Abstract
Cell culture work has identified the tumor suppressor p53 as a component of the S-phase checkpoint control system, while in vivo studies of this role of p53 in whole-vertebrate systems were limited. Here, we describe zebrafish mutants in the DNA polymerase delta catalytic subunit 1, based on the positional cloning of the flathead (fla) gene. fla mutants display specific defects in late proliferative zones, such as eyes, brain and cartilaginous elements of the visceral head skeleton, where cells display compromised DNA replication, followed by apoptosis, and partial or complete loss of affected tissues. Antisense-mediated knockdown of p53 in fla mutants leads to a striking rescue of all phenotypic traits, including completion of replication, survival of cells, and normal differentiation and tissue formation. This indicates that under replication-compromised conditions, the p53 branch of the S-phase checkpoint is responsible for eliminating stalled cells that, given more time, would have otherwise finished their normal developmental program.
Similar content being viewed by others
Introduction
During S phase of every cell cycle, the entire genome must be replicated in an error-free manner. This not only requires DNA-replicating proteins with very high fidelity but also the ability of the cell to check for damage caused by endogenous and environmental forces. To date, 19 different DNA polymerases have been identified. Among the ‘classical’ polymerases (α, β, γ, δ, ɛ), Polα, Polδ, and Polɛ are believed to be responsible for the majority of nuclear DNA synthesis, each with specific and unique roles, while Polγ largely acts during mitochondrial replication, and Polβ during DNA repair (reviewed in Hindges and Hübscher,1 and Hübscher et al.2). The Polδ holoenzyme consists of several subunits, the catalytic subunit (Polδ1; p125), which is highly conserved among eukaryotes (50–60% between mammals and yeast), and two to four more divergent smaller subunits with structural and regulatory functions.2 Polδ plays a role in both leading and lagging strand synthesis after taking over for Polα in replication initiation, as demonstrated in the in vitro SV40 replication system.3 Disruptions of Polδ1 in various organisms support its proposed role in DNA replication. In fission yeast, temperature-sensitive mutations of Polδ lead to cell cycle arrest in S phase.4 Furthermore, studies with Xenopus egg extracts have shown that disruption of Polδ via immuno-depletion leads to an accumulation of single-stranded DNA gaps, highlighting Polδ's role in lagging strand synthesis, and the inability of Polɛ to functionally replace Polδ.5 In addition, Polδ1 displays 3′–5′ exonuclease activity and can function in DNA repair.1, 2 The importance of this 3′–5′ exonuclease activity has been highlighted by genetic analyses in mouse, showing that a point mutation specifically disrupting the 3′–5′ exonuclease proofreading activity of Polδ1 causes cancer susceptibility.6, 7 The phenotype of mouse mutants lacking entire Polδ1 function, however, has not been reported.
In the event of dysfunctions at the replication fork during S phase, a replication checkpoint is activated. The function of this checkpoint is four-fold. First, it blocks additional origin firing in the event of DNA damage. Second, elongation is slowed to allow for the assembly and function of repair machinery. Third, the structure of the stalled replication forks is maintained to prohibit additional damage to the DNA, and finally, the affected cells are prohibited from undergoing mitosis (for reviews see Nyberg et al.8 and Bartek et al.9). If replication forks are not repaired and/or stalled for too long, cells either proceed to G2/M arrest10, 11 or undergo apoptosis by the instructions of the tumor suppressor and transcription factor p53 (reviewed in Haupt et al.12). Phosphorylation, stabilization, and activation of p53 are accomplished at least in part by members of the replication checkpoint, namely the PI3K-like protein kinase ATR (Ataxia telangiectasia mutated (ATM) and Rad3 related)13, 14 and its substrate checkpoint kinase 1 (Chk1),15 which have been shown to play a major role during the elimination of cultured cells in response to DNA damage and replication blocks.16, 17, 18, 19, 20 Studies with Xenopus egg extracts have further demonstrated that immunodepletion of Polδ leads to an increase in the phosphorylation of Chk1.5 Together, this suggests that disruption of Polδ might lead to the activation of the replication checkpoint and downstream effectors like p53 via the function of ATR. However, in vivo studies of the interplay of Polδ1 and p53 during replication, cell cycle progression, and apoptosis in a whole-vertebrate system had been missing thus far.
Here, we describe combined Polδ1 and p53 loss-of-function studies in zebrafish larvae. Large-scale genetic screens led to the isolation of hundreds of zebrafish mutants, providing an excellent resource for traditional forward genetic approaches in studying gene function.21, 22 Via positional cloning, we identified the flathead (fla) gene, initially defined according to the craniofacial defects of its mutants,23 as zebrafish Polδ1. Most likely due to early compensation by maternally supplied polδ1 gene products, zygotic zebrafish polδ1 null mutants display rather late phenotypes, characterized by compromised DNA replication and subsequent apoptosis of cells in late proliferative zones, such as the chondrocyte precursors forming the branchial arches. Furthermore, applying morpholino antisense technology, we studied the role of p53 under polδ1-deficient conditions, demonstrating that loss of p53 function leads to a rescue of all phenotypic traits of polδ1 mutants, including apoptosis and cell differentiation, indicating that in this particular case, the p53 control system might be too ‘impatient’, and that cells slowed down in replication can give rise to normal tissues when their p53-dependent elimination is blocked.
Results
fla mutants are characterized by defects in various tissues derived from late proliferative zones
Zebrafish flathead (fla) mutants were originally isolated during the Tuebingen 1996 large-scale ethylnitrosourea (ENU) mutagenesis screen, based on their severe craniofacial defects.21, 23 Specifically, the cartilage of the neurocranium and of pharyngeal arches 1–3 was reduced, while arches 4–7 were completely absent in fla mutants at 120 hpf (Figure 1g, h). Cartilage of the pharyngeal arches is largely derived from neural crest.42 However, according to in situ hybridization analyses with dlx2, marking prechondrogenic neural crest cells,29 there was no difference in the amount of neural crest populating the pharyngeal arches of mutants and wild-type siblings at 36 hpf (data not shown), while at 48 hpf, the number of dlx2-positive cells in the fla mutants was moderately reduced (Figure 1c, d). Similarly, the endodermal pouches, which play an important role in patterning and survival of the pharyngeal arches,43 were formed normally in fla mutants, as indicated by the normal staining intensity and pattern of Zn5 immunostainings before 48 hpf (data not shown). However, craniofacial defects became more apparent during the next day of development, when the arch-populating crest cells undergo increased cell proliferation, as revealed by in situ hybridizations for the replication marker pcna (proliferating-cell-nuclear-antigen; see below; Figure 3j), and by BrdU incorporation analyses (see below; Figure 3k). Thus, at 72 hpf, expression of sox9,30 a marker for differentiating neural crest-derived chondrocytes, was particularly compromised in the gill arches (pharyngeal arches 3–7) of fla mutants (Figure 1e, f), suggesting that cells either fail to specify, or that they have become severely reduced in number due to decreased proliferation and/or enhanced apoptosis. Consistent with the latter notion, fla mutants also showed defects in other tissues known to be highly proliferative during late developmental stages, such as the eyes and the brain, which were of normal size during the first 40 h of development, but much smaller than in wild-type siblings during later stages of development (see Figure 1a, b for 120 hpf). In contrast, other regions and tissues of fla mutant larvae appeared morphologically normal.
