Abstract
Background:
Hepatocellular carcinoma (HCC) is one of the most important sanitary problems for its prevalence and poor prognosis. To date, no information is available on the prognostic value of the ov-serpin SERPINB3, detected in primary liver cancer but not in normal liver. The aim of the study was to analyse SERPINB3 expression in liver cancer in relation with molecular signatures of poor prognosis and with clinical outcome.
Methods:
Liver tumours of 97 patients were analysed in parallel for SERPINB3, TGF-β and β-catenin. In a subgroup of 67 patients with adequate clinical follow-up, the correlation of molecular findings with clinical outcome was also carried out.
Results:
High SERPINB3 levels were detectable in 22% of the patients. A significant correlation of this serpin with TGF-β at transcription and protein level was observed, whereas for β-catenin a strong correlation was found only at post-transcription level. These findings were in agreement with transcriptome data meta-analysis, showing accumulation of SERPINB3 in the poor-prognosis subclass (S1). High levels of this serpin were significantly associated with early tumour recurrence and high SERPINB3 was the only variable significantly associated with time to recurrence at multivariate analysis.
Conclusions:
SERPINB3 is overexpressed in the subset of the most aggressive HCCs.
Similar content being viewed by others
Main
Hepatocellular carcinoma (HCC) is one of the leading causes of cancer-related death worldwide. The majority of the patients, who are usually diagnosed at an advanced stage, still lack effective treatment and have an extremely dismal prognosis. Hepatocellular carcinoma nearly always develops in the setting of liver cirrhosis and hepatitis B and C viral infections; alcohol abuse and metabolic syndrome are the main risk factors (El-Serag and Rudolph, 2007; Zhang and Friedman, 2012). Liver surgical resection, transplantation and percutaneous ablation are potentially curative therapies in early stages of the disease, applicable to about one-third of the patients (Llovet et al, 2003). The occurrence of tumour recurrence is a frequent event in treated patients, and better stratification of the disease stage, according to the risk of recurrence and survival, remains a still unmet clinical need. Tumour molecular markers predictive of recurrence might become a useful tool to allocate the correct tumour stage that better reflects patient’s prognosis in order to provide the best treatment choice to individual patients. In addition, novel therapeutic approaches, especially for the adjuvant setting, could be supplied to selected patients, optimising the expected results.
In the past years data about molecular mechanisms of liver carcinogenesis, signal transduction pathways and potential therapeutic targets have been accumulated, providing new encouraging treatment options (Llovet and Bruix, 2008). A meta-analysis of gene expression profiles in datasets from nine independent patient cohorts across the world (Hoshida et al, 2009, 2010) allowed the identification of molecular subclasses of HCCs correlated with their histological, molecular and clinical features. In this setting, transforming growth factor β (TGF-β) signalling associated with Wnt target gene expression was identified as one of the most important features of more aggressive HCCs. Wnt signalling is one of the best characterised oncogenic pathways in HCC (Zucman-Rossi et al, 2007), detected in ∼50% of the patients (Villanueva et al, 2007). It is worth noting that cytoplasmic accumulation of β-catenin (the principal downstream effector of Wnt canonical activation) was found to be independent of β-catenin mutations in the subclass of more aggressive tumours (Hoshida et al, 2009). TGF-β is a polyfunctional regulator of cell growth and differentiation, but sometimes has inhibitory functions (Pasche, 2001). Overexpression of this cytokine has been found in hepatic tumours and correlated with carcinogenesis, progression and prognosis (Blobe et al, 2000; Bissell et al, 2001).
Microarray studies have shown transcriptional overexpression of Smad4, the main effector of TGF-β signalling (Zhong et al, 2010), in a subset of HCCs (Shirota et al, 2001; Xu et al, 2001). Recent data indicate that miR-146b-5p, which binds 3′UTR of SMAD4 and decreases TGF-β signalling (Geraldo et al, 2012), is downregulated in several cancers (Bhaumik et al, 2008; Man et al, 2011), including HCC (Gramantieri et al, 2007).
Recent data from our group have shown that the expression of TGF-β can be increased by the ov-serpin SERPINB3 (Turato et al, 2010). This molecule is frequently upregulated in several malignancies of epithelial origin (Takeshima et al, 1992; Cataltepe et al, 2000). In the liver, SERPINB3 and its isoform SERPINB4, previously known as squamous cell carcinoma antigen (SCCA), are undetectable in normal hepatocytes, but their expression progressively increases in chronic liver diseases (Beneduce et al, 2005), dysplastic nodules (Guido et al, 2008) and HCC (Pontisso et al, 2004; Trerotoli et al, 2009), suggesting that they may be involved in relatively early events of hepatocarcinogenesis, although their specific role(s) have not been defined yet (Biasiolo et al, 2012). In vitro studies have shown that SERPINB3 protects neoplastic cells from apoptotic death induced by several kinds of stimuli (Vidalino et al, 2009). Recent data have revealed that SERPINB3 induces epithelial-mesenchimal transition (EMT) and cell proliferation associated with dowregulation of E-caderin, and increased β-catenin expression (Quarta et al, 2010). In addition, this serpin has been found in the majority of hepatoblastomas, where the highest levels were found in tumours of more advanced stage (Turato et al, 2012). To date, no information is available on the prognostic role of SERPINB3 in HCC. The aim of the present study was to analyse the expression of this serpin in human HCC in relation to TGF-β and β-catenin expression and to clinical outcome.
