Abstract
Gram-negative bacteria are surrounded by an inner cytoplasmic membrane and by an outer membrane, which serves as a protective barrier to limit entry of many antibiotics. The distinctive properties of the outer membrane are due to the presence of lipopolysaccharide1. This large glycolipid, which contains numerous sugars, is made in the cytoplasm; a complex of proteins forms a membrane-to-membrane bridge that mediates transport of lipopolysaccharide from the inner membrane to the cell surface1. The inner-membrane components of the protein bridge comprise an ATP-binding cassette transporter that powers transport, but how this transporter ensures unidirectional lipopolysaccharide movement across the bridge to the outer membrane is unknown2. Here we describe two crystal structures of a five-component inner-membrane complex that contains all the proteins required to extract lipopolysaccharide from the membrane and pass it to the protein bridge. Analysis of these structures, combined with biochemical and genetic experiments, identifies the path of lipopolysaccharide entry into the cavity of the transporter and up to the bridge. We also identify a protein gate that must open to allow movement of substrate from the cavity onto the bridge. Lipopolysaccharide entry into the cavity is ATP-independent, but ATP is required for lipopolysaccharide movement past the gate and onto the bridge. Our findings explain how the inner-membrane transport complex controls efficient unidirectional transport of lipopolysaccharide against its concentration gradient.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Whitfield, C. & Trent, M. S. Biosynthesis and export of bacterial lipopolysaccharides. Annu. Rev. Biochem. 83, 99–128 (2014).
Okuda, S., Sherman, D. J., Silhavy, T. J., Ruiz, N. & Kahne, D. Lipopolysaccharide transport and assembly at the outer membrane: the PEZ model. Nat. Rev. Microbiol. 14, 337–345 (2016).
Chng, S. S., Gronenberg, L. S. & Kahne, D. Proteins required for lipopolysaccharide assembly in Escherichia coli form a transenvelope complex. Biochemistry 49, 4565–4567 (2010).
Sherman, D. J. et al. Lipopolysaccharide is transported to the cell surface by a membrane-to-membrane protein bridge. Science 359, 798–801 (2018).
Narita, S. & Tokuda, H. Biochemical characterization of an ABC transporter LptBFGC complex required for the outer membrane sorting of lipopolysaccharides. FEBS Lett. 583, 2160–2164 (2009).
Ruiz, N., Gronenberg, L. S., Kahne, D. & Silhavy, T. J. Identification of two inner-membrane proteins required for the transport of lipopolysaccharide to the outer membrane of Escherichia coli. Proc. Natl Acad. Sci. USA 105, 5537–5542 (2008).
Sperandeo, P. et al. Characterization of lptA and lptB, two essential genes implicated in lipopolysaccharide transport to the outer membrane of Escherichia coli. J. Bacteriol. 189, 244–253 (2007).
Davidson, A. L., Dassa, E., Orelle, C. & Chen, J. Structure, function, and evolution of bacterial ATP-binding cassette systems. Microbiol. Mol. Biol. Rev. 72, 317–364 (2008).
Thomas, C. & Tampé, R. Multifaceted structures and mechanisms of ABC transport systems in health and disease. Curr. Opin. Struct. Biol. 51, 116–128 (2018).
Okuda, S., Freinkman, E. & Kahne, D. Cytoplasmic ATP hydrolysis powers transport of lipopolysaccharide across the periplasm in E. coli. Science 338, 1214–1217 (2012).
Sherman, D. J. et al. Decoupling catalytic activity from biological function of the ATPase that powers lipopolysaccharide transport. Proc. Natl Acad. Sci. USA 111, 4982–4987 (2014).
Sperandeo, P. et al. Functional analysis of the protein machinery required for transport of lipopolysaccharide to the outer membrane of Escherichia coli. J. Bacteriol. 190, 4460–4469 (2008).
Wu, T. et al. Identification of a protein complex that assembles lipopolysaccharide in the outer membrane of Escherichia coli. Proc. Natl Acad. Sci. USA 103, 11754–11759 (2006).
Freinkman, E., Chng, S. S. & Kahne, D. The complex that inserts lipopolysaccharide into the bacterial outer membrane forms a two-protein plug-and-barrel. Proc. Natl Acad. Sci. USA 108, 2486–2491 (2011).
Chng, S. S., Ruiz, N., Chimalakonda, G., Silhavy, T. J. & Kahne, D. Characterization of the two-protein complex in Escherichia coli responsible for lipopolysaccharide assembly at the outer membrane. Proc. Natl Acad. Sci. USA 107, 5363–5368 (2010).
Bos, M. P., Tefsen, B., Geurtsen, J. & Tommassen, J. Identification of an outer membrane protein required for the transport of lipopolysaccharide to the bacterial cell surface. Proc. Natl Acad. Sci. USA 101, 9417–9422 (2004).
