Abstract
A dinucleotide deletion in human ubiquitin (Ub) B messenger RNA leads to formation of polyubiquitin (UbB)+1, which has been implicated in neuronal cell death in Alzheimer's and other neurodegenerative diseases. Previous studies demonstrate that UbB+1 protein causes proteasome dysfunction. However, the molecular mechanism of UbB+1-mediated neuronal degeneration remains unknown. We now report that UbB+1 causes neuritic beading, impairment of mitochondrial movements, mitochondrial stress and neuronal degeneration in primary neurons. Transfection of UbB+1 induced a buildup of mitochondria in neurites and dysregulation of mitochondrial motor proteins, in particular, through detachment of P74, the dynein intermediate chain, from mitochondria and decreased mitochondria–microtubule interactions. Altered distribution of mitochondria was associated with activation of both the mitochondrial stress and p53 cell death pathways. These results support the hypothesis that neuritic clogging of mitochondria by UbB+1 triggers a cascade of events characterized by local activation of mitochondrial stress followed by global cell death. Furthermore, UbB+1 small interfering RNA efficiently blocked expression of UbB+1 protein, attenuated neuritic beading and preserved cellular morphology, suggesting a potential neuroprotective strategy for certain neurodegenerative disorders.
Similar content being viewed by others
Main
Ubiquitin (Ub) is a highly conserved 76-amino acid protein that facilitates 26S proteasome-mediated proteolysis in cells. Ub covalently tags a wide variety of proteins by the sequential actions of an E1-activating enzyme, an E2-conjugating enzyme that transfers Ub to a target protein, and an E3 ligase that ubiquitinates the target protein.1 Polyubiquitinated proteins are then degraded through the 26S proteasome complex. To generate an efficient proteasome-targeting signal, Ub is first linked to a given substrate by formation of an isopeptide bond between the C-terminal glycine of Ub (G76) and an internal lysine (K) residue of the substrate. Polyubiquitination occurs through the formation of K48-G76 isopeptide bonds between successive Ub molecules. The Ub-proteasome pathway (UPP) is the major extralysosomal pathway responsible for protein degradation in eukaryotes.1 Converging lines of evidence have shown that a variety of intracellular regulatory molecules as well as misfolded proteins are substrates of the UPP. The UPP thus plays a critical role in vital cellular processes such as the cell cycle and cell division, immune and inflammatory responses, cell differentiation, DNA damage repair and cell death.1, 2
Abnormal UPP function has been implicated in many pathological conditions, including human neurodegenerative diseases.2, 3 Insoluble ubiquitinated protein aggregates are present in the pathological hallmarks of Alzheimer's disease (AD), particularly neuritic plaques and neurofibrillary tangles. Failure of UPP function has been linked to beta-amyloid (Aβ) toxicity in the central nervous system (CNS).4 UPP dysfunction is also caused by the Ub mutant, UbB+1, which accumulates in AD and other tauopathies.5, 6, 7 UbB+1 is generated by a non-DNA-encoded dinucleotide deletion in the first coding unit of human polyubiquitin B (UbB) transcripts. This leads to a +1 frameshift mutation resulting in replacement of the C-terminal G76 of Ub with a 20-residue extension.7 The altered C-terminus abrogates the ability of UbB+1 to tag other substrates without affecting its polyubiquitination.8, 9 Polyubiquitinated UbB+1 is refractory to deubiquitination and impairs proteasome function.8 Accordingly, induction of UbB+1 results in UPP dysfunction and neuronal cell death.9, 10 However, the underlying molecular basis of UbB+1 formation and its role in neuronal degeneration remain unclear. Here, we demonstrate the novel finding that UbB+1 causes clogging of mitochondria with beading in neurites, followed by mitochondrial stress and neuronal cell death. Biochemical and ultrastructural analyses suggest that neuritic beading might result from dysregulation of mitochondrial transport proteins and impaired mitochondria–microtubule (MT) interactions. In addition, small interfering RNA (siRNA)-mediated suppression of UbB+1 attenuates the development of neuritic beading and preserves neuronal morphology. Mitochondrial stress induced by a mutant form of Ub may therefore be a novel mechanism contributing to neuronal loss in AD and other neurodegenerative diseases.
Results
UbB+1 causes neuritic beading and mitochondrial buildup in neuronal processes
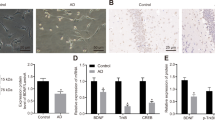
To investigate the pathophysiological effects of UbB+1, primary neurons were transfected with plasmids expressing either enhanced green fluorescent protein (EGFP)–UbB+1 or EGFP–Ub fusion proteins, or EGFP alone. Fluorescence microscopy revealed a pattern consistent with neuritic beading in the processes of most cells by 16 h after EGFP–UbB+1 transfection (Figure 1a, panel 1). The majority of cells transfected with either EGFP-Ub or EGFP showed normal morphology without development of neuritic beading (Figure 1a, panels 2 and 3 in upper row). Compared to EGFP-alone, cells transfected with EGFP–Ub exhibited a heterogeneous pattern of fluorescence consistent with the non-uniform distribution of endogenous Ub and Ub-conjugates.11 Quantification of the results revealed a statistically significant increase in the number of cells containing neuritic beads with time as well as the number of beads per beading cell following pEGFP–UbB+1 transfection compared with controls (Figure 1a, bottom row). There were no significant differences between results obtained in rat and mouse neurons (data not shown).
UbB+1 causes neuritic beading and mitochondrial buildup in neuronal processes. (a) Fluorescence images of rat cortical neurons 16 h after transfection with pEGFP–UbB+1, pEGFP-Ub or pEGFP alone. Quantification shows a significant increase in the percentage of beading cells and the number of beads per cell following transfection with pEGFP-UbB+1. * and Δ indicate significant differences (P's *<0.05, **<0.01, N=4; Δ<0.05, N=20, as per ANOVA and Fisher's PLSD). Bars depict mean±S.E. (b) Neurons were co-transfected with pEGFP–UbB+1 and pDsRed2-Mito. Confocal microscopy shows colocalization of the mitochondrial marker in EGFP–UbB+1-associated neuritic beads. (c) A transmission electron micrograph reveals beading structures in an axon from a pEGFP–UbB+1-transfected cell. (d and e) At higher magnification, the boxed areas in (c) show the presence of microtubules (MT), mitochondria or mitochondria-like structures (Mi), spheroids (broad arrows in e) and smaller vesicular bodies. (f) A neuritic bead containing mitochondria in clumps, MT and vesicular bodies. An adjacent process is demarcated by delimiting membranes (DM, arrowheads). Scale bar=50 μm (a), 20 μm (insets a), 150 μm (inset b) shown at higher magnification in other panels (30 μm), 1 μm (c) and 0.4 μm in (d–f)
As neuritic beading has been associated with jamming of intracellular organelles,12 we next examined the nature of the neuritic beads by co-transfecting primary neurons with pEGFP–UbB+1 and either a mitochondrial (pDsRed2–Mito), endoplasmic reticulum (pEYFP–ER) or Golgi (pEYFP–Golgi)-targeting reporter. Analysis by confocal laser optical dissection revealed that almost all of the EGFP–UbB+1-positive beads contained mitochondria (Figure 1b), whereas endoplasmic reticulum and Golgi bodies were absent (data not shown). To further examine the structure and composition of the neuritic beads, neurons were analyzed by transmission electron microscopy 16 h after pEGFP–UbB+1 transfection. Consistent with our previous observations mitochondria, or mitochondria-like structures, were the primary components in beads. In addition, the beads contained MTs, vesicular bodies and smaller structures that were possibly aggresomes and other organelles (Figure 1c–e).