Materials and Methods
Patients and samples
In total, 97 out of 279 patients with HCC undergoing surgical resection as first-line therapy (without any pre-operative anticancer treatment and distant metastases), evaluated at the Hepatobiliary Surgery and Liver Transplantation Unit of the Padua University Hospital between January 2004 and December 2010, were included in the study. All the patients met the following inclusion criteria: (1) HCC diagnosis based on AASLD radiological criteria (Bruix et al, 2005, 2011) or histology, when appropriate, (2) written informed consent and (3) patient’s eligibility for radical therapies according to published schedules (Bruix et al, 2001; Cillo et al, 2004). After discharge, the patients were followed up regularly, with monthly clinical and laboratory assessment, and US and CT scan and/or MRI every 3rd and 6th month, respectively. When HCC recurrence was noted, the patients were further treated with the most appropriate procedure, including repeated resection, percutaneous/laparoscopic/toracoscopic ablation, chemoembolisation or liver transplantation (Cillo et al, 2007).
Tumour tissue samples were collected at the time of surgical resection and part was formalin fixed and paraffin embedded, whereas the remaining part was immediately frozen at −80 °C for further analysis. A subgroup of 67 patients was also considered for the prognostic study on the basis of the following criteria: (a) tumour tissue adequate for molecular analysis; (b) confirmed pathologic diagnosis and surgical radicality; (c) complete response to therapy at 1 month CT scan; and (d) complete clinical-pathological and follow-up data.
Baseline characteristics of these patients are reported in Table 1.
Real-time PCR analysis in liver tumours
mRNA expression of SERPINB3, TGF-β1and β-catenin was assessed in parallel by real-time PCR in the 97 frozen liver tumours collected at the time of surgery.
Total RNA was extracted using RNeasy Mini Kit (Qiagen GmbH, Hilden, Germany) according to the manufacturer’s instructions. After determination of the purity and the integrity of total RNA, real-time amplification of SERPINB3 and TGF-β was carried out as previously described (Turato et al, 2010), whereas for β-catenin amplification the following set of primers was used: sense, 5′-TGGTGCCCAGGGAGAACCCC-3′; reverse 5′ TGTCACCTGGAGGCAGCCCA-3′.
The housekeeping genes HPRT1 and glyceraldehyde-3-phosphate dehydrogenase were amplified in parallel in all amplification sets. mRNA amounts were calculated according to the threshold cycle of individual genes and their relative expression was quantified by serial dilutions of the amplified products compared with external standard curves of the reference genes containing known amounts of each gene product. The results were expressed as a relative ratio of the target to the housekeeping gene using the Light Cycler Relative Quantification software 4.05 (Roche Diagnostics, Monza, Italy).
Samples were run in triplicate, and mRNA expression was generated for each sample. Specificity of the amplified PCR products was determined by melting curve analysis and confirmed by agarose gel electrophoresis and ethidium bromide staining.
Analysis of miRNA 146b-5p
miRNA 146b-5p was assessed by qRT-PCR in HepG2 cells stably transfected with the human SERPINB3 gene or with the empty vector alone as control, obtained as previously described. (Quarta et al, 2010). The 67 frozen tumour samples from the patients resected for HCC were also studied for miRNA 146b-5p. Briefly, 10 ng of total RNA was reverse transcribed using a TaqMan MicroRNA Reverse Transcription kit (Applied Biosystem, Foster City, CA, USA) and miR-146b-5p expression was detected in the cDNA product, using specific TaqMan MiRNA Assay (Applied Biosystem), according to manufacturer’s instruction. PCR amplification was carried out using the CFX96 Real-Time instrument (Bio-Rad Laboratories Inc, Hercules, CA, USA) and U6 small nuclear 2 RNA (RNU6B) was used for normalisation of the results. Samples were run in triplicate and fold changes were generated for each sample by calculating 2−ΔΔCt (Livak and Schmittgen, 2001).
Immunohistochemistry
Five-micrometre-thick paraffin-embedded sections from tumour samples were deparaffinised, rehydrated, immersed in 10 mM l−1 sodium citrate buffer, pH 6 and microwave heated for 4–5 min cycles at 750 W. After rinsing, endogenous peroxidase activity was inactivated with 3% H2O2. Sequential sections were then incubated overnight at 4 °C with specific anti-SERPINB3 monoclonal antibody (1 : 20, SCCA-1 clone 8H11, Santa Cruz Biotechnology, Santa Cruz, CA, USA), anti-TGF-β1 monoclonal antibody (1 : 300, TGFβ1 clone 3C11, Santa Cruz Biotechnology) and anti-β-catenin monoclonal antibody (1 : 500, β-catenin clone N1N2-2, GeneTex, Irvine, CA, USA). After incubation with the primary antibody, immunostaining was performed using the ready-to-use, peroxidase-conjugated EnVision reagent (DakoCytomation, Carpinteria, CA, USA). For quantitative analysis, 10 images of representative fields of 9 HCC samples (5 with low SERPINB3 mRNA expression and 4 with high SERPINB3 mRNA expression) were captured by Leica Qwin Plus v3 software (Cambridge, UK), under a CCD camera connected to a Leica microscope (Microsystem Imaging Solutions Ltd, Cambridge, UK). The average sum of intensities and stained-area percentage of each sample was calculated using ImageJ software (http://rsb.info.nih.gov/ij). Negative controls, obtained by replacing primary antibodies with PBS, were always run in parallel.
