (a) showed the gidB mutant and WT strains grown in LB motility plates (without supplementation with nalidixic acid), with no significant difference in their migration distance (P>0.05). Panel (b) showed a significant reduction in gidB mutant motility compared with the WT and complemented strains when grown in nalidixic acid-containing LB motility plates (P<0.005).
Lack of 16S rRNA methylation in gidB mutant
Autoradiogram analysis showed that the gidB mutant and the WT termination patterns were different in the reverse transcription at nucleotide C526, which lies immediately upstream of G527 on the 16S rRNA templates (Figure 5). The band formed in the WT at C526, and not in the gidB mutant, indicated the presence of the N7-methylated guanosine in the WT, which resulted in its cleavage, and that the lack of such methylation in the gidB mutant was due to the loss of GidB activity. This data indicated that GidB is responsible for N7 methylation of G527 of the 16S rRNA in Salmonella.
Identification of GidB methylation site. Gel autoradiogram shows the primer extension products of the NaBH4 and acetic acid–aniline-treated gidB mutant and WT RNAs. Only WT RNA was cut in the phosphodiester backbone at m7G527, giving a band corresponding to C526. This band was not detected in the gidB mutant due to the lack of methylation at G527.
Discussion
To investigate the potential effects of the gidB deletion on Salmonella growth and overall fitness, we performed conventional and competitive growth assays. This assay has been used widely to measure the overall fitness and determine ability of bacterial mutants to survive and compete with the WT under stressful conditions.19, 40, 45 Consistent with what has been shown in T. thermophilus, deletion of gidB significantly decreased the overall ability of Salmonella to survive and compete with the WT under stressful growth conditions. This could be due to the physiological cost of the methylation (m7G527) deficiency in the gidB mutant,19 or alterations in biosynthetic pathways due to gidB deletion, which needs further investigation.
The Salmonella gidB mutant showed filamentous cells in the presence of nalidixic acid. Filamentation is the continued cell growth without cell division, mostly associated with metabolic changes, DNA damage and mutations in cell division machinery.46 It has been shown that filamentation is a common strategy for survival in many bacteria under stress conditions, such as nutrient limitations, host effectors and antimicrobial therapies.46, 47 For example, magnesium deficiency induces filamentation in Clostridium welchii by a mechanism involving inhibition of cell division.48, 49 In E. coli, filamentation was found to serve as a survival strategy to evade neutrophil phagocytosis.50 The β-lactam antibiotics have also been found to induce filamentation in bacteria.51, 52, 53 Burkholderia pseudomallei showed filamentous cells when exposed to ceftazidime.54 In Salmonella, a filamentous phenotype was found to be associated with the in vitro survival within the macrophages. It was suggested that nitric oxide radicals inhibited cell division and that filamentation was mediated by the DNA-damage response (SOS).55 Nalidixic acid has also been implicated in the filamentation of E. coli cells.56, 57 In the present study, nalidixic acid did not cause significant filamentation in the WT Salmonella. Therefore, it is most likely that gidB has a potential role in responses to stress conditions and induction of a filamentous morphology in Salmonella.
Alterations in bacterial colony morphology could be either in size or shape of the colony. Changes in colony shape have been related to the bacterial response to different stress stimuli, such as starvation, high temperature, presence of antibiotics in a sub-IC and genetic mutations.58, 59, 60 Several genes were shown to be involved in colony morphological change in bacteria.61, 62 In E. coli, the flu (coding for the O-antigen) and fim (coding for type-1 fimbriae) genes significantly influenced the colony morphology.62 In Salmonella, smaller size colonies have been detected in an lpd (coding for dihydrolipoamide dehydrogenase) mutant, hemL (coding for glutamate-1-semyaldehyde aminotransferase) mutant, and aroD (coding for 3-dehydroquinate dehydratase) mutant. Interestingly, these Salmonella mutants were also less susceptible to aminoglycoside antibiotics.61 Small colonies were also observed in Staphylococcus aureus when exposed to gentamicin.63, 64, 65, 66, 67, 68, 69, 70, 71, 72 Smaller colonies have shown a slower rate of growth in many bacteria, including S. aureus.73 In this study, the gidB mutant formed small colonies in the presence of nalidixic acid without showing an effect on the growth rate.