Neuritic beading is correlated with mitochondrial motor dysregulation and altered mitochondria–MT interactions
Mitochondria are bidirectionally transported between the neuronal soma and terminals by the axoplasmic transport machinery.13, 14 As the accumulation of mitochondria in neuritic beads could potentially result from impaired mitochondrial transport, we evaluated the effects of UbB+1 on mitochondrial movements in neurites and expression of mitochondrial molecular motors. Neurons were co-transfected with the mitochondrial reporter, pDsRed2-Mito and either pEGFP, pEGFP-Ub or pEGFP-UbB+1, and the movements of labeled aggregates containing mitochondria were analyzed by time-lapse photomicrography (Figure 2a). Quantification of the results revealed a striking decrease in the number of mitochondria-containing aggregates that moved retrogradely in EGFP–UbB+1- compared with either EGFP–Ub- or EGFP-expressing cells (Figure 2b). In contrast, there was no significantly statistical difference in the anterograde movement of aggregates from EGFP–UbB+1-expressing cells compared with control EGFP–Ub- or EGFP-expressing cells (Figure 2b).
Time-lapse analysis of mitochondrial movements. (a) Comparison of photomicrographs at two different times after transfection demonstrates retrograde (arrowheads shifted up) and anterograde (arrows shifted down) motions of mitochondria-containing aggregates. Scale bar=10 μm. (b) Quantification of mitochondrial aggregate motions in neuronal processes. The number of labeled mitochondrial aggregates that moved retrogradely was significantly reduced in EGFP–UbB+1-transfected cells (P's *< 0.01, **<0.005, N=25, as per ANOVA and Fisher's PLSD). No significant difference was detected between the number of mitochondrial aggregates that moved retrogradely in cells expressing EGFP-Ub and Ub alone (P's Δ> 0.45, N=25, as per ANOVA); no statistically significant change was detected in the number of mitochondrial aggregates that moved anterogradely in EGFP–UbB+1-expressing cells or in cells expressing either EGFP–Ub or EGFP alone (P's ∇>0.50, N=25, as per ANOVA). Bars depict mean±S.E.
Mitochondria are transported as cargo along neuronal processes through direct interactions between the motor molecules, KIF1B kinesin and dynein, and the MT cytoskeletal system.15, 16 Therefore, we examined the overall expression levels of KIF1B, a member of the kinesin superfamily that is a major motor for anterograde transport of mitochondria,15 and dynein intermediate chain P74, which regulates mitochondrial retrograde transport, in transfected neurons. Western blot analyses using a panel of antibodies revealed that EGFP, EGFP–Ub and EGFP–UbB+1 expression increased between 8 and 30 h post-transfection (Figure 3a, row 1). UbB+1 immunoreactivity was confirmed with a specific antibody against UbB+1 (Figure 3a, row 2). Whereas there was no statistically significant change in the abundance of P74 in whole-cell lysates, a significant decrease in mitochondria-bound P74 was detected 16 h after UbB+1 transfection, along with a corresponding increase in cytosolic dynein (Figure 3a, rows 4, 6 and 9; Figure 3b). Both whole-cell and mitochondria-associated levels of KIF1B remained unchanged (Figure 3a, rows 3 & 7). There were no changes in motor protein expression in cells transfected with either EGFP–Ub or EGFP alone.
Biochemical analyses of mitochondrial motors. (a) Whole-cell lysates, mitochondrial (mito) and cytosolic (cyto) fractions were prepared from primary neurons transfected with either pEGFP, pEGFP–UbB+1 or pEGFP–Ub. Western blotting was performed with a panel of antibodies at the indicated times. (b) The relative changes in dynein P74 abundance in both mitochondrial and cytosol fractions are shown based on the initial levels in EGFP-transfected cells. Bars represent the mean±S.E. from four independent experiments (*P<0.05, as per Student's t-test). (c) Fluorescence microscopy revealed neuritic beading of mitochondria (red) in primary neurons co-transfected with pDsRed2-Mito and UbB+1 (green) in pcDNA3.1. (d). Whole–cell lysates (whole cell), mitochondrial (mito) and cytosolic (cyto) fractions were prepared from neurons 24 h after transfection with empty pcDNA3.1 (ctl) or with Ub or UbB+1 cDNA cassette. Western blotting was performed with indicated antibodies. (e) Cell-free reconstitution assay. Isolated mitochondria were incubated with increasing amounts of Ub species or bovine serum album (ctl), followed by P74 Western blotting. * indicates a significant difference of the group incubated with either 2.0 μg UbB+1 or Ub5+1 relative to the control (P's<0.05, N=4, as per ANOVA and Fisher's PLSD)
To rule out any possible effects of EGFP in the fusion proteins, primary neurons were co-transfected with DsRed2-Mito and a pcDNA3.1 plasmid expressing nontagged Ub or UbB+1. Dual fluorescence of DsRed2–Mito and UbB+1 immunoreactivity revealed similar beading pattern of mitochondria in neurites in cells expressing UbB+1 (Figure 3c). Western blots also demonstrated a similar decrease in abundance of mitochondria-associated P74 with a corresponding increase in the level of cytoplasmic P74 in neurons transfected with UbB+1, whereas empty vector and Ub controls showed no change (Figure 3d).
As mitochondrial transport is facilitated by direct interactions with molecular motors, we investigated whether UbB+1 protein interfered with binding of P74 to mitochondria cargo. In a cell-free reconstitution assay, isolated mitochondrial fractions from untreated neurons were incubated with wild-type Ub, UbB+1 or Ub5+1, that is, UbB+1 conjugated with a tetra-polyUb chain through K48. Increasing amounts of UbB+1 or Ub5+1 resulted in a significant decrease in mitochondria-associated P74 and a corresponding increase in the buffer fraction (Figure 3e). These results indicate that both free and polyubiquitinated forms of UbB+1 protein may interfere with P74 binding to mitochondria and result in detachment of P74 from its cargo.