Statistical analysis
The primary end point of the present study was to assess the association of SERPINB3 with TGF-β1 and β-catenin expression in liver tumours of surgically resected patients. In addition, clinical significance of SERPINB3 in tumour samples was assessed by analysing its expression in relation to time to recurrence, where early recurrence was defined as HCC reappearance within 24 months from radical resection. Values for continuous variables were presented as medians (ranges) and values for categorical–nominal variables as frequencies (%).
For subgroup comparisons, quantitative variables were compared using Wilcoxon rank sums tests and categorical variables were compared using χ2 or Fisher’s exact tests.
The length of the follow-up after resection was calculated from the date of the operation to the date of the patient’s death or of the latest follow-up. Time to recurrence was defined from the date of the operation to the date of HCC recurrence, and latest follow-up and patient death before recurrence were considered as censor points. The overall time to recurrence curves were calculated using the Kaplan-Meier technique and compared using the log-rank test.
Univariate and multivariate logistic regression analysis was carried out to identify clinical and/or molecular predictors of early tumour recurrence. For univariate analysis, variables collected in our prospective database were considered (Table 1). As for liver function variables, we preferred to consider only complex variables (i.e. Child–Pugh class, model end-stage liver disease score) with respect to single laboratory values or clinical signs, as these scores are more commonly used for clinical decision.
Similarly, we preferred to consider only BCLC (Barcelona Clinic Liver Cancer) stage A vs B–C (corresponding to within and beyond Milan criteria) with respect to radiologic size and number of nodules as separate covariates. For molecular variables, a cutoff value greater than median was defined as ‘high mRNA expression’ and was considered as a dichotomous categorical variable, compared with the remaining cases, which were defined as ‘low mRNA expression’.
For multivariate analysis, only variables achieving a significant result in univariate analysis were included.
All statistical calculations were performed using JMP software (1989–2003 SAS Institute Inc., Cary, NC, USA).
Association of SERPINB3 expression with molecular HCC classification was evaluated by analysing the previously reported data set (NCBI Gene Expression Omnibus accession number GSE10186) (Hoshida et al, 2009). The molecular subclasses, S1 (TGF-β/WNT activation), S2 (MYC/AKT activation, AFP positivity) and S3 (preserved hepatocyte function), were determined by using the nearest template prediction algorithm (Hoshida, 2010) implemented in GenePattern genomic analysis toolkit (www.broadinstitute.org/genepattern). Samples with prediction confidence P-value <0.05 were evaluated for the association.
Results
Molecular profiles in liver tumours
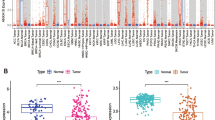
Among the 97 HCC samples analysed, high mRNA levels were detected in 22% of the patients for SERPINB3, in 48% for TGF-β1, and in 51% for β-catenin, respectively, with different relative levels in individual cases. Higher levels of TGF-β1 expression were found in tumour samples positive for SERPINB3, compared with negative samples (Figure 1A) and a positive correlation between the expression of the two molecules was also observed (Figure 1C), whereas β-catenin did not show any correlation at transcription level with the other molecular markers (Figure 1B and D). Different findings were detected at protein level, where a marked overlap of SERPINB3, TGF-β1 and β-catenin was observed. As shown in Figure 2 (panel B), in serial sections of cases with high SERPINB3 mRNA, a strong reactivity for cytoplasmic SERPINB3, TGF-β1 and also for β-catenin was observed by immunohistochemistry. The scenario was markedly different in cases with low SERPINB3 mRNA, where significantly lower expression of the three molecules was detected by quantitative analysis, and β-catenin was typically located in the nucleus, as shown in the example of Figure 2 (panel A).
SERPINB3, TGF- β 1 and β -catenin mRNA in human HCCs. TGF-β1 (A) and β-catenin (B) mRNA were analysed by real-time PCR in relation to the presence (SERPINB3+; N=44) or absence (SERPINB3−; N=53) of SERPINB3 mRNA. The y axis represents the relative mRNA amount of the normalised genes, calculated by dividing the non-normalised values by the housekeeping genes and expressed as arbitrary units. The analysis was performed with Mann–Whitney test. In the lower panels correlation of SERPINB3 with TGF-β1 (C) and β-catenin (D) mRNA in the same HCC sample are depicted.
Immunohistochemical analysis in human HCCs. Representative example of immunohistochemistry for SERPINB3, TGF-β1 and β-catenin, counterstained with hematoxylin, in serial sections of HCC specimens showing low SERPINB3 mRNA (A) or high SERPINB3 mRNA (B). Original magnification are reported. (C) Graphical representation of the quantitative analysis of each staining expressed as the mean of percentage of positive parenchyma staining per area in analysed tumour samples. Bars represent s.d. values. *P<0.0001 (HCCs with low SERPINB3 mRNA vs HCCs with high SERPINB3 mRNA).
