Abstract
Heritability may be estimated using phenotypic data collected in relatives or in distantly related individuals using genome-wide single nucleotide polymorphism (SNP) data. We combined these approaches by re-parameterizing the model proposed by Zaitlen et al and extended this model to include moderation of (total and SNP-based) genetic and environmental variance components by a measured moderator. By means of data simulation, we demonstrated that the type 1 error rates of the proposed test are correct and parameter estimates are accurate. As an application, we considered the moderation by age or year of birth of variance components associated with body mass index (BMI), height, attention problems (AP), and symptoms of anxiety and depression. The genetic variance of BMI was found to increase with age, but the environmental variance displayed a greater increase with age, resulting in a proportional decrease of the heritability of BMI. Environmental variance of height increased with year of birth. The environmental variance of AP increased with age. These results illustrate the assessment of moderation of environmental and genetic effects, when estimating heritability from combined SNP and family data. The assessment of moderation of genetic and environmental variance will enhance our understanding of the genetic architecture of complex traits.
Similar content being viewed by others
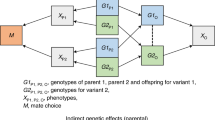
Introduction
Gene–environment (GxE) interaction is an important issue in genetics with potentially important empirical implications.1 In human genetics, the moderation of genetic effects by environmental risk factors has been considered with respect to, for example, psychiatric disorders (for reviews see refs 2, 3, 4, 5, 6), transcriptomics,7 and body mass index (BMI).8, 9 GxE interaction studies have often been criticized for lack of statistical power,10 poor choice of candidate genes, or genetic markers,11, 12 and poor replication.13 To judge whether GxE interaction studies may inform complex trait genetics, and may play a role in explaining missing heritability,5 more knowledge is required about the extent to which GxE interaction contributes to phenotypic variance. Studies that employ genetically informative designs (eg, the twin design),14, 15 and studies including genome-wide genotype data provide a means to evaluate GxE interaction effects. One example of the latter approach involves establishing whether the effect of a polygenic risk score is moderated by an environment variable.16, 17, 18 Such polygenic scores are usually based on a weighed linear combination of single-nucleotide polymorphisms (SNPs) which satisfy some significance level in a GWAS. An alternative approach to quantify the effect of a set of measured SNPs on a phenotype is to fit a random effects model using genetic relatedness matrix restricted maximum likelihood (GREML), as implemented in the GCTA software package.19 GCTA was developed to estimate ‘chip based’ heritability in large groups of distantly related individuals, and allows estimation of GxE interaction, given a dichotomous environmental exposure. Vinkhuyzen and Wray16 recently discussed the current options for GxE interaction research with dichotomous exposures, both for polygenic risk scores and GCTA analyses.
Here, we propose a model that can include a continuous, ordinal, or nominal moderator of genetic and environmental variance components. In addition, we extend the model to inclusion of data of closely related subjects, such as twin pairs, or members of extended pedigrees. To this end, we re-parameterized the model proposed by Zaitlen et al.1 This allows an evaluation of GxE interaction in terms of the moderation of the genetic variance attributable to the measured SNPs, the total additive genetic variance, and the residual (environmental) variance. Including distantly related and closely related subjects increases the power to resolve the moderator's effect on genetic and environmental variance components.
We conducted simulation studies to investigate the performance of the model. First, we simulated data given several null hypotheses to establish that the type-1 error rates for the significance tests used are correct. Second, we established that the estimated variance components were accurate when phenotypic data are normally distributed. GREML is usually based on the assumption of phenotypic multivariate normality. Speed et al.20 previously tested the performance of the model given non-normality due to non-normally distributed environmental effects, they reported little evidence for bias. Here we explored the effects of phenotypic positive skewness on the type-1 error rates and parameter estimates. Positive skewness is often observed in test scores obtained from questionnaires used to asses psychopathology due to an abundance of items with low endorsement rates. We applied our model to BMI and height, and to attention problems (AP) and anxious depression (AnxDep). We estimate the contribution of common genetic variants, all genetic effects, and the environment subject to moderation by age or birth year (for height). We considered birth year the appropriate moderator for height as the variance in adult height does not change with age. The largest variance in adult height is attributable to the mean increase in height throughout the 20th century. The distributions of BMI and height are normal, whereas AP and AnxDep are positively skewed.
Methods
Statistical methods
Genome-wide genetic similarities between individuals based on measured SNPs can be used to estimate the variance attributable to these measured SNPs (for the derivation, see ref. 19). The associated model is provided in Equations 1a–1d.