Reduced motility in the gidB mutant under the stress of nalidixic acid might be related to its filamentation. It was shown that filamentous Burkholderia pseudomallei exhibited a significant decrease in motility. This reduction in motility has been suggested to be due to suppression of energy metabolism in these filamentous cells, which is essential for flagella movement.54 Such a proposed mechanism may have also resulted in motility defect in the Salmonella gidB mutant in the presence of nalidixic acid.
In agreement with many other findings in E. coli, T. thermophilus, Bacillus subtilis and M. tuberculosis, gidB mutation conferred streptomycin resistance to Salmonella.17, 18, 19 Although it conferred a low level of streptomycin resistance in other bacteria, our mutant displayed a high level of resistance. Resistance to antimicrobial agents was independently confirmed by a commercially available automated MIC test. This test showed that the gidB mutant had, in addition to streptomycin, a high resistance to neomycin. Interestingly, streptomycin and neomycin both belong to the aminoglycoside antibiotic family. Both antibiotics act by inhibiting bacterial protein synthesis by binding to the 16S rRNA of the 30S subunit of the ribosome.33, 36, 74, 75, 76, 77 During this binding, streptomycin makes contact with the 16S rRNA residues C526 and G527, resulting in the genetic code misreading in the translation process.63, 78 It has been reported that the loss of methylation at G527 (E. coli numbering) as a result of a gidB mutation caused streptomycin resistance in B. subtilis, M. tuberculosis and other bacteria.17, 18 Our data showed a lack of methylation in the 16S rRNA of the gidB mutant, indicating a similar mechanism of resistance in Salmonella. On the other hand, neomycin binds to the 16S rRNA at a different site from that of streptomycin. The method by which GidB confers neomycin resistance in Salmonella is not clear. It is possible that lack of methylation in the gidB mutant may have affected the binding site of neomycin or by another mechanism yet to be determined.
In conclusion, we developed a Salmonella strain from which gidB was deleted from the chromosome. Subsequently, the gidB mutant was characterized using biological and virulence assays. Although GidB had no effect on the growth of Salmonella, a competitive growth assay indicated that deletion of gidB significantly affected the overall fitness of the bacteria, specifically under limited stressful growth conditions. In the presence of nalidixic acid, gidB mutant showed smaller sized colonies, filamentous cellular morphology and reduction in bacterial motility compared with the WT. Most importantly, the gidB mutant gained a high level of streptomycin resistance compared with the WT. These effects of GidB could be restored to the WT phenotype by complementation of the gidB mutant. The streptomycin resistance was associated with the loss of methylation at the G527 of the 16S rRNA of the gidB mutant, as demonstrated by the primer extension assay. The MIC data indicated that the gidB mutant was also resistant to another aminoglycoside antibiotic neomycin. Taken together, these data strongly suggest that gidB has an important role in response to stress conditions in Salmonella. It will be crucial to examine the effect of GidB on the ribosomal structure and function in Salmonella. Further investigation is needed to determine the role of GidB in stress response and in modulation of metabolic pathways. Such studies are now underway in our laboratory.
References
Centers for Disease Control and Prevention. CDC Estimates of Foodborne Illness in the United States: CDC 2011 Estimates: Findings (2011) http://www.cdc.gov/foodborneburden/2011-foodborne-estimates.html.
Centers for Disease Control and Prevention. Salmonella: General Information: Technical Information (2010) http://www.cdc.gov/salmonella/general/.
Salmonella: Salmonella Outbreak Investigations: Investigation Update: Outbreak of Salmonella Typhimurium Infections: Questions and Answers Related to the Salmonella Typhimurium Outbreak and Pets (2008–2009) http://www.cdc.gov/salmonella/typhimurium/update.html.