Transcriptome data meta-analysis
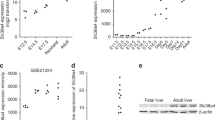
The above results were in agreement with the data of transcriptome meta-analysis. This approach identified molecular subclasses of HCC, S1, S2 and S3 (Hoshida et al, 2009), where the subclass S1 was characterised by the activation of TGF-β and WNT pathways, EMT-like phenotype and more disseminative clinical phenotype, evidenced as more frequent early tumour recurrence. HCC tumours with outstandingly high SERPINB3 expression were indeed accumulated in the S1 subclass (Figure 3), suggesting that SERPINB3 may contribute to the S1-like molecular and clinical phenotypes in the subset of S1 tumours.
SERPINB3 expression and human HCC molecular subclass. SERPINB3 expression level according to molecular subclass of HCC. SERPINB3 expression levels were extracted from publicly available transcriptome dataset (NCBI Gene Expression Omnibus accession number GSE10186) (n=97). Samples are ordered according to previously reported molecular subclasses of HCC determined by a meta-analysis of the following transcriptome datasets: S1 (activation of TGF-β/WNT pathways, EMT-like disseminative phenotype), S2 (MYC/AKT activation, AFP positivity) and S3 (preserved hepatocyte function) (Hoshida et al, 2009).
Modulatory effect of SERPINB3 on miR-146b-5p
Because of the central role of Smad4 in the TGF-β signalling pathway (Rooke and Crosier, 2001; Shi and Massague, 2003), we hypothesised that modification of miR-146b-5p, involved in TGF-β/SMAD signalling pathways, may contribute to TGF-β modulation mediated by SERPINB3. In keeping with this hypothesis, miR-146b-5p was 2.04 fold downregulated in SERPINB3-transfected cells (Figure 4A) and these features were confirmed in human HCCs, where downregulation of miR-146b-5p was significantly more consistent in SERPINB3-positive tumours, compared with SERPINB3-negative cases (Figure 4B).
miRNA 146b-5p in relation to SERPINB3 expression. miR-146b-5p levels in control HepG2 (SERPINB3−) and in HepG2 cells expressing SERPINB3 (SERPINB3+) (A). Data are represented as mean of three independent experiments and bars represent s.e. miR-146b-5p values distribution in liver tumour specimens positive or negative for SERPINB3 (B). Central bars represent mean and external bars represent s.e.
Correlation of SERPINB3 with HCC recurrence after surgical resection
In order to evaluate the potential relevance of SERPINB3 in the clinical setting, the association between high levels of SERPINB3 and time to recurrence was examined in the previously described subgroup of 67 patients.
Using 24 months as the cutoff value, all the recurrences were divided into early recurrences (⩽24 months), which are commonly regarded as a true metastasis owing to dissemination of the primary tumour cells, and late recurrences (>24 months), which are considered more likely to be de novo liver tumours occurring in a cirrhotic background (Imamura et al, 2003; Hoshida et al, 2008). As shown in Figure 5A, significantly higher levels of SERPINB3 were detected in liver tumours of patients with early recurrence. The levels of TGF-β showed a similar trend, although the similarity of median values of the two groups did not allow to reach a statistically significant difference. (Figure 5B). In agreement with these findings, Kaplan–Meier curves of time to recurrence according to SERPINB3 expression confirmed a lower recurrence-free survival in patients with high expression of the serpin, compared with patients with low or undetectable levels of the serpin in the resected tumour (Figure 6).
Correlation of molecular variables with clinical outcome. Using 24 months as the cutoff value, all the recurrences were divided into early recurrences (⩽24 months) and late recurrences (>24 months). Distribution of mRNA values of SERPINB3 (A) and of the molecular variable TGF-β1 (B), which was found significantly correlated at transcription level with SERPINB3, is reported. Central bars represent mean and external bars represent s.e. Statistical analysis was performed using unpaired t-test with Welch’s correction.
Kaplan–Meier curves of time to recurrence according to SERPINB3 expression. Patients with high (>median value) expression of SERPINB3 showed shorter recurrence-free survival, compared with patients with low or undetectable levels of SERPINB3. Statistical analysis was performed using log-rank (Mantel–Cox) test.
Molecular and clinical variables in relation to tumour recurrence
In a univariate logistic regression model, among all considered variables, including the clinical and histological data described in Table 1 and the molecular variables (high mRNA levels of SERPINB3, TGF-β and β-catenin), high SERPINB3, Child B class and BCLC B–C showed a significant association with early recurrence (Table 2). At multivariate analysis, only high SERPINB3 retained a statistically significant result, as reported in Table 2.
Discussion
The molecular mechanisms leading to HCC are still largely unknown. Hepatocarcinogenesis is a multistep phenomenon and during the progression phase activation of cellular oncogenes, overexpression of growth factors, and inactivation of tumour suppressor genes, possibly telomerase activation, may contribute to the development of the neoplastic phenotype. Alterations in gene expression patterns typical of different stages of growth, cell cycle progression, disease initiation and responses to environmental stimuli provide important clues to this complex process (Al-Sukhun and Hussain, 2003; Theodorescu, 2003).