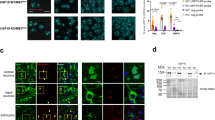
Although mechanisms of mitochondrial transport are not well understood, studies have revealed that motors actively move their cargo along MTs.17 We speculated that detachment of P74 from mitochondria may result in the dissociation of mitochondria from MTs. The effect of UbB+1 on the spatial relationship between mitochondria and MTs was investigated by an ultrastructural analysis of longitudinal sections through neurites. In EGFP- or EGFP–Ub-transfected neurons, the majority of mitochondria were flanked by MTs on two sides, that is, ‘MT-associated’. In contrast, most of the mitochondria in EGFP–UbB+1-transfected neurons were either not adjacent to or contiguous with MTs on only one side. Quantification of these results demonstrated a dramatic reduction of MT-associated mitochondria in EGFP–UbB+1-transfected cells (Figures 4a, b).
Ultrastructural analysis of the spatial relationships between mitochondria and MTs. (a) Electron micrographs taken from transfected neurons. The first two panels show mitochondria flanked by MTs (arrows) on two sides (i.e., MT-associated), whereas the third panel shows a mitochondrion with MTs on only one side. (b) There was a significant decrease in MT-associated mitochondria relative to the total number of mitochondria-like structures in EGFP–UbB+1-transfected cells. (*P's<0.01, N=4, as per ANOVA and Fisher's PLSD). Bars depict mean±S.E.
Neuritic beading is associated with mitochondrial stress
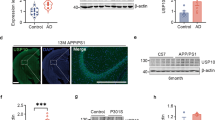
To explore the downstream effects of UbB+1 expression in more detail, several indicators of mitochondrial stress were employed. The loss of mitochondrial membrane potential was evaluated by staining with tetramethyl rhodamine methyl ester (TMRM) and mitochondrial superoxide production was evaluated by MitoSOX-Red (MSR). TMRM was observed to stain almost all the mitochondria in EGFP-transfected cells (data not shown) and EGFP–Ub-transfected cells (Figure 5a, second row). In contrast, cells expressing EGFP–UbB+1 exhibited reduced TMRM staining in both neuronal processes and the cell somata (Figure 5a, first row). Specifically, reduced TMRM fluorescence in neurites preceded the more global loss of TMRM fluorescence over the entire cell (Figure 5b). Consistent with the loss of mitochondrial membrane potential, MSR signal colocalized with EGFP–UbB+1 in neuritic beads (Figure 5c). Cells expressing EGFP–Ub were MSR-negative (Figure 5c). Quantification of these results demonstrated a progressive increase in the number of MSR-positive cells with time after EGFP–UbB+1-transfection (Figure 5d). Importantly, the loss of mitochondrial membrane potential was associated with the release of cytochrome c into the cytosol (Figure 5e). Local elevations in cytochrome c might then act as a trigger for caspase-3 activation in the vicinity of UbB+1-positive neuritic beads (Figure 5f).
Neuritic beading is associated with mitochondrial stress. (a) Neurons were stained with TMRM after either EGFP–UbB+1 or EGFP–Ub transfection. TMRM fluorescence (red) is present in both neurites (arrows) and cell soma (arrowheads) after EGFP–Ub transfection, but only in the cell soma after EGFP–UbB+1 transfection. (b) The number of cells with reduced TMRM signal in neurites (left) or whole cell (right) relative to EGFP-positive cells following transfection. P's<0.01 for each time point, N=4. (c) Mitochondrial superoxides were stained by MSR in EGFP–UbB+1-associated neuritic beads (arrows), but not in EGFP–Ub-transfected cells. (d) The number of MSR- relative to EGFP-positive cells following plasmid transfection. * Indicates significant differences (P’s *<0.05, **<0.01, N=4, as per ANOVA and Fisher's PLSD). Bars depict mean±S.E. (e) Cytochrome c abundance in cytosol following plasmid transfection. GAPDH was used as loading control. (f) Caspase-3 activation (green) and UbB+1-positive beads (red) are colocalized in some processes (merged, arrow). Scale bar=40 μm (inset f), 10 μm in other panels of (f), and 15 μm in (a and c)
UbB+1-induced proteasomal dysfunction is associated with activation of p53 and neuronal cell death
Although our results suggested that adverse effects on mitochondrial movements play a critical role in UbB+1-mediated neuronal death, we were interested in exploring other potential actions of UbB+1 in neurons. The extended C-terminus resulting from the dinucleotide deletion in UbB+1 messenger RNA leads to impaired UPP activity and protein stabilization.9 Therefore, we evaluated UPP function by examining the accumulation of polyUb conjugates in transfected neurons. Western blotting revealed a significant increase in abundance of polyUb conjugates in cells expressing EGFP–UbB+1 compared with either EGFP or EGFP–Ub (Figure 6a). In addition, a marked accumulation of the cell death protein, p53, a major UPP substrate, was also detected by 16 h after pEGFP–UbB+1 transfection (Figure 6b). Immunocytochemistry revealed a predominantly nuclear accumulation of p53 in beading neurons (Figure 6c, 1st row). However, we could not identify p53 associated with the mitochondrial fractions (data not shown). We previously showed that p53 accumulates, in part, due to decreased availability of free Ub.18, 19 In this regard, free Ub was unchanged in UbB+1-transfected cells (Figure 6b), suggesting that proteasome inhibition might contribute to the accumulation of p53 in this model.5 As p53 is a major transcriptional regulator of apoptosis, it was important to evaluate the transcriptional activity of stabilized p53. Studies with the p53 reporter, pp53-TA-DsRed2, revealed a significant and progressive increase in p53 transactivity (Figure 6c, 2nd row; Figure 6d) and cell death as detected by TUNEL (Figure 6c, 3rd row; Figure 6d) with time after transfection. In contrast, there were relatively few MSR- or TUNEL-positive cells following transfection with either EGFP–Ub or EGFP alone (data not shown). Additionally, the role of p53 in UbB+1-induced neuronal cell death was investigated by transfecting primary neurons from p53-null mice with EGFP–UbB+1, followed by either MSR or TUNEL staining. Although MSR staining was present (Figure 6c, 4th row), there was a striking decrease in the number of MSR-positive neurons in the absence of p53 (i.e., compare Figure 6e (23±4.0%) with Figure 5d (83.7±5.5%) at 48 h, P<0.01). Similarly, p53-deficiency resulted in a significant reduction in TUNEL-positive neurons (i.e., compare Figure 6e (19.3±3.9%) with Figure 6d (47.3±4.9%) at 48 h, P<0.05). To evaluate the importance of caspase-3 activation, primary neurons were transfected with EGFP–UbB+1 with or without the caspase-3-specific inhibitor, DEVD-CHO, followed by TUNEL staining. The results showed that DEVD-CHO significantly attenuated cell death due to EGFP–UbB+1 (Figure 6d).