Braun, M. & Silhavy, T. J. Imp/OstA is required for cell envelope biogenesis in Escherichia coli. Mol. Microbiol. 45, 1289–1302 (2002).
Tran, A. X., Dong, C. & Whitfield, C. Structure and functional analysis of LptC, a conserved membrane protein involved in the lipopolysaccharide export pathway in Escherichia coli. J. Biol. Chem. 285, 33529–33539 (2010).
Dong, H., Zhang, Z., Tang, X., Paterson, N. G. & Dong, C. Structural and functional insights into the lipopolysaccharide ABC transporter LptB2FG. Nat. Commun. 8, 222 (2017).
Luo, Q. et al. Structural basis for lipopolysaccharide extraction by ABC transporter LptB2FG. Nat. Struct. Mol. Biol. 24, 469–474 (2017).
Locher, K. P. Mechanistic diversity in ATP-binding cassette (ABC) transporters. Nat. Struct. Mol. Biol. 23, 487–493 (2016).
Ryu, Y. & Schultz, P. G. Efficient incorporation of unnatural amino acids into proteins in Escherichia coli. Nat. Methods 3, 263–265 (2006).
Villa, R. et al. The Escherichia coli Lpt transenvelope protein complex for lipopolysaccharide export is assembled via conserved structurally homologous domains. J. Bacteriol. 195, 1100–1108 (2013).
Putker, F., Bos, M. P. & Tommassen, J. Transport of lipopolysaccharide to the Gram-negative bacterial cell surface. FEMS Microbiol. Rev. 39, 985–1002 (2015).
Bertani, B. R., Taylor, R. J., Nagy, E., Kahne, D. & Ruiz, N. A cluster of residues in the lipopolysaccharide exporter that selects substrate variants for transport to the outer membrane. Mol. Microbiol. 109, 541–554 (2018).
Benedet, M. et al. The lack of the essential LptC protein in the trans-envelope lipopolysaccharide transport machine is circumvented by suppressor mutations in LptF, an inner membrane component of the Escherichia coli transporter. PLoS ONE 11, e0161354 (2016).
May, J. M. et al. The antibiotic novobiocin binds and activates the ATPase that powers lipopolysaccharide transport. J. Am. Chem. Soc. 139, 17221–17224 (2017).
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D 69, 1204–1214 (2013).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Freinkman, E., Okuda, S., Ruiz, N. & Kahne, D. Regulated assembly of the transenvelope protein complex required for lipopolysaccharide export. Biochemistry 51, 4800–4806 (2012).
Simpson, B. W. et al. Identification of residues in the lipopolysaccharide ABC transporter that coordinate ATPase activity with extractor function. MBio 7, e01729-16 (2016).
Matano, L. M. et al. Antibiotic that inhibits the ATPase activity of an ATP-binding cassette transporter by binding to a remote extracellular site. J. Am. Chem. Soc. 139, 10597–10600 (2017).
Chifflet, S., Torriglia, A., Chiesa, R. & Tolosa, S. A method for the determination of inorganic phosphate in the presence of labile organic phosphate and high concentrations of protein: application to lens ATPases. Anal. Biochem. 168, 1–4 (1988).
Acknowledgements
The authors thank J. Chen and M. Oldham (Rockefeller University) for helpful conversations in the early stages of the project. This work used GM/CA and NE-CAT beamlines (P30 GM124165) at the APS synchrotron source (DE-AC02-06CH11357) and an Eiger detector (S10OD021527). We thank the NE-CAT and GM/CA beamline staff for support during X-ray data collection. This work was supported by grants R01GM100951 to N.R., U19 AI109764 to A.C.K., and R01 GM066174 and R01 AI081059 to D.K.
Reviewer information
Nature thanks Russell Bishop, Alessandra Polissi, Robert Tampé and Bert van den Berg for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
D.K. and T.W.O. conceptualized the project. T.W.O. performed crystallographic experiments; T.W.O. and A.C.K. solved and refined the structures. T.W.O., R.J.T. and K.S.P. performed biochemical experiments that B.R.B., N.R. and D.K. helped to conceptualize. B.R.B. performed in vivo functional assays conceptualized by B.R.B. and N.R. T.W.O., R.J.T., K.S.P. and D.K. wrote the manuscript and all authors contributed to editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Purification and reconstitution of wild-type and inactive complexes of V. cholerae and E. cloacae LptB2FGC.