Hepatocellular carcinoma is one of the most aggressive cancers, with poor prognosis (Sherman, 2010). Tumour resection may offer an opportunity to improve the long-term survival in early detected patients (Cillo et al, 2007); however, with current diagnostic approaches, only ∼10–20% of HCC patients are eligible for resection (Bruix et al, 2011). Early tumour recurrence is one of the major factors influencing patient survival (Marrero, 2013); however, clinical and molecular features to be used as prognostic parameters in clinical practice are still lacking. Several reports indicate that liver tumours characterised by progenitor cell signatures tend to have worse prognosis (Villanueva et al, 2010). In agreement with these findings, the proteinase inhibitor SERPINB3 has been recently described to be overexpressed in human hepatoblastoma, especially in those with more advanced stage (Turato et al, 2012). In HCC, both SERPINB3 and SERPINB4 isoforms have been identified (Pontisso et al, 2004), but no information on the relationship between the expression of these serpin isoforms and clinical outcome is available yet. Recent findings indicate that, upon target recognition, novel structural aspects of the serpin family members have been identified, leading to non-inhibitory functions, such as chaperone, tumour modulation or transport function (Silverman et al, 2010).
Regarding SERPINB3, previous studies have reported that mice transgenic for SERPINB3 showed higher liver regenerative potential compared with wild-type mice, suggesting a role of this protein in promoting cell growth and proliferation (Villano et al, 2010). We have indeed recently reported that SERPINB3 is highly expressed in hepatic stem/progenitor cell compartment of both fetal and adult livers (Villano et al, 2014). In addition, this serpin is able to induce cell proliferation in human HepG2 cells and to trigger EMT (Quarta et al, 2010), a physiological process involved in embryogenesis that has been proposed to contribute also to increased invasiveness of cancer cells and to the development of metastasis and cancer progression (Thiery, 2003). In keeping with these data, HepG2 cells genetically manipulated to overexpress SERPINB3 produced increased number of colony formation in soft agar, compared with controls (Quarta et al, 2010). The present study has been addressed to assess the relationship of SERPIRNB3 with molecular markers of the subclass of liver tumours with poor prognosis (the S1 subclass) in HCCs, where TGF-β signalling associated with Wnt target gene expression was typically upregulated (Hoshida et al, 2009). SERPINB3 was found to be overexpressed in 22% of the HCCs, and in this subset a significant increase of TGF-β1 at both transcription and protein level was observed. These features were associated with a remarkable increase of cytoplasmic expression of β-catenin, without any correlation at RNA level, suggesting a post-transcriptional modulatory effect of this serpin on this Wnt target protein. By contrast, liver tumours with low/undetectable levels of SERPINB3 showed not only low TGF-β expression, but also low β-catenin with prevalent nuclear localisation. It is worth noting that two different Wnt-related molecular classes have been recently identified: (a) the CTNNB1 class, characterised by upregulation of liver-specific Wnt-targets, nuclear β-catenin and glutamine-synthetase immunostaining, and enrichment of CTNNB1 mutation (Chiang et al, 2008), and (b) the Wnt-TGFβ class, associated with TGF-β activation, cytoplasmic β-catenin staining, vascular invasion, satellitosis and greater risk of early recurrence after surgical resection (Lachenmayer et al, 2012).
As regards TGF-β, a direct correlation of SERPINB3 on the transcription and translation of this cytokine has been reported in the present study in liver tumours. In addition, in SERPINB3-positive cases a concomitant reduction of miR-146b-5p has been detected, in keeping with the finding of an in vitro model. As this miRNA is implicated in TGF-β pathway downregulation and human hepatocarcinogenesis (Borel et al, 2012; Geraldo et al, 2012), the presented results support a multiple effect of SERPINB3 leading to increased TGF-β signalling by an increased production of this cytokine and by downregulation of miR-146b-5p miRNA that targets SMAD4, the main effector of TGF-β pathway.
Transforming growth factor-β has emerged as a major microenvironmental factor playing a role in HCC progression. In fact, in spite of its activity in the early phase of tumorigenesis as tumour suppressor, mediating growth arrest and apoptosis, in end-stage tumours it appears to take on the opposite role, where it promotes metastasis through different mechanisms (Padua and Massague, 2009). The paradoxical role of TGF-β in cancer is believed to be a consequence of the context dependence of the TGF-β signalling pathway on tumour cells (Giannelli et al, 2011; Morris et al, 2012). One of the most typical effects associated with the presence of TGF-β in tumoral cells is the enhancement of Wnt signalling by increasing the intracellular pool of β-catenin (Fischer et al, 2007).
In vitro experiments have shown previously that SERPINB3 determines morphological modifications characterised by elongated shape and decrease of desmosomal junctions associated with a remarkable accumulation of β-catenin and E-cadherin downregulation (Quarta et al, 2010). The findings of the present study support the central role of SERPINB3 in upregulation of TGF-β associated with cytoplasmic β-catenin accumulation, the molecular profile that characterises HCCs at poor prognosis. Indeed, high expression of SERPINB3 in liver tumours was significantly associated with early tumour recurrence, showing a better prognostic significance, compared with clinical and histological variables.