In Equation 1a, Y (n × 1) is a random vector of phenotypic scores as observed in n individuals, X (n × m) is the matrix of m fixed covariates, and β (m × 1) is the vector of fixed effects. The matrix W (n × p) is the matrix of p standardized SNPs, u (p × 1) is a zero mean vector of random effects (regression coefficients), and e is the n × 1 vector of zero mean residuals. The phenotype Y is assumed to be a random multivariate normal vector with mean vector μ and covariance matrix V (Equation 1b). The GRM is a matrix of pairwise genetic similarities computed as WW′/p. The parameter  is an estimate of the variance explained by the SNPs included in W and SNPs in strong LD with SNPs included in W and I is an identity matrix. The GCTA software can be used to fit the model above, with extensions allowing for dichotomous GxE moderation, among other options.19
is an estimate of the variance explained by the SNPs included in W and SNPs in strong LD with SNPs included in W and I is an identity matrix. The GCTA software can be used to fit the model above, with extensions allowing for dichotomous GxE moderation, among other options.19
The continuous moderation model
The continuous moderation model allows the magnitude of the variance components to vary with respect to a moderator M. Parameter βsnp quantifies the effect of moderation by M of the variance explained in Y by common SNPs. Parameter σsnp quantifies the effect of measured SNPs on Y given βsnp=0. Parameter βe quantifies the effect of moderator M of the residual variance of Y. Parameter σe quantifies the effect of residual variance given βe=0.

In Equation 2, ‘*’ indicates a matrix product and ‘.’ indicates element-wise multiplication. This model allows for moderation of the genetic and residual (co)variances by M. Equation 2 is a variation on the moderation model introduced by Purcell in the context of twin studies (2002).14 Note that we assume that the moderator M is also included in the matrix X, that is, as a fixed covariate with a main effect on the phenotype. We have presented the moderator M as continuous, but it may be discrete (ordinal or nominal). Given a binary moderator M, coded √0.5 and −√0.5, the model in Equation 2 is equivalent to the GxE approach used in GCTA.
Related individuals in the sample
It is a standard practice in GCTA to exclude genetically closely related individuals to avoid confounding of the total heritability and the SNP heritability. An extension proposed by Zaitlen et al.1 allows for inclusion of closely related individuals to estimate the variance due to the SNPs, as well as the total additive genetic variance. In Equation 3 below, the matrix GRMIBS is equivalent to the GRM in Equations 1 and 2, but now includes closely related individuals. Closely related in our analysis is defined as genetic relatedness greater than 0.05, as in Zaitlen et al.1 The matrix GRMIBS>0.05 equals the matrix GRMIBS in which all relatedness coefficients below 0.05 set to zero. Note that the values of these coefficients in closely related individuals tend towards the expected proportion of alleles shared identically by decent (~IBD; we denote the expected proportion with pi-hat) (ie, full siblings are characterized by pi-hat=0.5 IBD, and ~0.5 in the GRMIBS>0.05). Using an IBD matrix or IBS>0.05 matrix yields very similar results.1

In Equation (3), parameter  represents the difference between the total additive genetic variance
represents the difference between the total additive genetic variance  and the variance explained by SNPs, and
and the variance explained by SNPs, and  reflects the variance attributable to residual effects. Inspection of the parameter correlation matrix derived from the Hessian revealed very strong negative parameter correlations between
reflects the variance attributable to residual effects. Inspection of the parameter correlation matrix derived from the Hessian revealed very strong negative parameter correlations between  and
and  which complicates any moderation of these terms. To ensure low-parameter correlations, and to allow for separate moderation of
which complicates any moderation of these terms. To ensure low-parameter correlations, and to allow for separate moderation of  and
and  we re-parameterized the Zaitlen et al. model (Equation 3) as shown in Equation 4.
we re-parameterized the Zaitlen et al. model (Equation 3) as shown in Equation 4.

In Equation 4, the first GRM,  includes values only for pairs of distantly related individuals with IBS<0.05, whereas the other values including the diagonal elements are set to zero. This provides an estimate of variance attributable to SNPs exclusively based on the covariance between distantly related individuals. The second GRM,
includes values only for pairs of distantly related individuals with IBS<0.05, whereas the other values including the diagonal elements are set to zero. This provides an estimate of variance attributable to SNPs exclusively based on the covariance between distantly related individuals. The second GRM,  contains only values for pairs of individuals that are closely related, with IBS>0.05, reflecting all genetic variance as a function of approximate IBD. Note that this model requires the presence of closely related individuals to reliably estimate
contains only values for pairs of individuals that are closely related, with IBS>0.05, reflecting all genetic variance as a function of approximate IBD. Note that this model requires the presence of closely related individuals to reliably estimate  The re-parameterized model (Equation 4) and the Zaitlen et al. model (Equation 3) produce the same estimates of
The re-parameterized model (Equation 4) and the Zaitlen et al. model (Equation 3) produce the same estimates of  and
and  This equivalence was established empirically by simulating data for a wide range of
This equivalence was established empirically by simulating data for a wide range of  and
and  under the Zaitlen et al. model, and subsequently fitting both models, and obtaining the same −2*log-likelihood and parameter estimates (Supplementary Table S1). We note that in the unlikely scenario that
under the Zaitlen et al. model, and subsequently fitting both models, and obtaining the same −2*log-likelihood and parameter estimates (Supplementary Table S1). We note that in the unlikely scenario that  the equivalence does not hold. However, if
the equivalence does not hold. However, if  is true, separate moderation of
is true, separate moderation of  is undesirable given the absence of the variance component to be moderated. Furthermore, note that
is undesirable given the absence of the variance component to be moderated. Furthermore, note that  is not added to the total variance of a trait. This is due to our reparameterization of the Zaitlen et al. model, given that we estimate
is not added to the total variance of a trait. This is due to our reparameterization of the Zaitlen et al. model, given that we estimate  in the matrix diagonal and entries for covariance between closely related subjects is set to 0,
in the matrix diagonal and entries for covariance between closely related subjects is set to 0,  no longer influences the total phenotypic variation.