Buzby, J. C. & Frenzen, P. D. Estimating the Cost of Illness Caused by Select Foodborne Pathogen, United States Department of Agriculture, Economic Research Service, (2010) http://www.fsis.usda.gov/PDF/Atlanta2010/Slides_FSEC_JBuzby.pdf.
von Meyenburg, K. & Hansen, F. G. The origin of replication, oriC, of the Escherichia coli chromosome: genes near to oriC and construction of oriC deletion mutations. In Mechanistic Studies of DNA Replication and Genetic Recombination. Mol. Cell. Biol. 19, 137–159 (1980).
von Meyenburg, K., Jorgensen, B. B., Nielsen, J. & Hansen, F. G. Promoters of the atp operon coding for the membrane-bound ATP synthase of Escherichia coli mapped by Tn10 insertion mutations. Mol. Gen. Genet. 188, 240–248 (1982).
Ogawa, T. & Okazaki, T. Concurrent transcription from the gid and mioC promoters activates replication of an Escherichia coli minichromosome. Mol. Gen. Genet. 230, 193–200 (1991).
Kinscherf, T. G. & Willis, D. K. Global regulation by gidA in Pseudomonas syringae. J. Bacteriol. 184, 2281–2286 (2002).
Gupta, R., Gobble, T. R. & Schuster, M. GidA posttranscriptionally regulates rhl quorum sensing in Pseudomonas aeruginosa. J. Bacteriol. 191, 5785–5792 (2009).
Sha, J. et al. Molecular characterization of a glucose-inhibited division gene, gidA, that regulates cytotoxic enterotoxin of Aeromonas hydrophila. Infect. Immun. 72, 1084–1095 (2004).
Cho, K. H. & Caparon, M. G. tRNA modification by GidA/MnmE is necessary for Streptococcus pyogenes virulence: a new strategy to make live attenuated strains. Infect. Immun. 76, 3176–3186 (2008).
Shippy, D. C., Eakley, N. M., Bochsler, P. N. & Fadl, A. A. Biological and virulence characteristics of Salmonella enterica serovar Typhimurium following deletion of glucose-inhibited division (gidA) gene. J. Microb. Pathog. 50, 303–313 (2011).
Via, L. E. et al. Polymorphisms associated with resistance and cross-resistance to aminoglycosides and capreomycin in Mycobacterium tuberculosis isolates from South Korean Patients with drug-resistant tuberculosis. J. Clin. Microbiol. 48, 402–411 (2010).
Shi, R. et al. Structure function analysis of Escherichia coli MnmG (GidA), a highly conserved tRNA modifying enzyme. J. Bacteriol. 191, 7614 (2009).
Moukadiri, I. et al. Evolutionarily conserved proteins MnmE and GidA catalyze the formation of two methyluridine derivatives at tRNA wobble positions. Nucleic Acids Res. 37, 7177–7193 (2009).
Yim, L., Moukadiri, I., Björk, G. R. & Armengod, M. E. Further insights into the tRNA modification process controlled by proteins MnmE and GidA of Escherichia coli. Nucleic Acids Res. 34, 5892–5905 (2006).
Okamoto, S. et al. Loss of a conserved 7-methylguanosine modification in 16S rRNA confers low-level streptomycin resistance in bacteria. J. Mol. Microbiol. 63, 1096–1106 (2007).
Nishimura, K. et al. Identification of the RsmG methyltransferase target as 16S rRNA nucleotide G527 and characterization of Bacillus subtilis rsmG mutants. J. Bacteriol. 189, 6068–6073 (2007).
Gregory, S. T. et al. Structural and functional studies of the Thermus thermophilus 16S rRNA methyltransferase RsmG. J. RNA 15, 1693–1704 (2009).
Wong, S. Y. et al. Mutations in gidB Confer Low-Level Streptomycin Resistance in Mycobacterium tuberculosis. J. Antimicrob. Agents Chemother. 55, 2515–2522 (2011).
Centers for Disease Control and Prevention. Antibiotic: Antimicrobial Resistance (2011) http://www.cdc.gov/drugresistance/index.html.