fla mutants display defects in craniofacial, brain, and eye development. All panels show fla mutants (b, d, f, h) in comparison to wild-type siblings (WT; a, c, e, g). (a, b) Live larvae at 120 hpf; lateral view over entire length of larvae. (c–f) Whole mount in situ hybridization with probes staining cartilage-forming neural crest cells at different stages of development. (c, d) 48 hpf, dlx2; (e, f) 72 hpf, sox9. Arrowheads point to crest cells forming the first two pharyngeal arches, mandibulare and hyoid, arrows to more posterior crest cells forming pharyngeal arches 3–7 (gill arches). Comparison of (e) and (f) reveals that sox9 expression in arches 1 and 2 is reduced in fla mutants, while staining in the gill arches is completely absent. Similarly, sox9 expression in the chondrocytes of the pectoral fin is absent in fla mutants (indicated with asterisks). (g, h) Alcian blue stainings of cartilaginous elements of the head skeleton at 120 hpf. ‘ep’ indicates the ethmoid plate of the neurocranium; numbers 1–7 indicate the pharyngeal arches (1=mandibulare; 2=hyoid; both are jaw-forming arches; 3–7=gill/branchial arches). Abbreviations: WT, wild type; h, hours post fertilization (hpf)
polδ1 is uniformly expressed early, whereas expression becomes restricted to zones of high proliferation during later stages of zebrafish development. All panels show wild-type embryos or larvae of ages indicated in lower right corner (in h=hours post fertilization/hpf). (a–j) Whole mount in situ hybridizations for polδ1 (a–i) or the replication marker pcna (j), respectively. (k, l) BrdU incorporation patterns. (a, b, d, f, g, i–k) Lateral views on entire embryo (a) or on the head region (others), and (c, e, h, l) dorsal views on head region. Arrows in (b,c) point to trunk region that has become devoid of polδ1 transcripts. Abbreviations: cmz, ciliary marginal zone of eyes; ey, eye; gi, gastrointestinal system; gt, gut; liv, liver; mbh, midbrain–hindbrain boundary; nc, neural crest cells; pcr, pancreas; pfb, pectoral fin buds; tec, tectum; vr, ventral retina; vz, ventricular zone of brain
fla mutations disrupt zebrafish polδ1 catalytic subunit
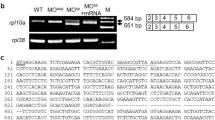
During the Tuebingen screen, five different alleles of fla were isolated, all of which yield morphologically indistinguishable phenotypes.23 We genetically mapped one of them, flata53c, to linkage group 3 between markers z8208 and z14228. For fine-mapping, additional markers were designed, based on genomic sequences in the vicinity of z14228 (Figure 2a; see also Materials and Methods and Supplementary Data). Ultimately, this led to the identification of Ensembl genomic contig ctg24833.1, which contains four putative open reading frames (ORFs), and which is flanked on either side by a north or a south marker, respectively, each with a distance of approximately 0.05 cM to the fla locus (NP3, NP4; one recombination in 1912 meioses) (Figure 2a). One of the putative ORFs was predicted to encode Polδ1 catalytic subunit, consistent with the phenotype of fla mutants in proliferative zones (see above). 5′ and 3′ RACE primers were designed to the putative predicted polδ1 transcript sequence. Sequencing of the RACE products revealed that this transcript encodes a protein of 1105 amino-acid residues (GenBank Accession Number AY883095), with 70.9% amino-acid residue identity to both human and mouse Polδ1 (calculated by Jotun Hein method, DNAstar software). We determined the genomic structure of polδ1 and amplified all 27 coding exons from genomic DNA of homozygous wild-type and mutant larvae for three different alleles, flata53c, flath5b, and flaty76a. For the flata53c allele, we identified a G to A transition in the splice acceptor site preceding coding exon 24 (989–1026), while the flaty76a allele contains a T to A transversion causing a nonsense mutation in exon 12 (C531*), and the flath5b allele a nonsense A to T transversion in exon 16 (L696*) (Figure 2b). C531* results in a truncation of the protein eliminating five of the six polymerase domains, all three conserved C-terminal (CT) domains, and both zinc-finger domains of Polδ1, while L696* mutant Polδ1 lacks three polymerase domains, all three CT domains, and both zinc-finger domains (Figure 2c). In order to examine the effects of the aberrant splice acceptor site, we PCR-amplified and sequenced the 3′ end (corresponding to coding exons 19–27) of polδ1 from wild type and mutant cDNA. We detected two isoforms of mutant transcript, resulting from the usage of alternative splice acceptor sites (Figure 2b). In one case, the splice acceptor preceding exon 25 is used, resulting in an in-frame deletion of exon 24. Exon 24 encodes the first half of the first zinc-finger domain of Polδ1. In the second case, the coding AG sequence at the beginning of exon 24 is used as an alternative splice site, producing a frame-shift and a premature termination of the polymerase removing both zinc-finger domains of Polδ1. The zinc-finger domains have been suggested to be important for internal stability of the protein.1 Furthermore, in the case of mouse Polα144 and yeast Polδ45 and Polɛ,46 these domains have been shown to be required for binding of the catalytic subunits to the B subunits (Cdc1/Polδ2) of the holoenzymes.44, 45, 46 Accordingly, mutational analyses in budding yeast revealed that mutations within the zinc-finger domains of Polδ and Polɛ abolish polymerase function in vivo.46, 47
Phenotype of fla mutants is caused by potential null mutations in zebrafish DNA polymerase delta1. (a) Positional cloning of fla allele ta53c. Meiotic segregation linkage analyses placed fla into an interval on linkage group 3, defined by two flanking polymorphic markers (NP3, NP4), both of which display a genetic distance of 0.05 centimorgan (cM) to the fla mutation. Numbers in middle diagram indicate found recombinations for investigated markers in 1912 meioses. Physically, the NP3/NP4 interval corresponds to 130 kb of DNA containing four predicted open reading frames (ORF), one of which is polδ1. (b) Sequencing of three independent fla alleles revealed a mutation in the splice acceptor site upstream of exon 24 in the case of the ta53c allele (left panel), and nonsense mutations in exon 12 (ty76; middle panel) or exon 16 (th5b; right panel), respectively. First amino-acid residues lost in the resulting truncated proteins are indicated with an asterisk. (c) Cartoon demonstrating the pattern of false splicing leading to the identified mutant polδ1 transcripts (A, B) of the ta53c allele, and cartoons demonstrating the domain structures of wild type and the various mutant Polδ1 proteins encoded by the three fla alleles ta53c, ty76, and th5b. The region deleted in Polδ1 encoded by transcript A of flata53c is indicated in black
In sum, given the importance of the domains lost in the different mutant versions of zebrafish Polδ1, together with the indistinguishable phenotypic strengths of the three alleles, we suspect that they all represent polδ1 null mutations.
polδ1 is restricted to areas with high cell proliferation
Given the rather late and restricted defects caused by loss of Polδ1 function, we examined polδ1 expression by RT-PCR analyses and whole mount in situ hybridization during normal zebrafish development up to 72 hpf. polδ1 mRNA could be detected throughout all investigated stages, including earliest stages before midblastula transition (MBT),48 indicating that it is both maternally and zygotically supplied (data not shown). During cleavage, blastula, gastrula, and early segmentation stages, polδ1 transcripts were ubiquitously and uniformly distributed throughout the entire embryo (Figure 3a; and data not shown). However, during mid-segmentation stages, polδ1 expression started to become restricted. At 24 hpf, expression had largely ceased in the posterior half of the embryo, while it was prominent in the brain, the eye vesicles, and the neural crest (Figure 3b, c). At 32 hpf (data not shown), 48 hpf (Figure 3d–f), and 57 hpf (Figure 3g, h), polδ1 expression had become further restricted, and was largely confined to a ventral patch of the retina and the ciliary marginal zone of the eyes, the midbrain–hindbrain boundary, neural crest cells of the branchial arches, the pectoral fin buds, and the endoderm of the gastrointestinal system. This expression pattern was very similar to that of the replication marker pcna,31 which also showed prominent and restricted expression in the ciliary marginal zone, the midbrain–hindbrain boundary, the tectum, and the neural crest cells of the branchial arches at 57 hpf (Figure 3j). Furthermore, it resembled the pattern obtained in BrdU incorporations studies to label proliferating cells at 48 hpf (Figure 3k, l). At 72 hpf, polδ1 expression had ceased in most tissues except the digestive organs (Figure 3i). In sum, these data indicate that during zebrafish development, polδ1 expression is restricted to proliferating cells, outlining proliferative zones during later stages of development, whereas expression is switched off in differentiating cells.