no longer influences the total phenotypic variation.
We extended Equation 4 to include moderation, as shown in Equation 4a. Parameter β~IBD in Equation 4a reflects the change in additive genetic variance as a function of the moderator M. Parameter βsnp reflects moderation of genetic variance attributable to SNPs as a function of M.

In fitting model 4a, we assume that we have a sufficient number of related individuals to accurately estimate  Although the main innovation presented here is continuous moderation of the variance attributable to SNPs, our model allows for moderation of the other variance components. It is important to consider moderation of all variance components. First, moderation of each of the variance components influences standardized variance components as these are expressed as a ratio. Accordingly, moderation of variance components featured in the denominator of a ratio results in moderation of the entire ratio. Second, allowing moderation of all genetic variance components yields an internally consistent model, as moderation of the variance of a trait explained by common SNPs should be accompanied by moderation of the total additive variance of the trait.
Although the main innovation presented here is continuous moderation of the variance attributable to SNPs, our model allows for moderation of the other variance components. It is important to consider moderation of all variance components. First, moderation of each of the variance components influences standardized variance components as these are expressed as a ratio. Accordingly, moderation of variance components featured in the denominator of a ratio results in moderation of the entire ratio. Second, allowing moderation of all genetic variance components yields an internally consistent model, as moderation of the variance of a trait explained by common SNPs should be accompanied by moderation of the total additive variance of the trait.
Model estimation
All models were fitted in R (Vienna, Austria) using full information maximum likelihood optimization with exact derivatives. The implementation in R includes two FORTRAN routines to speed up calculation of the likelihood function and derivatives. Optimization using exact derivatives is done using the optim() function available in R. Scripts to perform the optimization and the required FORTRAN routines are available online (https://sites.google.com/site/mgnivard/multigrm). We tested the significance of parameters by means of the likelihood ratio test. We adopted a significance level of 0.05 for each test. The four outcome phenotypes were regressed on the first 6 principal components that reflect the population structure.21 The standardized residuals were further analyzed to reduce the number of fixed effects in the final model, and thus reduce computational burden. Fixed effects covariates entered into all analyses were sex and standardized age, and age squared. For height, year of birth rather than age was included as a covariate.
Simulations
Type-1 error
It is well known that the distribution of the test statistic associated with bounded variance components deviate from the standard central χ2 distribution. The approximate null distribution is a 0.5  mixture distributions, but the exact distribution of the test statistic is dependent on the Eigen spectrum of the design matrix (GRM).22 In GCTA, this 0.5:0.5 mixture distribution is used. To establish that this assumed distribution is reasonable, we simulated data sets, based on the empirical GRMs as observed in the NTR data set. First, data were simulated with σsnp=0, σ~IBD=0.2 and σe=0.8 and βsnp=β~IBD=βe=0, and we tested the type-1 error associated with the null hypothesis σsnp=0. Assuming the mixture null distribution, we expected the type-1 error rates to be 0.025 and 0.05 (ie, the nominal significance level divided by 2). In our second scenario, we simulated a data set where: σsnp=0.2, σ~IBD=0.4 and σe=0.6 and βsnp=β~IBD=βe=0. Here we tested the type-1 error associated with the (omnibus) null hypotheses βsnp=0, β~IBD=0 and βe=0 at significance levels 0.1 and 0.05. The moderation parameters are not bounded and therefore should follow the standard distribution
mixture distributions, but the exact distribution of the test statistic is dependent on the Eigen spectrum of the design matrix (GRM).22 In GCTA, this 0.5:0.5 mixture distribution is used. To establish that this assumed distribution is reasonable, we simulated data sets, based on the empirical GRMs as observed in the NTR data set. First, data were simulated with σsnp=0, σ~IBD=0.2 and σe=0.8 and βsnp=β~IBD=βe=0, and we tested the type-1 error associated with the null hypothesis σsnp=0. Assuming the mixture null distribution, we expected the type-1 error rates to be 0.025 and 0.05 (ie, the nominal significance level divided by 2). In our second scenario, we simulated a data set where: σsnp=0.2, σ~IBD=0.4 and σe=0.6 and βsnp=β~IBD=βe=0. Here we tested the type-1 error associated with the (omnibus) null hypotheses βsnp=0, β~IBD=0 and βe=0 at significance levels 0.1 and 0.05. The moderation parameters are not bounded and therefore should follow the standard distribution  To gauge the effect of violations of phenotypic normality, the simulated normal data were transformed by mapping the data onto the empirical distribution of AnxDep scores, while retaining the rank of the data. We then repeated our test type-1 error based on the empirically distributed data.