Accumulation of polyUb conjugates, p53 activation and neuronal cell death. (a) Whole-cell lysates were prepared from primary neurons transfected with EGFP or EGFP fusion constructs 24 h after transfection. Western blotting using a Ub antibody showed increased abundance of polyUb conjugates in UbB+1-transfected cells. Lysates from neurons treated with MG115 (5 μM, 24 h) were used as a positive control. Bottom: changes in abundance of total polyUb conjugate ladders are expressed relative to those in EGFP control cells. The data depict the mean number of pixels±S.E.M from four independent experiments. (b) p53 and free Ub abundance at indicated times following plasmid transfection. All the lanes shown in each panel were from the same gel. β-Actin was used as loading control. (c) Representative examples in EGFP–UbB+1-transfected neurons of: nuclear p53 immunoreactivity (arrow, top row); activated p53 following co-transfection with pp53-TA-DsRed2 (second row); positive nuclear TUNEL staining (red, third row) with neuritic beads; positive MSR staining (red, fourth row) in p53-null neurons. Scale bar=60 μm in second row and 30 μm in other panels. (d) The number of TUNEL- or p53 reporter-positive cells relative to EGFP-positive cells after EGFP–UbB+1 transfection. Bars are mean±S.E. TUNEL studies were performed in the presence or absence of DEVD-CHO. (e) p53-null neurons were transfected with EGFP–UbB+1, and the number of MSR- or TUNEL-positive cells relative to EGFP-positive cells was counted. Bars are mean±S.E.
siRNA-mediated suppression of UbB+1 is neuroprotective
siRNA is a powerful tool that is commonly used for selective suppression of gene expression including allele-specific silencing.20 Therefore, we explored the efficacy of siRNA in reversing UbB+1-mediated toxicity. A sequence-specific UbB+1 siRNA construct was first validated in N2A cells by Western blot analysis (see Supplementary Data). This sequence was then incorporated into a vector co-expressing a short interfering hairpin RNA (shRNA) that targeted UbB+1 (pRFP.UbB+1sh) and red fluorescent protein (RFP), DsRed2, to monitor transfection (Figure 7a). An shRNA coding for an irrelevant RNA sequence (pRFP.Controlsh) was used as a control. Studies performed with both N2A cells (see Supplementary Data) and primary neurons showed that pRFP.UbB+1sh effectively and selectively suppressed EGFP-UbB+1 expression (Figure 7b, right panels). In contrast, pRFP.Controlsh had no effect on EGFP–UbB+1 expression (Figure 7b, left panels) and pRFP.UbB+1sh had no effect on UbB expression (see Supplementary Data). Quantitation revealed that there was a significant decrease in the number of neurons expressing UbB+1 after co-transfection with UbB+1 shRNA compared with the control shRNA (Figure 7c). Importantly, blocking UbB+1 expression in primary neurons prevented the development of neuritic beading and preserved neuronal morphology (Figure 7c, right panels).
shRNA-mediated suppression of UbB+1. (a) Schematic of RFP/DsRed2.U6.silencer vector encoding a specific shRNA directed by murine U6 promoter and a CMV-directed fluorescent reporter. UbB+1 shRNA targets UbB+1 mRNA, which differs by only two nucleotides (highlighted in red) from UbB mRNA. (b) Mouse cortical neurons were co-transfected with indicated plasmids and visualized by dual fluorescence. Specific suppression of EGFP–UbB+1 by RFP.UbB+1sh was detected in the cells 24 h after transfection (right column). Note: one EGFP–UbB+1-positive cell with neuritic beading in the right lower two panels is RFP.UbB+1sh-negative. Scale bar represents 30 μm for all the panels. (c) The number of EGFP-positive cells (left panel) or cells with neuritic beads (right panel) relative to RFP-positive cells following co-transfection of EGFP-UbB+1 and either RFP.Controlsh or RFP.UbB+1sh. The bars represent mean±S.E. **P's<0.001
Discussion
Neuritic beading has previously been shown to occur in association with neurodegeneration induced by excitotoxins, β-amyloid peptide and other neurotoxic agents. 21, 22, 23 Here, we demonstrate that the mutant protein, UbB+1, causes rapid induction of neuritic beading and subsequent cell death. Although protein overexpression may nonspecifically lead to neuritic morphology, changes consistent with neuritic beading were evident only in UbB+1-expressing cells. Further, Western blotting demonstrated that the amounts of EGFP–Ub and EGFP–UbB+1 expressed with time after transfection were roughly equivalent. Despite abundant levels of EGFP–Ub on Western blots, we did observe a lower intensity of fluorescence signal in EGFP–Ub-expressing cells (Figure 1a, panel 2). This discrepancy might be due to modulation of fluorescence intensity by interactions between EGFP and UPP-related proteins.24 Moreover, neuritic beading of mitochondria was also induced by nontagged UbB+1 (Figure 3c). Taken together, these observations indicate that the demonstrated neuritic effects were specific to UbB+1. A slight increase in the number of beading cells was detected in pEGFP–Ub- or pEGFP-transfected cultures. These changes were not significant and possibly due to minor toxic effects of overexpressed proteins as well as transfection stress.