fla mutants display a transient subtle increase in the number of cells in S phase
Different approaches were taken to study the proliferation and cell cycle state of cells before and during the appearance of morphological defects in fla mutants. At 48 hpf, the staining intensity obtained via BrdU incorporations was strongly reduced in fla mutants (Figure 4b) compared to wild-type siblings (Figure 4a), suggesting that DNA replication was compromised. Consistent results were obtained in FACS analyses determining the DNA content of dissociated cells from heads of single fla mutants and wild-type siblings, accompanied by genotyping of the embryos with genomic DNA extracted from the corresponding trunk and tail region. At 36 hpf, before morphological defects become apparent, fla mutants displayed a subtle, but consistent increase in the fraction of cells with a DNA content higher than 2N (17.5±0.86%; n=5) (Figure 4d), compared to wild-type siblings (14.5±1.18%; n=14) (Figure 4c). The difference is statistically significant (P=0.0002). Given that at these stages, most cells have initiated terminal differentiation (DNA content=2N), while only a rather small subset of cells is proliferative (see pcna and polδ1 in situ hybridizations and BrdU incorporation studies; Figure 3), these results suggest that within this population of proliferating cells, loss of polδ1 leads to a slower progression through S phase.
fla mutants display compromised replication and a transient accumulation of cells with a DNA content higher than 2N, followed by an accumulation of cells with a DNA content smaller than 2N, which can be rescued by p53 deficiency. (a,b) BrdU incorporation in wild-type (a) and fla mutant (b) at 48 hpf, revealing strongly reduced staining intensity in the proliferative zones of the mutant. (c–h) Representative histograms from FACS analyses after DAPI staining of dissociated cells from single heads of embryos at 36 hpf (c, d) or 50 hpf (g, h), and from pooled heads of five embryos at 52 hpf (e, f). At 52 hpf, mutants (f) and wild-type siblings (e) were distinguished by their morphological phenotype, while at 36 hpf (c, d), and at 50 hpf after injection of p53 MO (g, h), when the morphological defects of mutants are very subtle, the genotype was determined via PCR of the corresponding posterior part of the specimen, as described in Materials and Methods. Numbers show the percentages of cells with DNA contents lower (<) or higher (>) than 2N in the area indicated by bars
In contrast, no such increase in the relative ratio of cells in S phase was observed in fla mutants at a later stage, when the defects have become morphologically visible. Rather, at 52 hpf, the number of cells with a DNA content higher than 2N in fla mutants (11.3±0.46%; n=15; three pools with five embryos each; Figure 4f) was similar to that of wild-type siblings (13.1±1.7%; n=15; Figure 4e; difference statistically nonsignificant; P=0.2%). Instead, fla mutants at 52 hpf displayed a significant increase in the fraction of cells with a DNA content lower than 2N, compared both to wild-type siblings of the same stage (18.7±1.68% in fla versus 4.3±1.54% in wild type; P=0.0006) (Figure 4e, f), and to fla mutants at 36 hpf (3.8±1.03%; n=5; compare Figure 4f with Figure 4d; P=0.0009). This <2N fraction is commonly regarded to represent apoptotic cells, suggesting that at these later stages, cells arrested or slowed down in S phase might undergo programmed cell death.
fla mutants display increased apoptosis in late proliferative zones
To directly examine apoptosis in fla mutants, we carried out TUNEL (terminal deoxynucleotidyl transferase-mediated deoxyuridin triphosphate nick-end labeling; Figure 5a–d) and acridine orange stainings (Figure 5e–p), both of which yielded very similar results. No increased apoptosis rates were found in fla mutants prior to 36 hpf. At 38 hpf, TUNEL stainings revealed higher numbers of apoptotic cells particularly in the tectum of fla mutants (Figure 5a–d). At 50 hpf, fla mutants displayed high numbers of acridine orange-positive cells in all late proliferative zones, such as the midbrain–hindbrain boundary (Figure 5f, j), the eye vesicles (Figure 5n), and the branchial arch region (Figure 5j), while wild-type embryos only showed dispersed acridine orange-positive cells (Figure 5e, i, m). Actually, the pattern of acridine orange staining in fla mutant embryos resembled very much the pattern obtained via BrdU incorporation studies in wild-type embryos (compare Figure 5f with Figure 3l). In sum, this suggests that in fla mutant embryos, cells slowed down or arrested in S phase due to the loss of Polδ1 activity, are eliminated by apoptosis.
fla mutants display increased apoptosis in late proliferative zones, which can be completely rescued by p53 deficiency. (a–d) TUNEL stainings at 38 hpf, all other panels show acridine orange stainings at 50 hpf. The genotype of larvae (wild type or fla), and the injected morpholino antisense oligonucleotides (p53 MO or four mismatch control morpholino p53 4mm MO) are shown in lower left corner. (a, b, i–l) Lateral view, (c, d) frontal view, (e–h) dorsal view on head region; (m–p) magnified lateral view on eye. In (j, k) cells forming the cartilage of the gill arches (arrowheads) and of the pectoral fin buds (arrows) are indicated (compare with Figure 1e, f). Abbreviations: mbh, midbrain–hindbrain boundary; tc, tectum; vz, ventricular zone of tectum
Disruption of Polδ1 leads to an upregulation of genes involved in the p53 pathway
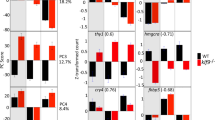
Studies in cell culture systems have indicated that the tumor suppressor protein p53 is part of the S-phase checkpoint control systems, eliminating replication-arrested cells by inducing programmed cell death. In order to examine if the p53 pathway is activated in response to disruption of Polδ1 function, we examined wild type and flata53c mutant single embryos for p53 transcript levels. Quantitative real-time RT-PCR analysis showed that p53 transcript levels were elevated approximately two-fold in mutants (Figure 6a). Additionally, p53 protein activity appeared to be increased, as indicated by the two-fold increased mRNA levels of the direct p53 target gene p21.36, 49 p21 encodes a CDK inhibitor that mediates DNA damage-induced cell cycle arrest both during G1/S and G2/M transition.50 In contrast, the reported p53 target gene bax,39, 51 which is involved in mediating p53-dependent apoptosis via the mitochondrial pathway,12, 52 showed unaltered mRNA levels in fla mutants (Figure 6a). These in vivo results support previous data obtained in cell culture studies showing a role of p53 for G2/M arrest after replication blockage,10, 11 while its role to mediate apoptosis remains elusive.