Accession codes
Change history
27 May 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Al-Sukhun S, Hussain M (2003) Current understanding of the biology of advanced bladder cancer. Cancer 97: 2064–2075.
Beneduce L, Castaldi F, Marino M, Quarta S, Ruvoletto M, Benvegnù L, Calabrese F, Gatta A, Pontisso P, Fassina G (2005) Squamous cell carcinoma antigen-immunoglobulin M complexes as novel biomarkers for hepatocellular carcinoma. Cancer 103 (12): 2558–2565.
Bhaumik D, Scott GK, Schokrpur S, Patil CK, Campisi J, Benz CC (2008) Expression of microRNA-146 suppresses NF-kappaB activity with reduction of metastatic potential in breast cancer cells. Oncogene 27: 5643–5647.
Biasiolo A, Tono N, Ruvoletto M, Quarta S, Turato C, Villano G, Beneduce L, Fassina G, Merkel C, Gatta A, Pontisso P (2012) IgM-linked SerpinB3 and SerpinB4 in sera of patients with chronic liver disease. PLoS One 7: e40658.
Bissell DM, Roulot D, George J (2001) Transforming growth factor beta and the liver. Hepatology 34: 859–867.
Blobe GC, Schiemann WP, Lodish HF (2000) Role of transforming growth factor beta in human disease. N Engl J Med 342: 1350–1358.
Borel F, Konstantinova P, Jansen PL (2012) Diagnostic and therapeutic potential of miRNA signatures in patients with hepatocellular carcinoma. J Hepatol 56: 1371–1383.
Bruix J, Sherman M American Association for the Study of Liver Diseases (2011) Management of hepatocellular carcinoma: an update. Hepatology 53: 1020–1022.
Bruix J, Sherman M, Llovet JM, Beaugrand M, Lencioni R, Burroughs AK, Christensen E, Pagliaro L, Colombo M, Rodes J EASL Panel of Experts on HCC (2001) Clinical management of hepatocellular carcinoma. Conclusions of the Barcelona-2000 EASL conference. European Association for the Study of the Liver. J Hepatol 35: 421–430.
Bruix J, Sherman M Practice Guidelines Committee, American Association for the Study of Liver Diseases (2005) Management of hepatocellular carcinoma. Hepatology 42: 1208–1236.
Cataltepe S, Gornstein ER, Schick C, Kamachi Y, Chatson K, Fries J, Silverman GA, Upton MP (2000) Co-expression of the squamous cell carcinoma antigens 1 and 2 in normal adult human tissues and squamous cell carcinomas. J Histochem Cytochem 48: 113–122.
Chiang DY, Villanueva A, Hoshida Y, Peix J, Newell P, Minguez B, LeBlanc AC, Donovan DJ, Thung SN, Sole M, Tovar V, Alsinet C, Ramos AH, Barretina J, Roayaie S, Schwartz M, Waxman S, Bruix J, Mazzaferro V, Ligon AH, Najfeld V, Friedman SL, Sellers WR, Meyerson M, Llovet JM (2008) Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res 68: 6779–6788.
Cillo U, Vitale A, Bassanello M, Boccagni P, Brolese A, Zanus G, Burra P, Fagiuoli S, Farinati F, Rugge M, D’Amico DF (2004) Liver transplantation for the treatment of moderately or well-differentiated hepatocellular carcinoma. Ann Surg 239: 150–159.
Cillo U, Vitale A, Brolese A, Zanus G, Neri D, Valmasoni M, Bonsignore P, Grigoletto F, Burra P, Farinati F, D’Amico DF (2007) Partial hepatectomy as first-line treatment for patients with hepatocellular carcinoma. J Surg Oncol 95: 213–220.
El-Serag HB, Rudolph KL (2007) Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology 132: 2557–2576.
Fischer AN, Fuchs E, Mikula M, Huber H, Beug H, Mikulits W (2007) PDGF essentially links TGF-beta signaling to nuclear beta-catenin accumulation in hepatocellular carcinoma progression. Oncogene 26: 3395–3405.
Geraldo MV, Fuziwara CS, Friguglieti CU, Costa RB, Kulcsar MA, Yamashita AS, Kimura ET (2012) MicroRNAs miR-146-5p and let-7f as prognostic tools for aggressive papillary thyroid carcinoma: a case report. Arq Bras Endocrinol Metabol 56: 552–557.
Giannelli G, Mazzocca A, Fransvea E, Lahn M, Antonaci S (2011) Inhibiting TGF-beta signaling in hepatocellular carcinoma. Biochim Biophys Acta 1815: 214–223.
Gramantieri L, Ferracin M, Fornari F, Veronese A, Sabbioni S, Liu CG, Calin GA, Giovannini C, Ferrazzi E, Grazi GL, Croce CM, Bolondi L, Negrini M (2007) Cyclin G1 is a target of miR-122a, a microRNA frequently down-regulated in human hepatocellular carcinoma. Cancer Res 67: 6092–6099.
Guido M, Roskams T, Pontisso P, Fassan M, Thung SN, Giacomelli L, Sergio A, Farinati F, Cillo U, Rugge M (2008) Squamous cell carcinoma antigen in human liver carcinogenesis. J Clin Pathol 61: 445–447.