Ultrastructural studies revealed that the beads contained one or more mitochondria. Although the time frame used for transport measurements in our study was relatively long, significant changes in retrograde movement of mitochondria-associated aggregates were nonetheless observed, whereas changes in anterograde movement were not significant. More specific analyses of mitochondrial velocities and movements over shorter periods of time may help to delineate the dynamics of mitochondrial movement in injured neurons. Whereas there was no effect of UbB+1 expression on the overall levels of P74 dynein and KIF1B kinesin, a significant decrease in the abundance of mitochondria-associated P74 was observed. This most likely resulted from detachment of P74 from mitochondrial cargo based on the findings from a cell-free reconstitution assay. The importance of P74 is demonstrated by its role as a key adaptor molecule involved in linking two 530 kDa dynein heavy chains to cargo for retrograde transport.25 Accordingly, inhibition of dynein function causes what has been described as ‘organelle jams’ or ‘clogs’.26 Our results therefore support the hypothesis that UbB+1-mediated detachment of P74 from mitochondrial cargo disturbs the physiological balance between anterograde and retrograde transport. The finding that mitochondria–MT associations were significantly reduced in the processes of UbB+1-transfected neurons provides further evidence in support of this hypothesis. There was a minor decrease in retrograde axonal transport that was not statistically significant in EGFP–Ub-transfected cells. This suggests that the overexpression of wild-type Ub might also lead to a slight accumulation of polyUb conjugates, which interferes with motor–MT interactions. The combined effects of UbB+1-induced detachment of P74 from mitochondrial cargo and altered MT associations may profoundly disable dynein-driven retrograde motion, thereby leading to an imbalance between anterograde and retrograde movements and a buildup of mitochondria in neurites. Moreover, both UbB+1 and dynein play key roles in the formation of aggresomes during the cellular response to misfolded proteins.27, 28 Although we did not specifically identify aggresomes in this study, it is possible that aggresomes may compete with mitochondria for P74 binding and thereby facilitate mitochondrial detachment and impairment of retrograde transport.28 UbB+1-mediated actions may also be modulated by heat-shock protein family members,29 which may be important regulators of axonal transport.30
In addition to mitochondria, the transport machinery conveys other membranous and nonmembranous cargo between the neuronal soma and its processes.17 Although we did not detect either ER or Golgi bodies in neuritic beads, the distribution of these organelles and other cargo in neuronal processes might still be perturbed by UbB+1-mediated disruption of axonal transport. Thus, impaired axonal transport with abnormal organelle clustering in neurites has been implicated in the pathophysiology of several neurodegenerative diseases.31 Moreover, induction of neuritic beading by impairment of the UPP suggests that the adverse effects of UbB+1 on axonal transport may result, in part, from inhibition of proteasome function. In this regard, proteasome inhibition induces neuritic beading in cultured neurons (Tan et al., unpublished observation), and UbB+1 as well as polyubiquitinated protein aggregates accumulate in the pathological hallmarks of AD, such as neurofibrillary tangles and neuritic plaques.7 In addition, axon damage has been implicated in the initial stages of Alzheimer's-related neuronal degeneration.32, 33 Although amyloid precursor protein and presenilin mutations, as well as tau isoforms, may disrupt axonal transport,34, 35 UbB+1-mediated clogging of mitochondria in neurites may also contribute to the mechanism of axon injury and neuropathology in AD.
The pathophysiological significance of neuritic beading is underscored by the detection of mitochondrial stress in this study. The results suggest that there is a temporal relationship between activation of mitochondrial stress in neurites and neuronal cell death. Notably, mitochondrial stress indicated by the loss of mitochondrial membrane potential seemed to start in peripheral neurites and then spread toward the cell soma. Loss of mitochondrial membrane potential may lead to a local release of cytochrome c in neurites that triggers the mitochondrial cell death cascade through caspase-3 activation. Accordingly, about 70% of UbB+1-positive cells exhibited a loss of mitochondrial membrane potential in peripheral neurites relatively early on, which preceded a more global loss of mitochondrial membrane potential, mitochondrial oxidative stress and cell death indicated by TUNEL staining. Our findings therefore support the idea that overwhelming mitochondrial stress in neuronal processes, possibly triggered, in part, by impaired transport, may activate a cell death-signaling cascade that is propagated from the periphery to the cell body in a dying-back fashion.
Our observation that UbB+1 induced a significant accumulation of polyUb conjugates is in agreement with prior studies that demonstrated inhibition of proteasome function by UbB+1.8, 9 Impaired UPP function likely contributed to the accumulation of tumor suppressor p53, which is a major UPP substrate.36 Our results showed that activation of the p53 pathway plays an important role in UbB+1-induced neuronal degeneration, as cell death was substantially decreased in neurons from p53-null mice. In addition, absence of p53 resulted in a significant reduction in the number of cells exhibiting mitochondrial oxidative stress. These findings suggest that p53 might enhance mitochondrial stress either through direct interaction, as has previously been shown, or an indirect mechanism yet to be defined.37 A link between impaired proteasome activity and mitochondrial oxidative stress has recently been demonstrated.38 In the present study, UbB+1-mediated UPP dysfunction may therefore facilitate mitochondrial superoxide production and contribute to mitochondrial stress.
The use of siRNA is an efficient way to block expression of specific genes, including alleles that differ by only one nucleotide.20 Our results demonstrate that siRNA/shRNA can effectively knockdown expression and prevent the toxic effects of UbB+1 protein in cultured neurons. Thus, siRNA might be a potential therapeutic approach to counteract the accumulation of UbB+1 protein in certain neurological disorders. In addition to UbB+1 mRNA, dinucleotide deletions in other transcripts may increase with aging due to a process of molecular misreading.39 RNA interference may therefore be an effective tool to control the expression of abnormal transcripts associated with aging.
In summary, our results demonstrate that UbB+1 causes a buildup of mitochondria in neurites and suggest that this results from dysregulation of motor protein–MT interactions. Impaired transport may then lead to mitochondrial stress and activation of both the mitochondrial and p53 cell death pathways. siRNA-mediated knockdown of UbB+1 expression prevents neuritic beading and may be an effective therapeutic strategy for certain neurodegenerative diseases. Current investigations are aimed at understanding specific interactions between UbB+1 and other polyubiquitinated protein aggregates and axonal transport motors, and relevant effects on mitochondrial movements. These studies may also help to elucidate molecular mechanisms of axon damage in age-related neurodegenerative diseases.
Materials and Methods
Antibodies, plasmids and reagents
Rabbit polyclonal antibodies directed against EGFP and full length p53, mouse monoclonal antibodies for P74, that is, dynein intermediate chain, goat polyclonal IgG against KIF1B, goat polyclonal IgG against kinesin heavy chain, A/G plus agarose beads and biotinylated or HRP-conjugated secondary antibodies were obtained from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Mouse monoclonal IgG against β-actin, rabbit polyclonal IgGs against Ub, porin/VDAC1 and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) were from Sigma-Aldrich (St Louis, MO, USA). The Cytochrome c Releasing Kit was purchased from Biovision (Mountain View, CA, USA). Rabbit polyclonal IgG for UbB+1 as well as recombinant UbB+1 were obtained from R&D Systems (Minneapolis, MN, USA). Semisynthetic recombinant Ub5+1 protein was from BIOMOL International (Plymouth Meeting, PA, USA). Organelle-targeting reporters and cDNA expression constructs including pEGFP-C1, pDsRed2-Mito, pEYFP-ER and pEYFP-Golgi were purchased from Clontech (Palo Alto, CA, USA). pEGFP or pCDNA3.1 containing either a single copy of human Ub or UbB+1 cDNA was subcloned as described.9 pEGFP-UbB was subcloned using polymerase chain reaction (PCR) and cDNA from human brain tissue as template. The fluorescent reporter of p53 activation (pp53-TA-DsRed2) was subcloned from the pp53-TA-Luc vector (Clontech). All of the DNA oligonucleotides used for subcloning were synthesized by Integrated DNA Technologies (Coralville, IA, USA). Cell culture media and Lipofectamine 2000 transfection reagents were obtained from Invitrogen (Carlsbad, CA, USA). CombiMag Transfection Enhancing reagents were purchased from Boca Scientific (Boca Raton, FL, USA). Purified bovine Ub was from Sigma-Aldrich. PCR reagents and related restriction enzymes were purchased from Promega (Madison, WI, USA) and New England Biolabs (Ipswich, MA, USA), respectively. Silencer siRNA construction kit and pSilencer-1.0 U6 vector were obtained from Ambion (Houston, TX, USA). ECL reagents for Western blotting were obtained from Amersham Biosciences (Piscataway, NJ, USA). DEVD-CHO was purchased from Biosource (Camarillo, CA, USA). In Situ Cell Death Detection Kit – TMR Red was obtained from Roche (Indianapolis, IN, USA).