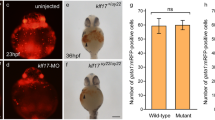
fla mutants display increased expression of p53 and p53 target genes, while morphological structures can be completely rescued by p53 deficiency. (a) Quantitative real time RT-PCR of fla and wild-type sibling mutants at 46 hpf, showing increased transcript levels of p53 and its target gene p21 in fla mutants, while bax transcript levels are normal. Mean relative expression ratios compared to ef1α (fold), standard deviations (Stdev), and P-values for comparison of wild type and fla numbers (P; statistically significant difference when < 0.05) are listed on the right side. (b–m) Injection of p53 MO, but not the four mismatch control MO, leads to a rescue of fla morphology. The genotype of larvae, and the injected morpholino antisense oligonucleotides are shown in upper right corner. (b–e) Dorsal views on the head region of live larvae at 120 hpf; fla mutants injected with p53 MO display a significant normalization of eye size. (f–m) Ventral view on heads stained with alcian blue at 72 hpf, (f–i) or 120 hpf (j–m). Numbers indicate pharyngeal arches (compare with Figure 1). At 120 hpf fla mutant injected with p53 MO (l) has regained pharyngeal arches 4–7 absent in regular or control-injected fla mutants (k, m). At 72 hpf, the differentiation of rescued chondrocytes in pharyngeal arches 4–7 of the fla mutant injected with p53 MO (h) is slightly delayed compared to the non-injected wild-type sibling (f) and to the wild-type sibling injected with p53 MO (i)
p53 deficiency rescues both apoptotic and morphological traits of the fla phenotype
To investigate whether despite the normal bax mRNA levels in fla mutants, the observed elimination of replication-compromised cells is p53-dependent, we generated polδ1-p53 double deficient zebrafish. Previous studies have demonstrated that p53 protein levels can be efficiently decreased and p53-induced apoptosis can be inhibited by the injection of p53 antisense MOs (p53-MO). p53 morphant embryos themselves show no developmental defects (data not shown; Langheinrich et al.36), similar to p53-deficient mice.53 This is in contrast to p53-deficient Xenopus embryos, which display compromised cell differentiation during early stages of development.54, 55, 56 Here, p53 is required for proper transforming growth factor beta (TGFβ) signaling and mesoderm formation, consistent with a physical interaction of p53 with Smad proteins.55, 56 The reasons for the different requirements of p53 during early mammalian, fish and amphibian development are unclear, but might be due to the presence of other p53 family members (p63, p73) in mammals and zebrafish, whereas they appear to be absent during early stages of Xenopus development.56
Two different p53 MOs were injected into clutches of flata53c or flath5b mutant embryos at the one-cell stage. Consistent with a role of p53 as a proapoptotic factor in replication-compromised cells, we found a significant rescue of apoptosis in fla mutants injected with p53 MOs. Acridine orange stainings revealed that apoptosis rates in all investigated proliferative zones (tectum, midbrain–hindbrain boundary, eye vesicles, craniofacial domain) of p53-deficient fla mutants (Figure 5g, k, o) were indistinguishable from those in wild-type siblings (Figure 5e, i, m) during days 2 and 3 of development, and strongly reduced compared to regular fla mutants (Figure 5f, j, n) or mutants injected with a 4-mismatch control MO (Figure 5h, l, p).
We also investigated cell proliferation behavior of polδ1-deficient cells in the absence of p53, carrying out DNA content-dependent flow cytometric experiments with heads of single genotyped embryos at 50 hpf, as described above. Consistent with the aforementioned results obtained via acridine orange stainings, the ratio of cells with a DNA content smaller than 2N, representing apoptotic cells, was significantly reduced in p53-deficient fla mutants (5.8±0.37%; n=6) compared to noninjected fla mutants (18.7±1.46%) (compare Figure 4h with Figure 4f; P=0.002), and was similar to that of wild-type embryos injected with p53 MO (4.6±0.71%; n=8) (compare Figure 4h with Figure 4g). However, the ratio of cells with a DNA-content higher than 2N was not increased accordingly. Rather, it was only moderately higher (11.8±0.93%; Figure 4h) than that of the p53 MO-injected wild-type controls (10.2±0.87%; Figure 4g; difference statistically significant; P=0.008). This suggests that in the absence of p53, replication-compromised fla mutant cells neither die nor irreversibly arrest, but can progress through S phase, possibly with slightly reduced speed.
Such surviving p53-deficient fla mutant cells also appear to be able to finish their normal developmental program, as indicated by the rescue of all morphological traits of fla mutants upon p53 MO injection. Noninjected fla mutants or mutants injected with a four mismatch p53 control MO could easily be detected at 48 hpf by their characteristic small head/small eyes phenotype, which is even more apparent at 72 hpf (data not shown) and at 120 hpf (Figure 6c; and data not shown). However, p53 MO-injected clutches displayed no mutant phenotypes at 48 hpf, while at 72 and at 120 hpf, the size of the eyes and the head of p53-deficient fla mutants was much less reduced than in noninjected fla mutants (compare Figure 6d with Figure 6c). In addition, alcian blue stainings at 120 hpf revealed that the craniofacial cartilage defects of fla mutants were rescued to almost wild-type condition upon loss of p53 activity. Specifically, gill arches 4–7, while absent in noninjected fla mutants at 72 and 120 hpf (Figure 6g, k), and not fully developed in p53 MO-injected fla mutants at 72 hpf (Figure 6h), were present at normal sizes and shapes in injected mutants at 120 hpf (Figure 6l). These regained alcian blue-positive cartilaginous elements mostly consist of postmitotic, differentiated chondrocytes, indicating that in the absence of p53, polδ1-deficient cells rescued from apoptosis can undergo terminal differentiation. However, rescued fla mutants were not viable. Although they developed swim bladders and swam normally, they died around 12 dpf, while non-injected fla mutants lived until 8 or 9 dpf.
Discussion
Zygotic Polδ1 is essential for proper development of multiple tissues derived from late proliferative zones
Thus far, genetic analyses addressing the in vivo role of DNA Polδ1 in higher eukaryotic systems had been limited. Here, we identified zygotic polδ1 null mutants in zebrafish, based on the positional cloning of the gene causing the fla phenotype in three independent alleles.23 During development, polδ1 is expressed ubiquitously early, while expression becomes restricted to proliferative zones later (Figure 3). Consistently, defects of polδ1/fla mutants are restricted to tissues derived from these zones, such as eyes, brain, and cartilaginous elements of the head skeleton, all of which are strongly reduced in size. The gill arches are even completely missing, although their progenitor cells were present in normal numbers during earlier stages of development (Figure 1). In FACS analyses determining the DNA content of cells from dissociated zebrafish heads, we found only a subtle accumulation of cells in S phase, suggesting that replication is not completely arrested, but only slowed down (Figure 4). Later, these cells are eliminated by apoptosis, as indicated by the high numbers of acridine orange-positive cells in proliferative zones of fla mutants (Figure 5), and by the accumulation of signals with a DNA content lower than 2N in the FACS studies, while the fraction of cells in S-phase drops (Figure 4). In sum, these data indicate that zygotic Polδ1 is required in multiple proliferative zones of zebrafish larvae to ensure proper DNA replication, and the survival and subsequent differentiation of late proliferating cells.
Maternally supplied Polδ1 protein might be sufficient to drive DNA replication during early stages of zebrafish development
DNA replication is required during all stages of development. Thus, treatment of blastula stage zebrafish embryos with drugs blocking DNA replication leads to an arrest of early development.57 Given the essential role of Polδ1 during DNA replication revealed in lower eukaryotes (e.g. fission yeast4) and in higher in vivo-like systems (Xenopus egg extracts5), the rather late appearance of defects in zebrafish polδ1 zygotic null mutants seems surprising. We can only speculate about possible reasons. There could be a second polδ1 gene, resulting from the genome duplication during teleost evolution.58 However, we thus far failed to identify such a second gene in genomic and EST database searches. Also, functional redundancy with its closest relative Polɛ seems quite unlikely in light of results obtained via immuno-depletions in Xenopus egg extracts, demonstrating that the two polymerases cannot replace each other.5, 59
In this light, we favor a model in which zygotic Polδ1 might be dispensable for early DNA replication because of functional compensation by maternally supplied polδ1 gene products. mRNAs and proteins of many so-called house-keeping genes have been shown to be synthesized during oogenesis and deposited in the egg. Indeed, we could detect high levels of polδ1 mRNA in zebrafish embryos before the onset of MBT, when zygotic transcription of genes is initiated (4 hpf).48 However, maternal mRNAs have been shown to be usually degraded during the first 10–15 hpf,60 whereas the phenotype of fla mutants only becomes apparent during day 2 of development. Accordingly, knocking down both maternal and zygotic polδ1 mRNAs with antisense MOs targeting the translational start codon generated phenotypes not much stronger and earlier than those of fla mutants (NP and MH, unpublished observations).