To gauge the effect of violations of phenotypic normality, the simulated normal data were transformed by mapping the data onto the empirical distribution of AnxDep scores, while retaining the rank of the data. We then repeated our test type-1 error based on the empirically distributed data.
Parameter accuracy given distributional violations
In fitting the model, we usually assume that the data are multivariate normally distributed. However, in practice data are often non-normal. To test the effects of non-normality on parameter estimates, we simulated sets of multivariate normal data, in which the SNP heritability was increased from 0 to 0.5 in 0.01 steps, and the residual heritability was increased from 0 to 0.5 in 0.01 steps for each step. Forty data sets were simulated per step. The simulated normal data were transformed by mapping the data onto the empirical distribution of AnxDep, while retaining the rank of the data. We chose the distribution of AnxDep because its distribution is typical of the distributions arising from questionnaires concerning psychopathology. That is, such data are positively skewed (the AnxDep data has a skewness of 1.62, and the AP data a skewness of 0.91). We estimated the parameters both in the normal and the skewed data, and evaluated the effects of non-normality by a regression of the estimates based on the normal data and non-normal data on the true values. Unbiased estimates will give rise to a slope parameter equal to 1. We separately tested the bias in the SNP heritability variance component, and genetic variance not attributable to SNPs.
Subjects, genotypes and measures
Phenotype data were collected from participants in the Netherlands Twin Register (NTR)23 by mailed or online surveys, or during home visits. Adult participants received surveys in 10 consecutive waves over the past 25 years. Adolescent participants received self-report questionnaires, from the age of 14 onwards. AnxDep and AP scores were obtained from the Youth or Adult Self Report.24 as part of the Achenbach system for empirical assessment.24 AnxDep and AP were defined as described in previous work.25, 26 Height and BMI were assessed during a home visit for the NTR biobank projects.27 and by self-report. AnxDep scores were available in 6881, AP scores in 6618, BMI in 6395, and height in 6409 individuals. As our simulations show that a skewed distribution influences the parameter accuracy in the proposed models, we carried out a square-root transformation of the AP and AnxDep scores to reduce the skewness of the observed distributions. AP had a skewness of 0.91 before transformation and −0.30 after square-root transformation, AnxDep had a skewness of 1.62 before transformation and 0.25 after square-root transformation. Genotypes: DNA samples were obtained in different projects of the NTR.27, 28 Genotyping in the different projects was performed on the Affymetrix 6.0 chip (Affymetrix, Santa Clara, CA, USA). Genotypes were called, cleaned and processed in a single pipeline to ensure consistency across projects. SNPs that were genotyped in less than 95% of individuals were removed. Individuals with a contrast quality control (CQC) score below 0.40, who had less than 90% of SNPs successfully genotyped, or had excess genome-wide heterozygosity/inbreeding levels (F<−0.10 or F>0.10) were removed. Individuals of non-European descent were excluded. The resulting sample included genotypes of 8485 individuals. A genetic relatedness matrix (GRM) was computed on the basis of all autosomal SNPs with a minor allele frequency>0.01, and Hardy–Weinberg Equilibrium test P-value>1 × 10−6 using GCTA 1.24.2.19 Informed consent was obtained from all participants. The study was approved by the Central Ethics Committee on Research Involving Human Subjects of the VU University Medical Centre, Amsterdam, an Institutional Review Board certified by the U.S. Office of Human Research Protections (IRB number IRB-2991 under Federal-wide Assurance-3703; IRB/institute codes, NTR 03-180).
Results
Type-1 error
We simulated 1000 data sets in which the SNP-heritability variance component was zero. To establish the type-1 error rate, we tested the hypothesis that the variance component was zero using a naive chi.2 1) test of the likelihood ratio with significance levels set at 0.1 and 0.05, but expect type-1 error rate of 0.05 and 0.025, respectively, as the variance estimate is bounded by zero. This test produced the type-1 error rates of 0.057 and 0.029. These values are consistent with expectation, that is, they do not deviate significantly (given α=0.01) from the expected values of 0.025 and 0.05. Type-1 error rates calculated using the skewed data produced type-1 error rates of 0.035 and 0.059, neither of which deviated significantly (α=0.01) from the expected values of 0.025 and 0.05.