Animals, cell cultures and transfections
Timed-pregnant Sprague–Dawley rats and C57Bl/6 mice were purchased from Charles River Laboratories (Wilmington, MA, USA). p53-null (B6.129S2-Trp53tm1Tyj) breeding pairs were obtained from the Jackson Laboratory (Bar Harbor, ME, USA). All animals were housed in the University of California Irvine School of Medicine Vivarium on a 12-h light/dark cycle with food and water available ad libitum. p53-null genotype was confirmed by PCR according to the vendor's protocol. Primary cortical neurons were prepared from embryonic day 18 (E18) Sprague–Dawley rats and postnatal day 1 (P1) mice and maintained as described previously.18 For electron microscopy studies, neurons were grown on poly-D-lysine-coated Eppendorf CELLocate coverslips (Brinkmann Instruments, Westbury, NY, USA). All procedures were approved by the Institutional Animal Care and Use Committee of the University of California Irvine School of Medicine. Mouse N2A neuroblastoma cells were grown in MEM supplemented with heat-inactivated fetal bovine serum (10%), pyruvate and nonessential amino acids. Plasmid transfections were performed with Lipofectamine 2000 (Invitrogen) in six-well plates following the manufacturer's protocol. Briefly, mixtures of Lipofectamine 2000 diluted in DMEM and plasmid(s) were incubated at room temperature for 20 min and directly applied to each well with 2 ml fresh serum-free MEM for N2A cells or Neurobasal medium for neurons. Fresh media were substituted 8 h after transfection. To enhance the transfection efficiency of nontagged Ub and UbB+1 genes in pcDNA3.1 plasmids, magnetofection was performed using CombiMag transfection kit combined with Lipofectamine 2000 in six-well plates according to the manufacturer's instruction. Transfection efficiencies as determined by both phase contrast and fluorescence microscopy ranged from 70–90% in N2A cells and 15–30% in primary neurons.
In situ functional activity assays, immunocytochemistry and TdT-mediated dUTP nick end labeling (TUNEL)
The transcriptional activity of p53 protein was evaluated by co-transfection of pp53-TA-DsRed2 (0.4 μg/well) and either pEGFP–UbB+1 or pEGFP–Ub (1.6 μg/well) as described above. Changes in mitochondrial membrane potential, mitochondrial oxidative stress and caspase-3 activation were assayed using TMRM, MitoSox Red (MSR) (Molecular Probe/Invitrogen) and the CaspaTag Fluorescein Caspase Activity Kit (Intergen, Purchase, NY, USA), respectively, following the manufacturers’ protocols. Cells were then fixed with 4% paraformaldehyde in 1 × PBS (pH 7.4) at 4°C for 20 min and immunostained as described previously.18, 19 Primary antibody against p53 or UbB+1 protein was used at a concentration of 1 : 100. Neuronal cell death was evaluated by TUNEL using an In Situ Cell Death Detection kit following the manufacturer's protocol. In some experiments, TUNEL was evaluated in neurons transfected with EGFP–UbB+1 with or without the caspase-3-specific inhibitor, DEVD-CHO (20 μM).
Microscopy and image analysis
Fluorescence microscopy was performed with a Leica DMLB microscope. Images were recorded using a Spot II CCD camera with equal exposure times for specific fluorescence reporters or antibodies. For quantification, all EGFP-positive neurons were counted in 10 randomly chosen fields at a magnification of × 200. Those neurons showing three or more neuritic beads in their processes were counted as ‘beading cells’. Up to 60 beading cells were included in the analysis of the number of beads per beading cell. For each of the functional assays, that is, TMRM or MSR staining, p53 activation and caspase-3 activity, results were tabulated for a total of 60 randomly counted EGFP- or UbB+1-positive cells. Only those EGFP-UbB+1 neurons that showed lack of TMRM staining in at least three distinguishable EGFP-highlighted beads were counted as neuritic TMRM-negative cells. Colocalization studies of neuritic beads and mitochondria were performed with a BioRad confocal laser optical dissection system.
Time-lapse analysis of mitochondrial movements
Neurons were co-transfected with DsRed2-Mito and either EGFP, EGFP–Ub or EGFP–UbB+1. Movements of mitochondria-containing aggregates were analyzed in neuronal processes from 25 randomly chosen neurons 16–18 h after transfection. Anterograde and retrograde movements were quantified by a direct comparison of two photomicrographs taken of the same field 30 min apart. The results are depicted as the average number of labeled mitochondria moving retrograde or anterograde within a given field.
Electron microscopic analysis
Transfected neurons grown on glass coverslips were randomly selected and pre-positioned against an etched grid based on EGFP fluorescence. Cells were then sequentially fixed in 2% (w/v) paraformaldehyde and 2% (v/v) glutaraldehyde in 0.1 M phosphate buffer (PB, pH 7.4) at 4°C for 1 h, rinsed with PB, postfixed with 1% OsO4-buffered solution for 1 h, pre-stained with 1% (w/v) uranyl acetate for 1 h, dehydrated in ethylic alcohol and embedded in Epon 812 (50°C × 24 h then 60°C × 48 h). Glass coverslips on embedded blocks were removed following immersion in 49% (v/v) hydrofluoride for 10–15 min and embedded cells were cut out and sectioned at 70 nm (N=3–4 cells for each group), followed by en bloc staining with 2% (w/v) uranyl acetate for 2 min and Reynold's lead citrate for 2 min. Transmission electron micrographs were acquired using a Philips CM10 electron microscope. To collect images for analysis of neuritic beads, a neurite was identified by morphological features including continuity with a cell body. All of the neuronal images used in the analysis were nonoverlapping and acquired sequentially from the cell body through its processes. For quantification, mitochondria were classified on the basis of their relationship with nearby MTs in longitudinal sections. Mitochondria or mitochondria-like structures that were flanked by MTs on two sides were defined as ‘MT-associated’, whereas mitochondria with either no adjacent MTs or those with MTs on only one side were defined as ‘MT-dissociated’.