Thus, it appears more likely that it is maternally provided Polδ1 protein, rather than maternal polδ1 mRNA, that compensates for zygotic Polδ1 during early stages of zebrafish development. We do not know how much maternally supplied Polδ1 protein is present in early stage zebrafish embryos, since none of several tested antibodies against mammalian Polδ1 crossreacted with the zebrafish protein. However, in amphibian oocytes, DNA polymerases have been shown to be over 100 000-fold enriched compared to somatic cells, and we suspect that the same might be true for zebrafish. Maternal proteins can be much more stable than maternal mRNAs, as for instance revealed for Mcm5, another component of the replication complex, which is detectable in zygotic mcm5 mutants until day 2 of development (Soojin Ryu and Wolfgang Driever, personal communication). A similar slow and gradual loss of maternal Polδ1 protein would not only provide an excellent explanation for the late appearance of the fla mutant phenotype but also for the observed normalization of the fla phenotype upon p53 deficiency.
Zygotic Polδ1 becomes more dispensable under p53-deficient conditions
Thus far, implications of p53 deficiency on the rescue of apoptosis during vertebrate development have been mainly studied in the context of DNA repair, such as in the case of Polβ-deficient mice61 and of Fancd-morphant zebrafish.39 In the context of DNA replication stress, similar studies have been largely restricted to cell culture systems. Thus, it was shown that blockage of DNA replication through application of aphidicolin or hydroxyurea in mouse or human fibroblast cultures leads to an increase in p53 protein level and to an S-phase arrest, which is released in p53-deficient fibroblasts.10 Similarly, S-phase arrest and subsequent apoptosis of mouse ES cells deficient for the protein kinase Cdc7 are accompanied by an increase in p53 mRNA, protein and activity levels, while apoptosis can be repressed upon chemical inactivation of p53.62
Here, we show that polδ1 zebrafish mutants display an increase in p53 transcription and p53 activity (Figure 6), while loss of p53 allows completion of replication (Figure 4), as well as survival (Figure 5) and differentiation of replication-compromised cells (Figure 6). For the same reasons as described above, we think that completion of replication in rescued polδ1-p53 double-deficient cells is driven by maternally supplied Polδ1 protein. Assuming that in zygotic polδ1 mutants, such maternal protein only runs out slowly, p53-dependent apoptosis of replication-compromised cells might occur when maternal Polδ1 has dropped below a critical threshold, for instance when not all replication forks initiated under the participation of Polα can be directly taken over by Polδ. These transiently stalled replication forks, if present for too long, would then lead to the activation of p53 to eliminate the corresponding cells. However, there might still be enough maternal Polδ1 present to complete DNA replication in the absence of p53, when cells are given more time. This notion is supported by our findings that in polδ1-p53 double-deficient zebrafish, chondrocytes of the rescued gill arches differentiate slightly later than in regular p53 morphants (Figure 6). In this light, the p53 branch of the S-phase control checkpoint might be too tight and ‘impatient’, eliminating cells that would otherwise have caught up. However, polδ1-p53 double-deficient zebrafish are not perfectly rescued; death is just postponed for 3–4 days (compare with Kim et al.62). This could be due to the progressive reappearance of p53; MO-based gene knockdown normally starts to be attenuated after 3 dpf. In addition, it could mean that zygotic Polδ1 eventually becomes absolutely essential even under p53-deficient conditions, for instance when maternal protein has been completely lost.
It remains unclear how p53 mediates apoptosis of replication-compromised cells. Interestingly, we found bax mRNA levels to be normal in fla mutant embryos, suggesting that the mitochondrial death pathway is not involved. This is consistent with data obtained in mouse, showing that although DNA damage fails to induce apoptosis in p53-deficient cells, it can do so in cells deficient for Bax or death receptors (reviewed in Rich et al.63)
p53 is only one of the effectors of the S-phase checkpoint control system
Cell culture work has identified the kinase ATR as the central component of the S-phase checkpoint control, which upon replication stress activates other proteins in parallel to p53 (see Introduction and Nyberg et al.8 and Bartek et al.9 for reviews). We have preliminary data that in contrast to p53 itself, the ATR checkpoint control system in its entirety is required for normal cell cycle progression in replication-compromised zebrafish cells. Thus, when knocking down zebrafish ATR with antisense MOs, we observed a significant worsening of the fla mutant defects, with a loss of pharyngeal arches 2 and 3 in addition to arches 4–7 absent in regular fla mutants. In contrast, ATR deficiency in wild-type embryos had no visible effects (NP and MH, unpublished data). This suggests that cells of arches 2 and 3 of fla mutants also display (more moderately) compromised replication, but can complete S phase before they are caught by p53, whereas in the absence of ATR, they undergo precocious entry into mitosis, ultimately leading to their elimination. This is in clear opposition to the situation observed in p53/polδ1-double deficient animals, in which the absence of p53 appears to allow all replication-compromised cells of arches 2–7 to survive, while the presence of ATR enables them to enter mitosis only after they have finished replication. This implies that in developing zebrafish, other ATR target proteins of the S-phase checkpoint system regulate entry into mitosis, whereas p53 itself is dispensable for this process, in contrast to its reported role in cultured cells.10
Craniofacial development under Polδ1-compromised conditions
Within the pharyngeal arches, fla mutants display a deletion of arches 4–7, while arches 1–3 develop with rather normal size, which initially had prompted us to believe that fla might be involved in anterior–posterior (AP) patterning of the visceral head skeleton. What might account for these AP effects under replication-compromised conditions? Specification of cartilage-forming cephalic neural crest cells occurs in an AP-wave, with cells from anterior arches differentiating before cells from more posterior arches.64 In this light, chondrocytes forming arches 1–3 might already have completed their final round of replication before Polδ1 levels drop below the critical threshold for p53 activation, whereas chondrocytes forming arches 4–7 are caught by the checkpoint control system and eliminated. Along the same lines, the differential responses among arches 1–3 to ATR deficiency could be explained (see above). In any case, our data suggest that the defects of other craniofacial zebrafish mutants with fla-like phenotypes23, 65 might be caused by defects in genes involved in DNA replication and/or S-phase checkpoint control.
Materials and Methods
Maintenance and identification of flata53c, flaty76, and flath5b
Zebrafish were bred and maintained, and embryos were treated using standard conditions.24 flata53c, ty76, th5b mutant lines25 were obtained from the Tuebingen stock center and were maintained in the Tuebingen background. flata53c was outcrossed to the WIK line for genetic mapping.26 Mutant larvae were identified live at 3–5 days postfertilization (dpf) by their obvious craniofacial morphology and eye size reduction.
Genetic and physical mapping of the fla locus
The flata53c mutation was mapped to linkage group 3 via PCR analysis of simple sequence length polymorphism (SSLP) markers (Tuebingen version 4) on mutant (25 larvae) and wild-type (75 larvae) genomic DNA pools.27.Genomic sequence from the fla locus was identified by sequence similarity searches of markers previously mapped near z14228 (http://www.map.tuebingen.mpg.de/) against the Ensembl Zv2 genome database (http://www.ensembl.org/). New polymorphic markers were designed from this sequence for fine mapping of single mutant larvae (see Supplementary Data). Contig ctg24833.1 was identified by a sequence similarity search with the marker fi11h05.x1 against the Ensemble genomic database. All PCR fragments were amplified with a 55°C annealing temperature using standard protocols.