Chiu, C. H. et al. The emergence in Taiwan of fluoroquinolone resistance in Salmonella enterica serotype Choleraesuis. N. Engl. J. Med. 346, 413–419 (2002).
Cohen, M. L. & Tauxe, R. V. Drug-resistanct Salmonella in the United States: an epidemiologic perspective. J. Science 234, 964–969 (1986).
Hoge, C. W., Gambel, J. M., Srijan, A., Pitarangsi, C. & Echeverria, P. Trends in antibiotic resistance among diarrheal pathogens isolated in Thailand. J. Clin. Infect. Dis. 26, 341–345 (1998).
Reina, J., Gomez, J., Serra, A. & Borell, N. Analysis of the antibiotic resistance detected in 2043 strains of Salmonella enterica subsp. Enterica isolated in stool cultures of Spanish patients with acute diarrhea (1986–1991). J. Antimicrob. Chemother. 32, 765–769 (1993).
Saxena, S. N. et al. Surveillance of Salmonellae in India for drug resistance. Indian J. Med. Sci. 43, 145–150 (1989).
Su, L. H. et al. Secular trends in incidence and antimicrobial resistance among clinical isolates of Salmonella at a university hospital in Taiwan, 1983–1999. J. Epidemiol. Infect. 127, 207–213 (2001).
Tassios, P. T. et al. Molecular epidemiology of antibiotic resistance of Salmonella enteritidis during a 7-year period in Greece. J. Clin. Microbiol. 35, 1316–1321 (1997).
Threlfall, E. J., Frost, J. A., Ward, L. R. & Rowe, B. Epidemic in cattle and humans of Salmonella Typhimurium DT104 with chromosomally integrated multiple drug resistance. J. Vet. Rec. 134, 577 (1994).
Glynn, M. K. et al. Emergence of multidrug-resistant Salmonella enterica serotype Typhimurium DT104 infections in the United States. New Engl. J. Med. 338, 1333–1338 (1998).
Mingeot-Leclercq, M. P., Glupczynski, Y. & Tulkens, P. M. Aminoglycosides: Activity and resistance. Antimicrob. Agents Chemother. 43, 727–737 (1999).
Hermann, T. Aminoglycoside antibiotics: Old drugs and new therapeutic approaches. Cell. Mol. Life Sci. 64, 1841–1852 (2007).
Fourmy, D., Recht, M. I. & Puglisi, J. D. Binding of neomycin-class aminoglycoside antibiotics to the A-site of 16 S rRNA. J. Mol. Biol. 277, 347–362 (1998).
Shaw, K. J., Rather, P. N., Hare, R. S. & Miller, G. H. Molecular genetics of aminoglycoside resistance genes and familial relationships of the aminoglycoside-modifying enzymes. Microbiol. Rev. 57, 138–163 (1993).
Davies, J. & Wright, G. D. Bacterial resistance to aminoglycoside antibiotics. Trends Microbiol. 5, 234–240 (1997).
Carter, A. P. et al. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407, 340–348 (2000).
Igarashi, K., Ishitsuka, H. & Kaji, A. Comparative studies on the mechanism of action of lincomycin, streptomycin and erythromycin. Biochem. Biophys. Res. Commun. 37, 499–504 (1969).
Woodcock, J., Moazed, D., Cannon, M., Davies, J. & Noller, H. F. Interaction of antibiotics with A- and P-site-specific bases in 16S ribosomal RNA. EMBO J. 10, 3099–3103 (1991).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Armalyte, J., Seputiene, V., Melefors, O. & Suziedeliene, E. An Escherichia coli asr mutant has decreased fitness during colonization in a mouse model. J. Res. Microbiol. 159, 486–493 (2008).
Peattie, D. A. Direct chemical method for sequencing RNA. Proc. Natl Acad. Sci. USA 76, 1760–1764 (1979).
Wintermeyer, W. & Zachau, H. G. Tertiary structure interactions of 7-methylguanosine in yeast tRNA Phe as studied by borohydride reduction. FEBS Lett. 58, 306–309 (1975).