We simulated 1000 data sets in which moderation of the SNP, additive genetic, and environmental variance component was absent. We estimated a model with and without the 3 moderation parameters, and tested the likelihood ratio (a 3 df χ2 test) using a significance levels of 0.05 and 0.10. The observed type-1 error rates were 0.047 and 0.097, which do not deviate significantly (α=0.01) from the expected values of 0.05 and 0.10. Type-1 error rates for the omnibus test of moderation calculated using the skewed data were 0.181 and 0.266, revealing that distributional violations can result in inaccurate type-1 error rates.
Effects of the distribution on parameter estimates
We simulated 1000 data set with the SNP and additive heritable components increasing in 100 steps of 0.0033, whereas the environmental variance component decreased in 50 steps of 0.0066. We simulated 10 data sets per step. The parameter recovery was found to be good. The slope from the regression of the estimated parameter σsnp on the true parameter is 1.01 (SE=0.035) and for the regression of the estimated parameter σ~IBD on the true parameter the slope is 0.97 (SE=0.012) (Supplementary Figures S1 and S2). However, the parameter estimates obtained in the analysis of the skewed data were biased. Regression of the parameter estimates on the true parameter values yielded a slope of 0.81 (SE=0.033) for σsnp and a slope of 0.84 (SE=0.013) for σ~IBD. Regression of the true parameter difference (σ~IBD−σsnp, ie the genetic variance not attributable to SNPs) on the estimated difference yielded a slope of 0.93 (SE=0.04) for the normal data, and 0.88 (SE=0.039) for the transformed data (Supplementary Figure S3). We therefore conclude that the particular non-normality considered here resulted in an underestimation of the genetic variance attributable to SNPs and the genetic variance not attributable to SNPs, where the bias in the estimate of the genetic variance attributable to SNPs seemed to be greater.
Heritability and variance explained by SNPs
The variances attributable to total and SNP related genetic effects (Equation 3, see methods) on BMI, Height, AP and AnxDep are given in Table 1. All estimates are significant (P-values<0.05) except the variance explained by SNPs in AnxDep (P=0.078). SNP heritability for BMI and height were moderate and total heritability was substantial. SNP heritability of AP and AnxDep was low, and the total additive heritability was moderate. The percentage of the genetic variance of BMI, height, AP, and AnxDep explained by the SNPs was 56.2% (SE 8.6%), 59 % (SE 6.9%), 27.5% (SE 14.1%), and 24.3% (SE 14.1%), respectively. The genetic variance of AnxDep and AP may be underestimated, and type-1 error rates would be inflated due to the non-normality of the phenotypic data. As indicated in the methods, AP and AnxDep scores were square-root transformed to reduce the skewness. As the genetic variance explained by SNPs for BMI was somewhat higher than that reported in the literature.29 we repeated the analysis for BMI in GCTA, based on data of 3119 distantly related individuals only. Here the SNP heritability estimate was 49% (SE 11.4%). Analysis carried out in GCTA based on 6395 individuals (including closely related and distantly related subjects in the GRM, and therefore estimating a variance component dominated by the closely related subjects, approximately equal to σ~IBD) resulted in a heritability estimate of 75.4% (SE 1.3%). These results are very close to those obtained with our methods.
Genetic variance moderation
Next we fitted the model given in Equation 4a (see methods) to all four variables with age or birth year as the moderator of the variance components  Moderation of the variance components attributable to SNP effects were not significant for any of the phenotypes (Table 2). For BMI, the total genetic variation was significantly moderated by age. For BMI, height and AP moderation of the residual effects was significant; for AnxDep, no significant moderation by age was seen.
Moderation of the variance components attributable to SNP effects were not significant for any of the phenotypes (Table 2). For BMI, the total genetic variation was significantly moderated by age. For BMI, height and AP moderation of the residual effects was significant; for AnxDep, no significant moderation by age was seen.
Figures 1a, 2a and 3a, show  of as a function of age or birth year for BMI, height, and AP. Figures 1b, 2b and 3b show the heritability
of as a function of age or birth year for BMI, height, and AP. Figures 1b, 2b and 3b show the heritability  and the proportion of phenotypic variance attributable to SNPs
and the proportion of phenotypic variance attributable to SNPs  as a function of age or birth year. Note the denominator in these ratios does not include
as a function of age or birth year. Note the denominator in these ratios does not include  as our model parameterization
as our model parameterization  is estimated separately from the total additive genetic variance
is estimated separately from the total additive genetic variance  All figures show that the total heritability and the SNP heritability decrease as a function of age, or birth year.
All figures show that the total heritability and the SNP heritability decrease as a function of age, or birth year.