Hoshida Y (2010) Nearest template prediction: a single-sample-based flexible class prediction with confidence assessment. PLoS One 5: e15543.
Hoshida Y, Nijman SM, Kobayashi M, Chan JA, Brunet JP, Chiang DY, Villanueva A, Newell P, Ikeda K, Hashimoto M, Watanabe G, Gabriel S, Friedman SL, Kumada H, Llovet JM, Golub TR (2009) Integrative transcriptome analysis reveals common molecular subclasses of human hepatocellular carcinoma. Cancer Res 69: 7385–7392.
Hoshida Y, Toffanin S, Lachenmayer A, Villanueva A, Minguez B, Llovet JM (2010) Molecular classification and novel targets in hepatocellular carcinoma: recent advancements. Semin Liver Dis 30: 35–51.
Hoshida Y, Villanueva A, Kobayashi M, Peix J, Chiang DY, Camargo A, Gupta S, Moore J, Wrobel MJ, Lerner J, Reich M, Chan JA, Glickman JN, Ikeda K, Hashimoto M, Watanabe G, Daidone MG, Roayaie S, Schwartz M, Thung S, Salvesen HB, Gabriel S, Mazzaferro V, Bruix J, Friedman SL, Kumada H, Llovet JM, Golub TR (2008) Gene expression in fixed tissues and outcome in hepatocellular carcinoma. N Engl J Med 359: 1995–2004.
Imamura H, Matsuyama Y, Tanaka E, Ohkubo T, Hasegawa K, Miyagawa S, Sugawara Y, Minagawa M, Takayama T, Kawasaki S, Makuuchi M (2003) Risk factors contributing to early and late phase intrahepatic recurrence of hepatocellular carcinoma after hepatectomy. J Hepatol 38: 200–207.
Lachenmayer A, Alsinet C, Savic R, Cabellos L, Toffanin S, Hoshida Y, Villanueva A, Minguez B, Newell P, Tsai HW, Barretina J, Thung S, Ward SC, Bruix J, Mazzaferro V, Schwartz M, Friedman SL, Llovet JM (2012) Wnt-pathway activation in two molecular classes of hepatocellular carcinoma and experimental modulation by sorafenib. Clin Cancer Res 18: 4997–5007.
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25: 402–408.
Llovet JM, Bruix J (2008) Molecular targeted therapies in hepatocellular carcinoma. Hepatology 48: 1312–1327.
Llovet JM, Burroughs A, Bruix J (2003) Hepatocellular carcinoma. Lancet 362: 1907–1917.
Man YG, Fu SW, Liu AJ, Stojadinovic A, Izadjoo MJ, Chen L, Gardner WA (2011) Aberrant expression of chromogranin A, miR-146a, and miR-146b-5p in prostate structures with focally disrupted basal cell layers: an early sign of invasion and hormone-refractory cancer? Cancer Genomics Proteomics 8: 235–244.
Marrero JA (2013) Current Treatment Approaches in HCC. Clin Adv Hematol Oncol 11 (Suppl 5): 15–18.
Morris SM, Baek JY, Koszarek A, Kanngurn S, Knoblaugh SE, Grady WM (2012) Transforming growth factor-beta signaling promotes hepatocarcinogenesis induced by p53 loss. Hepatology 55: 121–131.
Padua D, Massague J (2009) Roles of TGFbeta in metastasis. Cell Res 19: 89–102.
Pasche B (2001) Role of transforming growth factor beta in cancer. J Cell Physiol 186: 153–168.
Pontisso P, Calabrese F, Benvegnù L, Lise M, Belluco C, Ruvoletto MG, Marino M, Valente M, Nitti D, Gatta A, Fassina G (2004) Overexpression of squamous cell carcinoma antigen variants in hepatocellular carcinoma. Br J Cancer 90: 833–837.
Quarta S, Vidalino L, Turato C, Ruvoletto M, Calabrese F, Valente M, Cannito S, Fassina G, Parola M, Gatta A, Pontisso P (2010) SERPINB3 induces epithelial-mesenchymal transition. J Pathol 221: 343–356.
Rooke HM, Crosier KE (2001) The smad proteins and TGFbeta signalling: uncovering a pathway critical in cancer. Pathology 33: 73–84.
Sherman M (2010) Epidemiology of hepatocellular carcinoma. Oncology 78 (Suppl 1): 7–10.
Shi Y, Massague J (2003) Mechanisms of TGF-beta signaling from cell membrane to the nucleus. Cell 113: 685–700.
Shirota Y, Kaneko S, Honda M, Kawai HF, Kobayashi K (2001) Identification of differentially expressed genes in hepatocellular carcinoma with cDNA microarrays. Hepatology 33: 832–840.
Silverman GA, Whisstock JC, Bottomley SP, Huntington JA, Kaiserman D, Luke CJ, Pak SC, Reichhart JM, Bird PI (2010) Serpins flex their muscle: I. Putting the clamps on proteolysis in diverse biological systems. J Biol Chem 285: 24299–24305.
Takeshima N, Suminami Y, Takeda O, Abe H, Okuno N, Kato H (1992) Expression of mRNA of SCC antigen in squamous cells. Tumour Biol 13: 338–342.