Cloning of zebrafish polδ1 and identification of mutations were carried out as described in Supplementary data.
Tissue labeling procedures
Whole mount in situ hybridization was carried out as previously described.28 For polδ1 in situ probe synthesis, plasmid pCRII-zfpolδ3’ was linearized with NotI and transcribed with SP6 RNA polymerase. In addition, riboprobes of the following cDNAs were used as described: dlx2,29 sox9,30 pcna.31
Whole mount immunostainings were carried out as previously described,32 using the Vectastain Elite ABC kit (Vector Laboratories), and the primary antibody Zn5 (Zebrafish International Resource Center; University of Oregon, Eugene) at 1 : 500. To visualize craniofacial cartilage, 120 hpf (hours postfertilization) embryos were stained with alcian blue (Sigma) as described.23
For detection of apoptotic cells, TUNEL (terminal desoxynucleotidyl transferase-mediated biotinylated UTP nick end labeling) stainings were performed using the In Situ Cell Death Detection Kit (Roche), essentially as described by Cole and Ross.33 Acridine orange stainings of apoptotic cells were performed as previously described.34
For labeling of proliferating cells via incorporation of 5-Bromo-2′-deoxyuridine (BrdU), BrdU was delivered by incubating live zebrafish embryos for 30 min in BrdU solution (10 mM BrdU; 15% DMSO in E3 medium) on ice, followed by three washes in E3 medium and further incubation for 1 h at 28.5°C. Afterwards, embryos were fixed in 4% PFA/PBS for 2 h at room temperature or over night at 4°C. After several washes in PBS, 0.1% Tween-20 (PBST), and one brief wash in distilled water, embryos were incubated in acetone at −20°C for 10 min. For further permeablization, embryos were digested for 6 min at room temperature in 0.25% trypsin in PBS without Mg2+ and Ca2+, and refixed in 4% PFA for 20 min. Embryos were then rinsed three times in 0.1% Tween-20 and two times with 2 N HCl, followed by incubation in 2 N HCl for 1 h. Afterwards, embryos were rinsed three times in PBST and blocked for 10 min in immunostaining blocking solution,32 followed by subsequent incubation with the primary anti-BrdU antibody (Boehringer 1170376; 1 : 100 in blocking solution) and the secondary AlexaFluor647-conjugated anti-mouse antibody (Molecular Probes, Eugene, OR, USA; 1 : 200 in blocking solution), using standard conditions.
Morpholino injections
Antisense morpholino oligonucleotides (MOs) were injected at the 1–4 cell stage and at a concentration of 0.1 mM, as previously described.35 The sequences of used MOs were: AGA ATT GAT TTT GCC GAC CTC CTC T (p53-MO1); GCG CCA TTG CTT TGC AAG AAT TG (p53-MO236); the mismatch control MO, with mismatched nucleotides denoted in lowercase, was GCa CCA TcG CTT gGC AAG cAT TG (p53-4mm36).
RNA extraction and quantitative real time RT-PCR analysis
Total RNA was isolated from 46 hpf whole wild-type or mutant embryos using 750 μl of Trizol reagent (Gibco/BRL). First strand cDNA synthesis was performed on 2 μg of total RNA using oligo dT primer and Superscript III reverse transcriptase (Invitrogen). Quantitative real time RT-PCR (QRT-PCR) was performed with the SYBR green PCR kit (Applied Biosystems) in a DNA Engine Opticon-Continuous Fluorescence Detection system (MJ Research), using the following primer sets: elongation factor 1α (ef1α)37 (control): sense, TCA CCC TGG GAG TGA AAC AGC; antisense, ACT TGC AGG CGA TGT CAG CAG; p53:38 sense, GCG ATG AGG AGA TCT TTA CCC; antisense, ACA AAG GTC CCA GTG GAG TG; p21:39, 40 sense, AGC TGC ATT CGT CTC GTA GC; antisense, TGA GAA CTT ACT GGC AGC TTC A; bax:39, 40 sense, GCA GTG GCA ATG ACC AGA TA; antisense, GGA AAA CTC CGA CTG TCT GC. All samples were quantified by the comparative cycle threshold (Ct) method for relative quantification of gene expression, normalized to ef1α.41 Differences between two groups were analyzed using a one-tailed Student's t-test assuming unequal variances.
Cell dissociation, genotyping, and fluorescence-activated cell sorting (FACS) analysis
Embryos were anaesthetized with triacine,24 and heads were cut immediately posterior to the eyes. Genomic DNA was isolated from the posterior part of each embryo, PCR-amplified with exon24 primers giving a 498 bp fragment, and digested overnight with Hpy188I. Digests were run on a 2% agarose gel and scored for their genotype. As the flata53c mutation results in the disruption of an Hpy188I site, the digest of wild-type DNA yields bands of 257, 174, and 67 bp, while mutant DNA results in fragments of 324 and 174 bp. The heads of single embryos were incubated at room temperature in 100 μl papain solution (2 mg/ml papain (Sigma), 0.3 mg/ml cysteine (Sigma) in magnesium/calcium-free PBS) for 30 min. An additional 100 μl of papain solution was added to the suspension for 30 min, cells were dissociated by trituration, and kept on ice. Triton X-100 was added to dissociated cells at a final concentration of 0.1%, followed by addition of 4′,6-Diamidino-2-phenylindole (DAPI) (Molecular Probes, Eugene, OR; Product number D-3571) at a final concentration of 3 μM. Cells were analyzed on an LSRII flow cytometer (BD Biosciences, San Jose, CA, USA). Data evaluation was carried out using FlowJo (Tree Star, Ashland, OR, USA). To avoid a loss of apoptotic cells by gating onto the forward/sideward scatter population, ungated histograms were used for the measurement of DNA content. To determine the ratio of cells with DNA contents smaller or higher than 2N, the area of the histograms with 10–80% (<2N) or 125–250% (>2N) of the peak channel were measured in comparison to the total area. Standard deviations (±) and probabilities associated with Student's t-test (P; Tail 2; Type 3) were calculated using Microsoft Exel software.