Zueva, V. S., Mankin, A. S., Bogdanov, A. A. & Baratova, L. A. Specific fragmentation of tRNA and rRNA at a 7-methylguanine residue in the presence of methylated carrier RNA. Eur. J. Biochem. 146, 679–687 (1985).
Jinks-Robertson, S., Gourse, R. & Nomura, M. Expression of rRNA and tRNA genes in Escherichia coli: evidence for feedback regulation by products of rRNA operons. J. Cell 33, 865–876 (1983).
Farrell, M. J. & Finkel, S. E. The growth advantage in stationary phase phenotype conferred by rpoS mutations is dependent on the pH and nutrient environment. J. Bacteriol. 185, 7044–7052 (2003).
Justice, S. S., Hunstad, D. A., Cegelski, L. & Hultgren, S. J. Morphological plasticity as a bacterial survival strategy. Nat. Rev. Microbiol. 6, 162–168 (2008).
Young, K. D. The selective value of bacterial shape. Microbiol. Mol. Biol. Rev. 70, 660–703 (2006).
Webb, M. Effects of magnesium on cellular division in bacteria. J. Science 118, 607–611 (1953).
Webb, M. The influence of magnesium on cell division. I. The growth of Clostridium welchii in complex media deficient in magnesium. J. Gen. Microbiol. 2, 275–287 (1948).
Justice, S. S. et al. Differentiation and developmental pathways of uropathogenic Escherichia coli in urinary tract pathogenesis. Proc. Natl Acad. Sci. USA 101, 1333–1338 (2004).
Comber, K. R., Boon, R. J. & Sutherland, R. Comparative effects of amoxycillin and ampicillin on the morphology of Escherichia coli in vivo and correlation with activity. J. Antimicrob. Agents Chemother. 12, 736–744 (1977).
Nakao, M., Nishi, T. & Tsuchiya, K. In vitro and in vivo morphological response of Klebsiella pneumoniae to cefotiam and cefazolin. J. Antimicrob. Agents Chemother. 19, 901–910 (1981).
Ryan, D. M. & Monsey, D. Bacterial filamentation and in vivo efficacy: a comparison of several cephalosporins. J. Antimicrob. Chemother. 7, 57–63 (1981).
Chen, K., Sun, G. W., Chua, K. L. & Gan, Y. H. Modified virulence of antibiotic-induced Burkholderia pseudomallei filaments. J. Antimicrob. Agents Chemother. 49, 1002–1009 (2005).
Rosenberger, C. M. & Finlay, B. B. Macrophages inhibit Salmonella Typhimurium replication through MEK/ERK kinase and phagocyte NADPH oxidase activities. J. Biol. Chem. 277, 18753–18762 (2002).
Donch, J., Green, M. H. L. & Greenberg, J. Interaction of the exr and lon genes in Escherichia coli. J. Bacteriol. 96, 1704–1710 (1968).
Kantor, G. J. & Deering, R. A. Effect of nalidixic acid and hydroxyurea on division ability of Escherichia coli Fil+ and Lon- strains. J. Bacteriol. 95, 520–530 (1968).
Chantratita, N. et al. Biological relevance of colony morphology and phenotypic switching by Burkholderia pseudomallei. J. Bacteriol. 189, 807–812 (2007).
Nicholls, L. Melioidosis, with special reference to the dissociation of Bacillus whitmori. Br. J. Exp. Pathol. 11, 393–399 (1930).
Ben-Jacob, E. et al. Bacterial cooperative organization under antibiotic stress. Physica A 282, 247–282 (2000).
Cano, D. A., Pucciarelli, M. G., Martínez-Moya, M., Casadesús, J. & García-del Portillo, F . Selection of small-colony variants of Salmonella enterica serovar Typhimurium in nonphagocytic eucaryotic cells. J. Infect. Immun. 71, 3690–3698 (2003).