Theodorescu D (2003) Phosphoinositide signaling in urological disease. J Urol 169: 2394–2396.
Thiery JP (2003) Epithelial-mesenchymal transitions in development and pathologies. Curr Opin Cell Biol 15: 740–746.
Trerotoli P, Fransvea E, Angelotti U, Antonaci G, Lupo L, Mazzocca A, Mangia A, Antonaci S, Giannelli G (2009) Tissue expression of Squamous Cellular Carcinoma Antigen (SCCA) is inversely correlated to tumor size in HCC. Mol Cancer 8: 29.
Turato C, Buendia MA, Fabre M, Redon MJ, Branchereau S, Quarta S, Ruvoletto M, Perilongo G, Grotzer MA, Gatta A, Pontisso P (2012) Over-expression of SERPINB3 in hepatoblastoma: a possible insight into the genesis of this tumour? Eur J Cancer 48: 1219–1226.
Turato C, Calabrese F, Biasiolo A, Quarta S, Ruvoletto M, Tono N, Paccagnella D, Fassina G, Merkel C, Harrison TJ, Gatta A, Pontisso P (2010) SERPINB3 modulates TGF-beta expression in chronic liver disease. Lab Invest 90 (7): 1016–1023.
Vidalino L, Doria A, Quarta S, Zen M, Gatta A, Pontisso P (2009) SERPINB3, apoptosis and autoimmunity. Autoimmun Rev 9: 108–112.
Villano G, Quarta S, Ruvoletto MG, Turato C, Vidalino L, Biasiolo A, Tono N, Lunardi F, Calabrese F, Dall’olmo L, Dedja A, Fassina G, Gatta A, Pontisso P (2010) Role of squamous cell carcinoma antigen-1 on liver cells after partial hepatectomy in transgenic mice. Int J Mol Med 25: 137–143.
Villano G, Turato C, Quarta S, Ruvoletto M, Ciscato F, Terrin L, Semeraro R, Paternostro C, Parola M, Alvaro D, Bernardi P, Gatta A, Pontisso P (2014) Hepatic progenitor cells express SerpinB3. BMC Cell Biol 15: 5.
Villanueva A, Hoshida Y, Toffanin S, Lachenmayer A, Alsinet C, Savic R, Cornella H, Llovet JM (2010) New strategies in hepatocellular carcinoma: genomic prognostic markers. Clin Cancer Res 16: 4688–4694.
Villanueva A, Newell P, Chiang DY, Friedman SL, Llovet JM (2007) Genomics and signaling pathways in hepatocellular carcinoma. Semin Liver Dis 27: 55–76.
Xu XR, Huang J, Xu ZG, Qian BZ, Zhu ZD, Yan Q, Cai T, Zhang X, Xiao HS, Qu J, Liu F, Huang QH, Cheng ZH, Li NG, Du JJ, Hu W, Shen KT, Lu G, Fu G, Zhong M, Xu SH, Gu WY, Huang W, Zhao XT, Hu GX, Gu JR, Chen Z, Han ZG (2001) Insight into hepatocellular carcinogenesis at transcriptome level by comparing gene expression profiles of hepatocellular carcinoma with those of corresponding noncancerous liver. Proc Natl Acad Sci USA 98: 15089–15094.
Zhang DY, Friedman SL (2012) Fibrosis-dependent mechanisms of hepatocarcinogenesis. Hepatology 56: 769–775.
Zhong H, Wang HR, Yang S, Zhong JH, Wang T, Wang C, Chen FY (2010) Targeting Smad4 links microRNA-146a to the TGF-beta pathway during retinoid acid induction in acute promyelocytic leukemia cell line. Int J Hematol 92: 129–135.
Zucman-Rossi J, Benhamouche S, Godard C, Boyault S, Grimber G, Balabaud C, Cunha AS, Bioulac-Sage P, Perret C (2007) Differential effects of inactivated Axin1 and activated beta-catenin mutations in human hepatocellular carcinomas. Oncogene 26: 774–780.
Acknowledgements
This work was supported in part by the following Research Grants: National Ministry of Education, University and Research (FIRB Project Prot. RBLA03S4SP_005), University of Padova (Project No CPDA110795) and Associazione Italiana per la Ricerca sul Cancro (AIRC Project No. 10235).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Turato, C., Vitale, A., Fasolato, S. et al. SERPINB3 is associated with TGF-β1 and cytoplasmic β-catenin expression in hepatocellular carcinomas with poor prognosis. Br J Cancer 110, 2708–2715 (2014). https://doi.org/10.1038/bjc.2014.246
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.246
Keywords
This article is cited by
-
Serum Squamous Cell Carcinoma Antigen-Immunoglobulin M complex levels predict survival in patients with cirrhosis
Scientific Reports (2019)
-
SerpinB3 Promotes Pro-fibrogenic Responses in Activated Hepatic Stellate Cells
Scientific Reports (2017)
-
SerpinB3 and Yap Interplay Increases Myc Oncogenic Activity
Scientific Reports (2015)
-
The effect of recombinant lentiviral vector encoding miR-145 on human esophageal cancer cells
Tumor Biology (2015)