Accession codes
Abbreviations
- AP:
-
anterior–posterior
- ATR:
-
Ataxia telangiectasia mutated (ATM) and Rad3 related
- BrdU:
-
5′ Bromo-2′desoxyuridine
- Chk1:
-
checkpoint kinase 1
- DAPI:
-
4′,6-Diamidino-2-phenylindole
- dpf:
-
days postfertilization
- ef1α:
-
elongation factor 1 alpha
- FACS:
-
fluorescence-activated cell sorting
- fla:
-
flathead mutant
- hpf:
-
hours postfertilization
- MO:
-
morpholino oligonucleotide
- pcna:
-
proliferating-cell nuclear antigen
- polδ1:
-
DNA polymerase delta1
- RT-PCR:
-
polymerase chain reaction after reverse transcription of RNA
- SSLP:
-
simple sequence length polymorphism
- TGFβ:
-
transforming growth factor beta
- TUNEL:
-
terminal desoxynucleotidyl transferase-mediated biotinylated UTP nick end labeling
- 4mm:
-
four mismatch control
References
Hindges R and Hübscher U (1997) DNA polymerase delta, an essential enzyme for DNA transactions. Biol. Chem. 378: 345–362
Hübscher U, Maga G and Spadari S (2002) Eukaryotic DNA polymerases. Annu. Rev. Biochem. 71: 33–63
Waga S and Stillman B (1994) Anatomy of a DNA replication fork revealed by reconstitution of SV40 DNA replication in vitro. Nature 369: 207–212
Francesconi S, Park H and Wang TS (1993) Fission yeast with DNA polymerase delta temperature-sensitive alleles exhibits cell division cycle phenotype. Nucleic Acids Res. 21: 3821–3828
Fukui T, Yamauchi K, Muroya T, Akiyama M, Maki H, Sugino A and Waga S (2004) Distinct roles of DNA polymerases delta and epsilon at the replication fork in Xenopus egg extracts. Genes Cells 9: 179–191
Goldsby AE, Lawrence NA, Hays LE, Olmsted EA, Chen X, Singh M and Preston BD (2001) Defective DNA polymerase-δ proofreading causes cancer susceptibility in mice. Nat. Med. 7: 638–639
Goldsby AE, Hays LE, Chen X, Olmsted EA, Slayton WB, Spangrude GJ and Preston BD (2002) High incidence of epithelial cancers in mice deficient for DNA polymerase δ proofreading. Proc. Natl. Acad. Sci. USA 99: 15560–15565
Nyberg KA, Michelson RJ, Putnam CW and Weinert TA (2002) Toward maintaining the genome: DNA damage and replication checkpoints. Annu. Rev. Genet. 36: 617–656
Bartek J, Lukas C and Lukas J (2004) Checking on DNA damage in S phase. Nat. Rev. Mol. Cell Biol. 5: 792–804
Taylor WR, Agarwal ML, Agarwal A, Stacey DW and Stark GR (1999) p53 inhibits entry into mitosis when DNA synthesis is blocked. Oncogene 18: 283–295
Taylor WR and Stark GR (2001) Regulation of the G2/M transition by p53. Oncogene 20: 1803–1815
Haupt S, Berger M, Goldberg Z and Haupt Y (2003) Apoptosis – the p53 network. J. Cell Sci. 116: 4077–4085
Tibbets RS, Brumbaugh KM, Williams JM, Sarkaria JN, Cliby WA, Shieh S-Y, Taya Y, Prives C and Abraham RT (1999) A role of ATR in the DNA damage-induced phosphorylation of p53. Genes Dev. 13: 152–157
Lakin ND, Hann BC and Jackson SP (1999) The ataxia-telangiectasia related protein ATR mediates DNA-dependent phosphorylation of p53. Oncogene 18: 3989–3995
Shieh SY, Ahn J, Tamai K, Taya Y and Prives C (2000) The human homologs of checkpoint kinases Chk1 and Cds1 (Chk2) phosphorylate p53 at multiple DNA damage-inducible sites. Genes Dev. 14: 289–300
Sanchez Y, Wong C, Thoma RS, Richman R, Wu Z, Piwnica-Worms H and Elledge SJ (1997) Conservation of the Chk1 checkpoint pathway in mammals: linkage of DNA damage to Cdk regulation through Cdc25. Science 277: 1497–1501
Liu Q, Guntuku S, Cui XS, Matsuoka S, Cortez D, Tamai K, Luo G, Carattini-Rivera S, DeMayo F, Bradley A, Donehower LA and Elledge SJ (2000) Chk1 is an essential kinase that is regulated by Atr and required for the G(2)/M DNA damage checkpoint. Genes Dev. 14: 1448–1459
Takai H, Tominaga K, Motoyama N, Minamishima YA, Nagahama H, Tsukiyama T, Ikeda K, Nakayama K and Nakanishi M (2000) Aberrant cell cycle checkpoint function and early embryonic death in Chk1(−/−) mice. Genes Dev. 14: 1439–1447
Guo Z, Kumagai A, Wang SX and Dunphy WG (2000) Requirement for Atr in phosphorylation of Chk1 and cell cycle regulation in response to DNA replication blocks and UV-damaged DNA in Xenopus egg extracts. Genes Dev. 14: 2745–2756
Feijoo C, Hall-Jackson C, Wu R, Jenkins D, Leitch J, Gilbert DM and Smythe C (2001) Activation of mammalian Chk1 during DNA replication arrest: a role for Chk1 in the intra-S phase checkpoint monitoring replication origin firing. J. Cell Biol. 154: 913–923
Haffter P, Granato M, Brand M, Mullins MC, Hammerschmidt M, Kane DA, Odenthal J, van Eeden FJM, Jiang Y-J, Heisenberg C-P, Kelsh RN, Furutani-Seiki M, Warga RM, Vogelsang E, Beuchle D, Schach U, Fabian C and Nüsslein-Volhard C (1996) The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development 123: 1–36
Driever W, Solnica-Krezel L, Schier AF, Neuhauss SCF, Malicki J, Stemple DL, Stanier DYR, Zwartkruis F, Abdeliah S, Rangini Z, Belak J and Boggs C (1996) A genetic screen for mutations affecting embryogenesis in zebrafish. Development 123: 37–46
Schilling TF, Piotrowski T, Grandel H, Brand M, Heisenberg CP, Jiang YJ, Beuchle D, Hammerschmidt M, Kane DA, Mullins MC, van Eeden FJ, Kelsh RN, Furutani-Seiki M, Granato M, Haffter P, Odenthal J, Warga RM, Trowe T and Nusslein-Volhard C (1996) Jaw and branchial arch mutants in zebrafish I: branchial arches. Development 123: 329–344
Westerfield M (1994) The Zebrafish Book: A Guide for the Laboratory use of Zebrafish (Eugene, Oregon: University of Oregon Press)
Schilling TF (1997) Genetic analysis of craniofacial development in the vertebrate embryo. BioEssays 19: 459–468
Rauch G-J, Granato M and Haffter P (1997) A polymorphic zebrafish line for genetic mapping using SSLPs on high-percentage agarose gels Technical Tips Online T01208
Geisler R (2002) In Zebrafish: A Practical Approach, Nüsslein-Volhard C, Dahm R, eds (Oxford: Oxford University Press) pp. 175–212
Hammerschmidt M, Pelegri F, Mullins MC, Kane DA, van Eeden FJM, Granato M, Brand M, Furutani-Seiki M, Haffter P, Heisenberg C-P, Jiang Y-J, Kelsh RN, Odenthal J, Warga RM and Nüsslein-Volhard C (1996) dino and mercedes, two genes regulating dorsal development in the zebrafish embryo. Development 123: 95–102
Akimenko M-A, Ekker M, Wegner J, Lin W and Westerfield M (1994) Combinatorial expression of three zebrafish genes related to distal-less: part of a homeobox gene code for the head. J. Neurosci. 16: 3475–3486
Chiang EF-L, Pai CI, Wyatt M, Yan Y-L, Postlethwait J and Chung B-C (2001) Two Sox9 genes on duplicated zebrafish chromosomes: expression of similar transcription activators in distinct sites. Dev. Biol. 231: 149–163
Lee J-S and Gye MC (1999) Zebrafish Danio rerio proliferating cell nuclear antigen (PCNA) cloning and characterization. Fish. Sci. 65: 955–958
Schulte-Merker S, Ho RK, Herrmann BG and Nüsslein-Volhard C (1992) The protein product of the zebrafish homologue of the mouse T gene is expressed in nuclei of the germ ring and the notochord of the early embryo. Development 116: 1021–1032
Cole LK and Ross LS (2001) Apoptosis in the developing zebrafish embryo. Dev. Biol. 240: 123–142
Herzog W, Sonntag C, von der Hardt S, Roehl HH, Varga ZM and Hammerschmidt M (2004) Fgf3 signaling from the ventral diencephalon is required for early specification and subsequent survival of the zebrafish adenohypophysis. Development 131: 3681–3692