Discussion
We presented a variance component model which included moderation of SNP genetic, total additive genetic and residual effects. The moderator is assumed to be a measured variable, which may be continuously or discretely distributed. The model can be used in cohorts of unrelated, and/or distantly related, individuals to estimate the moderation of the genetic effects attributable to SNPs. In pedigree data, the model separately estimates the moderation of SNP effects and of total additive genetic effects. By allowing for simultaneous moderation of SNP and total additive genetic variance, we can include data from twin and family cohorts, and estimate the genetic variance attributable to SNPs, and its moderation while retaining all participants in the analyses. We tested the type-1 error associated with a χ2 test and found that the type-1 error rate is accurate for the test of the unmoderated variance component estimate being larger than zero. For the (omnibus) test of continuous moderation of variance components, the type-1 error rate of the test was also accurate. We further investigated the effects of phenotypic non-normality on the (SNP) heritability estimates. We found that both the variance attributable to SNPs and the residual variance are underestimated by positive skewness of the phenotypic distribution. The variance attributable to SNPs was underestimated more than the additive genetic variance not captured by SNPs. Thus, our simulations showed good type-1 error rates for the χ2 test of variance components, and good type-1 error rates for the χ2 test of moderator effects. The type-1 error rates remained good when the data were transformed to match the empirical distribution of the AnxDep scores before square-root transformation. Type-1 error for the test of moderation was accurate if the distributional assumptions were met, violations of the distributional assumptions can induce false positive results.
The moderation models were fitted to data on BMI, height, AP, and AnxDep. Results showed differences between these phenotypes in genetic architecture that were not limited to differences in  but also were the results of differences in the degree to which these variance components were moderated by age or year of birth. The results indicated that for AnxDep, neither the genetic nor environmental effects were moderated by age. For AP, BMI, and height the residual variance,
but also were the results of differences in the degree to which these variance components were moderated by age or year of birth. The results indicated that for AnxDep, neither the genetic nor environmental effects were moderated by age. For AP, BMI, and height the residual variance,  was found to increase with age or birth year. For BMI, age had a positive moderating effect on the additive genetic variance. We found no evidence of moderation of
was found to increase with age or birth year. For BMI, age had a positive moderating effect on the additive genetic variance. We found no evidence of moderation of  in any of the phenotypes considered. These differences in results confirm differences in genetic architectures of these phenotypes. As established in twin studies, both AP and AnxDep are moderately heritable,25, 26 whereas BMI and height are strongly heritable.30, 31 The moderation findings for BMI (decreasing heritability with age) agree with a recent meta-analysis into the heritability of BMI30 and with evidence for GxE at the SNP level. For example, Rosenquist et al.8 reported evidence for an interaction between the FTO gene and birth cohort on BMI. Note the model fitted to the empirical data did not allow for a common environment within family, or for dominant genetic effects, these effects may be of interest and could be accommodated.32
in any of the phenotypes considered. These differences in results confirm differences in genetic architectures of these phenotypes. As established in twin studies, both AP and AnxDep are moderately heritable,25, 26 whereas BMI and height are strongly heritable.30, 31 The moderation findings for BMI (decreasing heritability with age) agree with a recent meta-analysis into the heritability of BMI30 and with evidence for GxE at the SNP level. For example, Rosenquist et al.8 reported evidence for an interaction between the FTO gene and birth cohort on BMI. Note the model fitted to the empirical data did not allow for a common environment within family, or for dominant genetic effects, these effects may be of interest and could be accommodated.32
The method presented here has some limitations. If related individuals are present in the sample, and the moderator itself is heritable, this needs to be accounted for as shown by van der Sluis et al.33 All GxE models are sensitive to scaling.34 and GxE interaction effects reported here also are conditional on the scale of the outcome variables; different scaling may yield different results. Where BMI and height have a natural scale (ie, kg/m2 and m or cm), the scale of the data based on self-report inventories is generally arbitrary. This problem is not limited to the current model and its solution is beyond the scope of this article (potential solutions are discussed elsewhere15, 34). Estimates of SNP heritability are unbiased if the residual variance is not normally distributed.20 Our simulation revealed, however, that even if the underlying variance components are normally distributed, some rating scales give rise to skewed distributions and this may result in parameter bias, and inflated type-1 error for the test of moderation. In view of our simulation results, it is possible that the (genetic) variance attributable to SNPs for AP and AnxDep are underestimated due to the skewness of the AP and AnxDep scores. As we found little evidence of age moderation for AP and AnxDep, the evidence we find (age moderation of the environment for AP) should be interpreted cautiously given the distributions of these phenotypes. To mitigate the distributional violations the AP and AnxDep scores were square-root transformed before analysis to reduce skew. In light of these limitations, presence or absence of age moderation of genetic variance components, for the traits AP and AnxDep need to be replicated across different self-report scales or diagnostic instruments, reducing the reliance on a single (arbitrary) scale.