Mitochondria/cytosol fractionation and reconstitution assay
Subcellular fractionation was performed using a Mitochondria/Cytosol Fractionation Kit (BioVision), following the manufacturer's protocol. Purified mitochondrial fractions were re-suspended in buffer and total protein determined using the Total Protein Kit (Sigma-Aldrich). For the reconstitution assay, 40 μg total protein from the mitochondrial fraction of untreated primary neurons was incubated with 0.5–2.0 μg Ub, UbB+1 or a polyubiquitinated form of UbB+1, Ub5+1, in a total volume of 40 μl for 20 min at room temperature. Following brief vortexing and centrifugation at 12 000 r.p.m. (4°C 15 min), the mitochondrial pellet and supernatant were each mixed with protein-loading buffer and analyzed by Western blotting for P74 abundance.
Western blotting
Western blotting was performed as described previously.18, 19 β-actin, porin and GAPDH were used as loading controls for whole-cell lysates, mitochondrial and cytosolic fractions, respectively. Primary antibodies were used at a concentration of 1 : 100 (kinesin heavy chain, KIF1B), 1 : 200 (Ub, UbB+1, EGFP, p53, p74, GAPDH) or 1 : 1000 (β-actin, porin). Quantification of gel pixel intensity was performed with UN-Scan-It software.
Small Interfering RNA-mediated suppression of UbB+1
siRNA corresponding to UbB+1 mRNA with the dinucleotide deletion was designed with two base overhangs on each strand and prepared using the Ambion siRNA construction kit and the following oligonucleotide templates: sense 5′-AAAGAGGGTATGCAGATCTTCCCTGTCTC-3′ and antisense 5′-AAGAAGATCTGCATACCCTCTCCTGTCTC-3′ according to the manufacturer's instructions. A construct expressing a hairpin siRNA and a fluorescent reporter under the direction of two different promoters was developed from Clontech pDsRed2-1 and Ambion pSilencer-1.0 U6 vectors as well as published data. Co-transfection of reporter plasmids and UbB+1 siRNA was performed using identical conditions as for plasmid transfections with Lipofectamine 2000. pEGFP–UbB was used as a control for specificity of UbB+1 RNA interference.
All of the results were confirmed by three or four independent experiments. Statistical analysis was performed using JMP 6 software (SAS Institute, Cary, NC, USA). Cell counts and gel pixel intensity were evaluated by analysis of variance (ANOVA) with post-hoc Fisher's protected least significant difference (PLSD) procedure or Student's t-test.
Abbreviations
- Ub:
-
ubiquitin
- UbB+1:
-
ubiquitin B+1
- AD:
-
Alzheimer's disease
- UPP:
-
uibiquitin-proteasome pathway
- siRNA:
-
small interfering RNA
- Aβ:
-
beta-amyloid
- CNS:
-
central nervous system
- MT:
-
microtubule
- EGFP:
-
enhanced green fluorescent protein
- EYFP:
-
enhanced yellow fluorescent protein
- ER:
-
endoplasmic reticulum
- TMRM:
-
tetramethyl rhodamine methyl ester
- MSR:
-
MitoSox Red
- TUNEL:
-
terminal dUTP nick-end labeling
- shRNA:
-
small hairpin RNA
- RFP:
-
red fluorescent protein
- GAPDH:
-
glyceraldehyde-3-phosphate dehydrogenase
- EM:
-
electron microscopy
- PCR:
-
DNA polymerase chain reaction
References
Hershko A, Ciechanover A . The ubiquitin system. Annu Rev Biochem 1998; 67: 425–479.
Schwartz AL, Ciechanover A . The ubiquitin-proteasome pathway and pathogenesis of human diseases. Annu Rev Med 1999; 50: 57–74.
Jesenberger V, Jentsch S . Deadly encounter: ubiquitin meets apoptosis. Nat Rev Mol Cell Biol 2002; 3: 112–121.
Scheper W, Hol EM . Protein quality control in Alzheimer's disease: a fatal saviour. Curr Drug Targets CNS Neurol Disord 2005; 4: 283–292.
Fischer DF, De Vos RA, Van Dijk R, De Vrij FM, Proper EA, Sonnemans MA et al. Disease-specific accumulation of mutant ubiquitin as a marker for proteasomal dysfunction in the brain. FASEB J 2003; 17: 2014–2024.
Hol EM, van Leeuwen FW, Fischer DF . The proteasome in Alzheimer's disease and Parkinson's disease: lessons from ubiquitin B+1. Trends Mol Med 2005; 11: 488–495.
van Leeuwen FW, de Kleijn DP, van den Hurk HH, Neubauer A, Sonnemans MA, Sluijs JA et al. Frameshift mutants of beta amyloid precursor protein and ubiquitin-B in Alzheimer's and Down's patients. Science 1998; 279: 242–247.
Lam YA, Pickart CM, Alban A, Landon M, Jamieson C, Ramage R et al. Inhibition of the ubiquitin-proteasome system in Alzheimer's disease. Proc Natl Acad Sci USA 2000; 97: 9902–9906.
Lindsten K, de Vrij FM, Verhoef LG, Fischer DF, van Leeuwen FW, Hol EM et al. Mutant ubiquitin found in neurodegenerative disorders is a ubiquitin fusion degradation ubstrate that blocks proteasomal degradation. J Cell Biol 2002; 157: 417–427.
De Vrij FM, Sluijs JA, Gregori L, Fischer DF, Hermens WT, Goldgaber D et al. Mutant ubiquitin expressed in Alzheimer's disease causes neuronal death. FASEB J 2001; 15: 2680–2688.
Qian SB, Ott DE, Schubert U, Bennink JR, Yewdell JW . Fusion proteins with COOH-terminal ubiquitin are stable and maintain dual functionality in vivo. J Biol Chem 2002; 277: 38818–38826.
Aridor M, Hannan LA . Traffic jams II: an update of diseases of intracellular transport. Traffic 2002; 3: 781–790.
Hollenbeck PJ, Saxton WM . The axonal transport of mitochondria. J Cell Sci 2005; 118: 5411–5419.
Goldstein LS, Yang Z . Microtubule-based transport systems in neurons: the roles of kinesins and dyneins. Annu Rev Neurosci 2000; 23: 39–71.
Nangaku M, Sato-Yoshitake R, Okada Y, Noda Y, Takemura R, Yamazaki H et al. KIF1B, a novel microtubule plus end-directed monomeric motor protein for transport of mitochondria. Cell 1994; 79: 1209–1220.