Nasevicius A and Ekker SC (2000) Effective targeted gene ‘knockdown’ in zebrafish. Nat. Genet. 26: 216–220
Langheinrich U, Hennen E, Stott G and Vacun G (2002) Zebrafish as a model organism for the identification and characterization of drugs and genes affecting p53 signaling. Curr. Biol. 12: 2023–2028
Nordness S, Krauss S and Johansen T (1994) cDNA sequence of zebrafish (Brachydanio rerio) translation elongation factor-1 alpha: molecular phylogeny of eukaryotes based on elongation factor-1 alpha protein sequences. Biochim. Biophys. Acta 1219: 529–532
Cheng R, Ford BL, O’Neal PE, Mathews CZ, Bradford CS, Thongtan T, Barnes DW, Hendricks JD and Bailey GS (1997) Zebrafish (Danio rerio) p53 tumor suppressor gene: cDNA sequence and expression during embryogenesis. Mol. Mar. Biol. Biotechnol. 6: 88–97
Liu TX, Howlett NG, Deng M, Langenau DM, Hsu K, Rhodes J, Kanki JP, DAndrea AD and Look AT (2003) Knockdown of zebrafish Fancd2 causes developmental abnormalities via p53-dependent apoptosis. Dev. Cell 5: 903–914
Nowak M, Köster C and Hammerschmidt M (2005) Perp is required for tissue-specific cell survival during zebrafish development. Cell Death. Diff. 12: 52–64
Livak KJ and Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25: 402–408
Schilling T, Walker C and Kimmel CK (1996) The chinless mutation and neural crest cell interactions in zebrafish jaw development. Development 122: 1417–1426
David NB, Saint-Etienne L, Tsang M, Schilling TF and Rosa FM (2002) Requirement for endoderm and FGF3 in ventral skeleton formation. Development 129: 4457–4468
Mizuno T, Yamagishi K, Miyazawa H and Hanaoka F (1999) Molecular architecture of the mouse DNA polymerase α-primase complex. Mol. Cell. Biol. 19: 7886–7896
Sanchez Garcia J, Cuifo LF, Kearsey SE and MacNeill SA (2004) The C-terminal zinc finger of the catalytic subunit of DNA polymerase δ is responsible for direct interaction with the B-subunit. Nucl. Acids Res. 32: 3005–3016
Dua R, Levy DL and Campbell JL (1999) Analysis of essential functions of the C-terminal protein/protein interaction domain of Saccharomyces cerevisiae pol e and its unexpected abilitiy to support growth in the absence of the DNA polymerase domain. J. Biol. Chem. 274: 22283–22288
Simon M, Giot L and Faye G (1993) A random mutagenesis procedure – application to the pol3 gene of Saccharomyces cerevisiae. Gene 127: 139–144
Kane DA and Kimmel CB (1993) The zebrafish midblastula transition. Development 119: 447–456
El-Deiry WS, Tokino T, Velculescu VE, Levy DB, Parsons R, Trent JM, Lin D, Mercer WE, Kinzler KW and Vogelstein B (1993) WAF1, a potential mediator of p53 tumor suppression. Cell 75: 817–825
Niculescu III AB, Chen X, Smeets M, Hengst L, Prives C and Reed SI (1998) Effects of p21 (Cip1/Waf1) at both the G1/S and the G2/M cell cycle transition: pRb is a critical determinant in blocking DNA replication and in preventing endoreduplication. Mol. Cell. Biol. 18: 629–643
Mayashita T and Reed JC (1995) Tumor suppressor p53 is a direct transcriptional activator of the human BAX gene. Cell 80: 293–299
Hengarter MO (2000) The biochemistry of apoptosis. Nature 407: 770–776
Donehower LA, Harvey M, Slagle BL, McArthur MJ, Montgomery Jr CA, Butel JS and Bradley A (1992) Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature 356: 215–221
Wallingford JB, Seufert DW, Virta VC and Vize PD (1997) p53 activity is essential for normal development in Xenopus. Curr. Biol. 7: 747–757
Takebayashi-Suzuki K, Funami J, Tokumori D, Saito A, Watabe T, Miyazono K, Kanda A and Suzuki A (2003) Interplay between the tumor suppressor p53 and TFGb signaling shapes embryonic body axes in Xenopus. Development 130: 3929–3939
Cordenonsi M, Dupont S, Maretto S, Insinga A, Imbriano C and Piccolo S (2003) Links between tumor suppressors: p53 is required for TGF-beta gene responses by cooperating with Smads. Cell 113: 301–314
Ikegami R, Rivera-Bennetts AK, Brooker DL and Yager TD (1997) Effects of inhibitors of DNA replication on early zebrafish embryos. Evidence for coordinate activation of multiple intrinstic checkpoints at midblastula transition. Zygote 5: 153–175
Postlethwait JH, Yan Y-L, Gates MA, Horne S, Amores A, Brownlie A, Donovan A, Egan ES, Force A, Gong Z, Goutel C, Fritz A, Kelsh R, Knapik E, Liao E, Paw B, Ransom D, Singer A, Thomson M, Abduljabbar TS, Yelik P, Beier D, Joly J-S, Larhammar D, Rosa F, Westerfield M, Zon LI, Johnson SL and Talbot WS (1998) Vertebrate genome evolution and the zebrafish gene map. Nat. Genet. 18: 345–349
Waga S, Masuda T, Takisawa H and Sugino A (2001) DNA polymerase e is required for coordinated and efficient chromosomal DNA replication in Xenopus egg extracts. Proc. Natl. Acad. Sci. USA 98: 4978–4983
Hammerschmidt M, Blader P and Strähle U (1999) In Methods in Cell Biology, Vol. 59, Detrich HW, Westerfield M, Zon LI (eds) (San Diego: Academic Press) pp. 87–115
Sugo N, Niimi N, Aratani Y, Takiguchi-Hayashi K and Koyama H (2004) p53 deficiency rescues neuronal apoptosis but not differentiation in DNA polymerase beta-deficient mice. Mol. Cell. Biol. 24: 9470–9477
Kim JM, Nakao K, Nakamura K, Saito I, Katsuki M, Arai K-i and Masai H (2002) Inactivation of Cdc kinase in mouse ES cells results in S-phase arrest and p53-dependent cell death. EMBO J. 21: 2168–2179
Rich T, Allen RL and Wyllie AH (2000) Defying death after DNA damage. Nature 407: 777–783
Schilling TF and Kimmel CB (1997) Musculoskeletal patterning in the pharyngeal segments of the zebrafish. Development 124: 2945–2960
Neuhauss SCF, Solnica-Krezel L, Schier AF, Zwartkruis F, Stemple DL, Malicki DL, Abdelilah S, Stainier DYR and Driever W (1996) Mutations affecting craniofacial development in zebrafish. Development 123: 357–367
Acknowledgements
We thank M Nowak and T Schilling for stimulating discussions, Soojin Ryu and Wolfgang Driever for communicating unpublished results, F van Eeden (pcna), Robert Kelsh (sox9) and Marc Ekker (dlx2) for reagents, and HG Fronhoefer from the Tuebingen stock center for fla mutant fish. NP is grateful to the Alexander von Humboldt Foundation and to EMBO for her long-term postdoctoral fellowships. Work in MH's laboratory was supported by the Max-Planck Society.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by V De Laurenzi
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Plaster, N., Sonntag, C., Busse, C. et al. p53 deficiency rescues apoptosis and differentiation of multiple cell types in zebrafish flathead mutants deficient for zygotic DNA polymerase δ1. Cell Death Differ 13, 223–235 (2006). https://doi.org/10.1038/sj.cdd.4401747
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401747
Keywords
This article is cited by
-
Antagonistic interactions safeguard mitotic propagation of genetic and epigenetic information in zebrafish
Communications Biology (2024)
-
Zebrafish Noxa promotes mitosis in early embryonic development and regulates apoptosis in subsequent embryogenesis
Cell Death & Differentiation (2014)
-
Plasticity of photoreceptor-generating retinal progenitors revealed by prolonged retinoic acid exposure
BMC Developmental Biology (2011)
-
Chromatin remodelling complex dosage modulates transcription factor function in heart development
Nature Communications (2011)
-
The zebrafish as a model organism for the study of apoptosis
Apoptosis (2010)