The present method can include data from distantly and closely related individuals, it can accommodate, categorical, ordinal or continuous moderators, and moderation multiple variance components simultaneously. The proposed method has been successfully applied to detect moderation of the (additive) genetic and environmental effects by age and sex for 400 000+ methylation probes.35 The analysis of these 400 000 phenotypes showed that large scale implementation is feasible. The addition of multiple genetic effects (GRMs) further allows for the separate moderation of different subsets of the genome. For example, one could limit the SNPs in a GRM to a single biological pathway (ie, SNPs in genes in the serotonin pathway), to a single class of SNPs (ie, coding variants), or to specific regions of the genome (ie, regulatory elements, the exome, etc.). Bearing in mind the assumptions discussed above the moderator can be genetic (GxG ie, a known risk variant), biological (ie, a gene expression; gut microbiota) or environmental (ie, early childhood trauma experiences).
References
Zaitlen N, Kraft P, Patterson N et al: Using extended genealogy to estimate components of heritability for 23 quantitative and dichotomous traits. PLoS Genet 2013; 9: e1003520.
Caspi A, Hariri AR, Holmes A, Uher R, Moffitt TE : Genetic sensitivity to the environment: the case of the serotonin transporter gene and its implications for studying complex diseases and traits. Am J Psychiatry 2010; 167: 509–527.
Carpenter RW, Tomko RL, Trull TJ, Boomsma DI : Gene-environment studies and borderline personality disorder: a review. Curr Psychiatry Rep 2013; 15: 1–7.
Dunn EC, Uddin M, Subramanian SV et al: Research review: gene-environment interaction research in youth depression-a systematic review with recommendations for future research. J Child Psychol Psychiatry 2011; 52: 1223–1238.
Uher R : Gene-environment interactions in common mental disorders: an update and strategy for a genome-wide search. Soc Psychiatry Psychiatr Epidemiol 2014; 49: 3–14.
Young-Wolff KC, Enoch MA, Prescott CA : The influence of gene-environment interactions on alcohol consumption and alcohol use disorders: a comprehensive review. Clin Psychol Rev 2011; 31: 800–816.
Buil A, Brown AA, Lappalainen T et al: Gene-gene and gene-environment interactions detected by transcriptome sequence analysis in twins. Nat Genet 2015; 47: 88–91.
Rosenquist JN, Lehrer SF, O’Malley AJ et al: Cohort of birth modifies the association between FTO genotype and BMI. Proc Natl Acad Sci USA 2014; 2: 354–459.
Huang T, Hu FB : Gene-environment interactions and obesity: recent developments and future directions. BMC Med Genomics 2015; 8: S2.
Munafo MR, Flint J : Replication and heterogeneity in gene x environment interaction studies. Int J Neuropsychopharmacol 2009; 12: 727–729.
Bosker FJ, Hartman CA, Nolte IM et al: Poor replication of candidate genes for major depressive disorder using genome-wide association data. Mol Psychiatry 2011; 16: 516–532.
Sullivan PF, Daly MJ, O’Donovan M : Genetic architectures of psychiatric disorders: the emerging picture and its implications. Nat Rev Genet 2012; 13: 537–551.
Duncan LE, Keller MC : A critical review of the first 10 years of candidate gene-by-environment interaction research in psychiatry. Am J Psychiatry 2011; 168: 1041–1049.
Purcell S : Variance components models for gene-environment interaction in twin analysis. Twin Res 2002; 5: 554–571.
Molenaar D, Dolan CV : Testing systematic genotype by environment interactions using item level data. Behav Genet 2014; 44: 212–231.
Vinkhuyzen AAE, Wray NR : Novel directions for G x E analysis in psychiatry. Epidemiol Psychiatr Sci 2015; 24: 1–8.
Wray NR, Yang J, Hayes BJ et al: Pitfalls of predicting complex traits from SNPs. Nat Rev Genet 2013; 14: 507–515.
Peyrot WJ, Milaneschi Y, Abdellaoui A et al: Effect of polygenic risk scores on depression in childhood trauma. Br J Psychiatry 2014; 205: 113–119.
Yang J, Lee SH, Goddard ME, Visscher PM : GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 2011; 88: 76–82.
Speed D, Hemani G, Johnson MR, Balding DJ : Improved heritability estimation from genome-wide SNPs. Am J Hum Genet 2012; 91: 1011–1021.
Abdellaoui A, Hottenga JJ, de Knijff P et al: Population structure, migration, and diversifying selection in the Netherlands. Eur J Hum Genet 2013; 21: 1277–1285.
Crainiceanu CM, Ruppert D : Likelihood ratio tests in linear mixed models with one variance component. J R Stat Soc Series B Stat Methodol 2004; 66: 165–185.