Morris RL, Hollenbeck PJ . Axonal transport of mitochondria along microtubules and F-actin in living vertebrate neurons. J Cell Biol 1995; 131: 1315–1326.
Hirokawa N, Takemura R . Molecular motors and mechanisms of directional transport in neurons. Nat Rev Neurosci 2005; 6: 201–214.
Tan Z, Tu W, Schreiber SS . Downregulation of free ubiquitin: a novel mechanism of p53 stabilization and neuronal cell death. Brain Res Mol 2001; 91: 179–188.
Tan Z, Qu W, Tu W, Liu W, Baudry M, Schreiber SS . p53 accumulation due to down-regulation of ubiquitin: relevance for neuronal apoptosis. Cell Death Differ 2000; 7: 675–681.
Miller VM, Xia H, Marrs GL, Gouvion CM, Lee G, Davidson BL et al. Allele-specific silencing of dominant disease genes. Proc Natl Acad Sci USA 2003; 100: 7195–7200.
Roediger B, Armati PJ . Oxidative stress induces axonal beading in cultured human brain tissue. Neurobiol Dis 2003; 13: 222–229.
Ivins KJ, Bui ET, Cotman CW . Beta-amyloid induces local neurite degeneration in cultured hippocampal neurons: evidence for neuritic apoptosis. Neurobiol Dis 1998; 5: 365–378.
Takeuchi H, Mizuno T, Zhang G, Wang J, Kawanokuchi J, Kuno R et al. Neuritic beading induced by activated microglia is an early feature of neuronal dysfunction toward neuronal death by inhibition of mitochondrial respiration and axonal transport. J Biol Chem 2005; 280: 10444–10454.
Berse M, Bounpheng M, Huang X, Christy B, Pollmann C, Dubiel W . Ubiquitin-dependent degradation of Id1 and Id3 is mediated by the COP9 signalosome. J Mol Biol 2004; 343: 361–370.
Hirokawa N . Kinesin and dynein superfamily proteins and the mechanism of organelle transport. Science 1998; 279: 519–526.
Martin M, Iyadurai SJ, Gassman A, Gindhart Jr JG, Hays TS, Saxton WM . Cytoplasmic dynein, the dynactin complex, and kinesin are interdependent and essential for fast axonal transport. Mol Biol Cell 1999; 10: 3717–3728.
Bardag-Gorce F, Riley N, Nguyen V, Montgomery RO, French BA, Li J et al. The mechanism of cytokeratin aggresome formation: the role of mutant ubiquitin (UBB+1). Exp Mol Pathol 2003; 74: 160–167.
Johnston JA, Illing ME, Kopito RR . Cytoplasmic dynein/dynactin mediates the assembly of aggresomes. Cell Motil Cytoskeleton 2002; 53: 26–38.
Hope AD, Lashley T, Lees AJ, De Silva R . Failure in heat-shock protein expression in response to UBB+1 protein in progressive supranuclear palsy in humans. Neurosci Lett 2004; 359: 94–98.
Tsai MY, Morfini G, Szebenyi G, Brady ST . Release of kinesin from vesicles by hsc70 and regulation of fast axonal transport. Mol Biol Cell 2000; 11: 2161–2173.
Hirokawa N, Takemura R . Molecular motors in neuronal development, intracellular transport and diseases. Curr Opin Neurobiol 2004; 14: 564–573.
Song MS, Saavedra L, De Chaves EI . Apoptosis is secondary to non-apoptotic axonal degeneration in neurons exposed to Abeta in distal axons. Neurobiol Aging 2005; 27: 1224–1238.
Mandelkow EM, Stamer K, Vogel R, Thies E, Mandelkow E . Clogging of axons by tau, inhibition of axonal traffic and starvation of synapses. Neurobiol Aging 2003; 24: 1079–1085.
Pigino G, Morfini G, Pelsman A, Mattson MP, Brady ST, Busciglio J . Alzheimer's presenilin 1 mutations impair kinesin-based axonal transport. J Neurosci 2003; 23: 4499–4508.
Gunawardena S, Goldstein LS . Disruption of axonal transport and neuronal viability by amyloid precursor protein mutations in Drosophila. Neuron 2001; 32: 389–401.
Maki CG, Huibregtse JM, Howley PM . In vivo ubiquitination and proteasome-mediated degradation of p53(1). Cancer Res 1996; 56: 2649–2654.
Mihara M, Erster S, Zaika A, Petrenko O, Chittenden T, Pancoska P et al. p53 has a direct apoptogenic role at the mitochondria. Mol Cell 2003; 11: 577–590.
Goldbaum O, Vollmer G, Richter-Landsberg C . Proteasome inhibition by MG-132 induces apoptotic cell death and mitochondrial dysfunction in cultured rat brain oligodendrocytes but not in astrocytes. Glia 2006; 53: 891–901.
van Leeuwen FW, Fischer DF, Kamel D, Sluijs JA, Sonnemans MA, Benne R et al. Molecular misreading: a new type of transcript mutation expressed during aging. Neurobiol Aging 2000; 21: 879–891.
Acknowledgements
This study was sponsored by NIH AG-26637 to Z.T., an Alzheimer's Association IIRG grant to S.S.S., NIH NS 27501 to D.K.O.D., NS 33213 to M.A.S., and grants from the Jan Dekker Stichting and Dr Ludgardine Bouwman Stichting (04–22), Internationale Stichting Alzheimer’s Onderzoek (04507/04830), the Van Leersum Foundation, the Hersenstichting Nederland (13F05.11) and the Hersenstichting Nederland-2005, the Matty Brand Stichting, the van Leersum Foundation (2006) and the Society for Progressive Supranuclear Palsy (440-4) to F.W.V.L.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by SA Lipton
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Tan, Z., Sun, X., Hou, FS. et al. Mutant ubiquitin found in Alzheimer's disease causes neuritic beading of mitochondria in association with neuronal degeneration. Cell Death Differ 14, 1721–1732 (2007). https://doi.org/10.1038/sj.cdd.4402180
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4402180
Keywords
This article is cited by
-
Promising protein biomarkers in the early diagnosis of Alzheimer’s disease
Metabolic Brain Disease (2022)
-
Improved neurocognitive performance in FIV infected cats following treatment with the p75 neurotrophin receptor ligand LM11A-31
Journal of NeuroVirology (2021)
-
Deposition of mutant ubiquitin in parkinsonism–dementia complex of Guam
Acta Neuropathologica Communications (2017)
-
Protein Homeostasis, Aging and Alzheimer’s Disease
Molecular Neurobiology (2012)
-
Role of the ubiquitin–proteasome system in nervous system function and disease: using C. elegans as a dissecting tool
Cellular and Molecular Life Sciences (2012)