Abstract
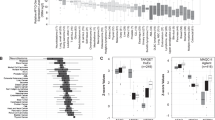
Deregulated expression of myc proto-oncogenes is implicated in several human neoplasias. We analysed the expression of c-myc, N-myc, L-myc, max and RB1 mRNAs in a panel of human gliomas and glioma cell lines and compared the findings with normal neural cells. The max and RB1 genes were included in the study because their protein products can interact with the Myc proteins, being thus putative modulators of Myc activity. Several gliomas contained c/L-myc mRNAs at levels higher than those in fetal brain, L-myc predominantly in grade II/III and c-myc in grade III gliomas. High-level N-myc expression was detected. In one small-cell glioblastoma and lower levels in five other gliomas. In contrast, glioma cell lines totally lacked N/L-myc expression. The in situ hybridisations revealed mutually exclusive topographic distribution of myc and glial fibrillary acidic protein (GFAP) mRNAs, and a lack of correlation between myc expression and proliferative activity, max and RB1 mRNAs were detected in most tumours and cell lines. The glioma cells displayed interesting alternative splicing patterns of max mRNAs encoding Max proteins which either suppress (Max) or augment (delta Max) the transforming activity of Myc. We conclude that (1) glioma cells in vivo may coexpress several myc genes, thus resembling fetal neural cells; but (2) cultured glioma cells expression only c-myc; (3) myc, max and RB1 are regulated independently in glioma cells; and (4) alternative processing of max mRNA in some glioma cells results in delta Max encoding mRNAs not seen in normal fetal brain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hirvonen, H., Salonen, R., Sandberg, M. et al. Differential expression of myc, max and RB1 genes in human gliomas and glioma cell lines. Br J Cancer 69, 16–25 (1994). https://doi.org/10.1038/bjc.1994.3
Issue Date:
DOI: https://doi.org/10.1038/bjc.1994.3
This article is cited by
-
Histone demethylase KDM4C controls tumorigenesis of glioblastoma by epigenetically regulating p53 and c-Myc
Cell Death & Disease (2021)
-
Molecular mechanisms underlying gliomas and glioblastoma pathogenesis revealed by bioinformatics analysis of microarray data
Medical Oncology (2017)
-
A mutant oncolytic adenovirus targeting the Rb pathway produces anti-glioma effect in vivo
Oncogene (2000)
-
Overexpression of E2F‐1 in glioma triggers apoptosis and suppresses tumor growth in vitro and in vivo
Nature Medicine (1998)
-
Genomic alterations of human gliomas detected by restriction landmark genomic scanning
Brain Tumor Pathology (1997)