Boomsma DI, de Geus EJ, Vink JM et al: Netherlands Twin Register: from twins to twin families. Twin Res Hum Genet 2006; 9: 849–857.
Achenbach, TM, Rescorla, LA, McConaughey, S, Pecora, P, Wetherbee, K, Ruffle, T: The Achenbach system of empirically based assessment; Handbook of Psychological and Educational Assessment of Children: Personality, Behavior, and Context. 2003, Vol 2, pp 406–432..
Kan KJ, Dolan CV, Nivard MG et al: Genetic and environmental stability in attention problems across the lifespan: evidence from the Netherlands twin register. J Am Acad Child Adolesc Psychiatry 2013; 52: 12–25.
Nivard MG, Dolan CV, Kendler KS et al: Stability in symptoms of anxiety and depression as a function of genotype and environment: a longitudinal twin study from ages 3 to 63 years. Psychol Med 2015; 45: 1039–1049.
Willemsen G, Vink JM, Abdellaoui A et al: The Adult Netherlands Twin Register: twenty-five years of survey and biological data collection. Twin Res Hum Genet 2013; 16: 271–281.
Willemsen G, de Geus EJ, Bartels M et al: The Netherlands Twin Register biobank: a resource for genetic epidemiological studies. Twin Res Hum Genet 2010; 13: 231–245.
Yang J, Manolio TA, Pasquale LR et al: Genome partitioning of genetic variation for complex traits using common SNPs. Nat Genet 2011; 43: 519–525.
Elks CE, den Hoed M, Zhao JH et al: Variability in the heritability of body mass index: a systematic review and meta-regression. Front Endocrinol 2012; 3: 2–16.
Silventoinen K, Sammalisto S, Perola M et al: Heritability of adult body height: a comparative study of twin cohorts in eight countries. Twin Res 2003; 6: 399–408.
Zhu Z, Bakshi A, Vinkhuyzen AA et al: Dominance genetic variation contributes little to the missing heritability for human complex traits. Am J Hum Genet 2015; 96: 377–385.
van der Sluis S, Posthuma D, Dolan CV : A note on false positives and power in G x E modelling of twin data. Behav Genet 2012; 42: 170–186.
Eaves L, Verhulst B : Problems and pit-falls in testing for G x E and epistasis in candidate gene studies of human behavior. Behav Genet 2014; 44: 578–590.
van Dongen J, Nivard MG, Willemsen G et al: Genetic and environmental influences interact with age and sex in shaping the human methylome. Nat Commun 2016; 7: 11115.
Greene WH : Econometric Analysis. Granite Hill Publishers: Upper Saddle River, NJ, USA, 2008.
Acknowledgements
This study was supported by the following: Genetic influences on stability and change in psychopathology from childhood to young adulthood (ZonMW 91210020); Genetics of Mental Illness (European Research Council ERC-230374); Biobanking and Biomolecular Resources Research Infrastructure (BBMRI—NL, 184.021.007); Community’s Seventh Framework Program (FP7/2007–2013); ENGAGE (HEALTH-F4-2007-201413); the Avera Institute, Sioux Falls, South Dakota (USA) and Grand Opportunity grants 1RC2 MH089951 and 1RC2 MH089995. GL was supported by the National Institute on Drug Abuse Grant No. DA018673. MGN is supported by a Royal Netherlands Academy of Science Professor Award (PAH/6635) to DIB. We thank Karin Verweij for critically reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Nivard, M., Middeldorp, C., Lubke, G. et al. Detection of gene–environment interaction in pedigree data using genome-wide genotypes. Eur J Hum Genet 24, 1803–1809 (2016). https://doi.org/10.1038/ejhg.2016.88
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2016.88
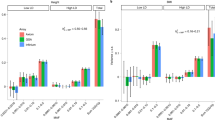




 in AP as a function of age. (b) the heritability
in AP as a function of age. (b) the heritability  and proportion of phenotypic variance attributable to SNPs
and proportion of phenotypic variance attributable to SNPs  in AP as a function of age. The histogram displays the distribution of the standardized moderator age.
in AP as a function of age. The histogram displays the distribution of the standardized moderator age.
 in BMI as a function of age. (b) The heritability
in BMI as a function of age. (b) The heritability  and proportion of phenotypic variance attributable to SNPs
and proportion of phenotypic variance attributable to SNPs  in BMI as a function of age. The histogram displays the distribution of the standardized moderator age.
in BMI as a function of age. The histogram displays the distribution of the standardized moderator age.
 in height as a function of birth year. (b) The heritability
in height as a function of birth year. (b) The heritability  and proportion of phenotypic variance attributable to SNPs
and proportion of phenotypic variance attributable to SNPs  in heigh as a function of birth year. The histogram displays the distribution of the standardized moderator birth year.
in heigh as a function of birth year. The histogram displays the distribution of the standardized moderator birth year.