Hasman, H., Schembri, M. A. & Klemm, P. Antigen 43 and type 1 fimbriae determine colony morphology of Escherichia coli K-12. J. Bacteriol. 182, 1089–1095 (2000).
Lacy, R. W. Dwarf-colony variants of Staphylococcus aureus resistant to aminoglycoside antibiotics and to a fatty acid. J. Med. Microbiol. 2, 187–197 (1969).
Proctor, R. A., van Langevelde, P., Kristjansson, M., Maslow, J. N. & Arbeit, R. D. Persistent and relapsing infections associated with small colony variants of Staphylococcus aureus. J. Clin. Infect. Dis. 20, 95–102 (1995).
Balwit, J. M., van Langevelde, P., Vann, J. M. & Proctor, R. A. Gentamicin-resistant menadione and hemin auxotrophic Staphylococcus aureus persist within cultured endothelial cells. J. Infect. Dis. 170, 1033–1037 (1994).
Baumert, N. et al. Physiology and antibiotic susceptibility of Staphylococcus aureus small colony variants. J. Microb. Drug Resist. 8, 253–260 (2002).
Miller, M. H., Edberg, S. C., Mandel, L. J., Behar, C. F. & Steigbigel, N. H. Gentamicin uptake in wild type and aminoglycoside-resistant small colony mutants of Staphylococcus aureus. J. Antimicrob. Agents Chemother. 18, 722–729 (1980).
Koo, S. P., Bayer, A. S., Sahl, H. G., Proctor, R. A. & Yeaman, M. R. Staphylocidal action of thrombin induced platelet microbicidal protein (tPMP) is not solely dependent on transmembrane potential. J. Infect. Immun. 64, 1070–1074 (1996).
Proctor, R. A. Bacterial energetics and antimicrobial resistance. J. Drug Resist. Updat. 1, 227–235 (1998).
Chambers, H. F. & Miller, M. M. Emergence of resistance to cephalothin and gentamicin during combination therapy for methicillin-resistant Staphylococcus aureus endocarditis in rabbits. J. Infect. Dis. 155, 581–585 (1987).
Musher, D. M., Baughn, R. E., Templeton, G. B. & Minuth, J. N. Emergence of variant forms of Staphylococcus aureus after exposure to gentamicin and infectivity of the variants in experimental animals. J. Infect. Dis. 136, 360–369 (1977).
Wilson, S. G. & Sanders, C. C. Selection and characterization of strains of Staphylococcus aureus displaying unusual resistance to aminoglycosides. J. Antimicrob. Agents Chemother. 10, 519–525 (1976).
Proctor, R. A. et al. Small colony variants: a pathogenic form of bacteria that facilitates persistent and recurrent infections. Nat. Rev. Microbiol. 4, 295–305 (2006).
Anderson, P., Davies, J. E. & Davis, B. D. The effect of spectinomycin on polypeptide synthesis in extracts of Escherichia coli. J. Mol. Biol. 29, 203–208 (1967).
Davies, J., Gorini, L. & Davis, B. D. Misreading of RNA codewords induced by aminoglycoside antibiotics. J. Mol. Pharmacol. 1, 93–106 (1965).
Weisberger, A. S. Inhibition of protein synthesis by chloramphenicol. J. Med. 18, 483–494 (1967).
Calvo, J. & Martínez-Martínez, L. Antimicrobial mechanisms of action]. Enferm. Infecc. MicrobioL. Clin. 27, 44–52 (2009).
Ogle, J. M. & Ramakrishnan, V. Structural insights into translational fidelity. Ann. Rev. Biochem. 74, 129–177 (2005).
Acknowledgements
We thank Pete Chandrangsu and Dr Richard Gourse, Department of Bacteriology, University of Wisconsin-Madison, for providing us with the Lambda Red system and technical assistance with the primer extension assay.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mikheil, D., Shippy, D., Eakley, N. et al. Deletion of gene encoding methyltransferase (gidB) confers high-level antimicrobial resistance in Salmonella. J Antibiot 65, 185–192 (2012). https://doi.org/10.1038/ja.2012.5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2012.5