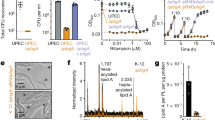
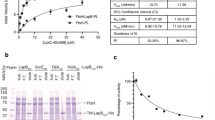
a, To assess the protein quality in purified LptB2FGC complexes, samples were boiled in SDS loading buffer, separated by PAGE using a 4–20% gradient polyacrylamide gel, and then stained with Coomassie blue dye. (1) Wild-type, E. cloacae; (2) catalytically inactive LptB(E163Q), E. cloacae; (3) wild-type, V. cholerae; and (4) catalytically inactive LptB(E163Q), V. cholerae. Note that V. cholerae LptF and LptG co-migrate in this gel, but can be separated under other conditions. Protein samples are each representative of ten or more replicates. b, Schematic of a reconstitution of LPS transport in which purified LptB2FGC and LPS have been incorporated into liposomes by the detergent-dilution method. Addition of LptA modified with a photocrosslinkable amino acid (I36pBPA for E. cloacae, V34pBPA for V. cholerae), along with ATP and Mg2+ leads to time-dependent accumulation of LPS in LptA. LPS transport is monitored by exposing the samples to UV light at the time point shown, quenching transport by addition of SDS loading buffer, PAGE to separate LPS-LptA adducts from LPS and, finally, western blotting for LPS. c, Results of the experiment described in b. Blotting for LPS shows time-dependent accumulation of LPS in LptA when the proteoliposomes contain wild-type LptB2FGC complexes, but accumulation of LPS is not observed when EDTA is substituted for Mg2+, or when the complexes contain LptB(E163Q) rather than wild-type LptB. His5 blots were used to assess the amount of LptA(XpBPA)–His in each sample. E. coli and E. cloacae LptA(I36pBPA)–His6 run as a doublet whereas V. cholerae LptA(I36pBPA)-His6 does not. Blots are representative of data from three biological replicates.
Extended Data Fig. 2 Structures V. cholerae LptB(E163Q)2FGC and E. cloacae LptB2FGC with their electron density maps.
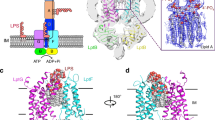
a, Ribbon diagram of V. cholera LptB(E163Q)2FGC (left) and corresponding 2Fo − Fc simulated-annealing composite omit map (right) contoured at 1.0σ. and carved 2.5 Å from the model. b, Ribbon diagram of E. cloacae LptB2FGC (left) and corresponding 2Fo − Fc simulated-annealing composite omit map (right) contoured at 1.0σ. and carved at 2.5 Å from the model. c, Electrostatic surface potential of V. cholerae LptB(E163Q)2FGC depicted with the most basic residues in blue and most acidic residues in red. The approximate edges of the inner membrane are denoted with grey dashed lines. The central panel is in the same orientation as in a. The left panel is a 180° rotation around the axis perpendicular to the membrane. In the right panel, the view in the central panel has been rotated 180° around an axis parallel to the membrane in the plane of the page, and the periplasmic domains are clipped away to give a view of the central cavity between the transmembrane helices. For reference, a ribbon diagram (analogous to Fig. 1c) is shown above the surface diagram.
Extended Data Fig. 3 Alignments of E. coli, E. cloacae, and V. cholerae LptB, LptF, LptG, LptC and LptA.
a, Sequences of Lpt proteins from E. coli K12, E. cloacae subsp. cloacae (ATCC 13047) and V. cholerae C6706. Per cent identity of the E. cloacae and V. cholerae sequences to the E. coli sequence are listed after each alignment. Note that public databases (including Uniprot and NCBI) contain a misannotated LptF sequence for E. cloacae subsp. cloacae (ATCC 13047), with bases GTGATAATCATAAGATATCTG missing from the beginning of LptF; LptF homologues from other E. cloacae strains do not appear to contain this error. Alignments were made using ClustalO and then coloured in MView (https://www.ebi.ac.uk/Tools/msa/clustalo/).
Extended Data Fig. 4 LptC(ΔTM) stably associates with LptB2FG, and facilitates LPS release to LptA as well as full-length LptC without altering the ATPase activity of LptB2FG.
a, In vivo photocrosslinking experiment from Fig. 2b, shown with the corresponding anti-LptC western blot. b, In vivo photocrosslinking experiments (Fig. 2) showed that a pBPA substitution at position R223 of LptF could form LptF–LptC adducts after exposure to UV. In this experiment, proteoliposomes containing LptF, LptB2F(R223pBPA)G were incubated with soluble ΔTM-LptC(ΔTM) at 1×, 2×, 5× or 10× molar excess of LptC and exposed to UV light to test whether LptC(ΔTM) associated with the LptB2F(R223pBPA)G complexes. Western blots for LptC (left) and LptF (right) were used to assess the amount of LptC–LptF adduct formed (top panels) as well as the amounts of LptC and LptF in each sample (bottom panel). The LptF and LptC blots were performed on the same set of samples. The intensity of the LptC–LptF adduct band does not increase with LptC(ΔTM) beyond 2× to 5× molar excess. This result suggests that at those concentrations of LptC(ΔTM), the binding site for LptC(ΔTM) at LptF(R223pBPA) is saturated. The intensity of the LptF band (bottom right panel) does not decrease further because residues R223pBPA can also form intramolecular crosslinks to other parts of the LptF β-jellyroll, and these may be favoured over intermolecular crosslinks to LptC(ΔTM). c, d, Fig. 2c, d shown with 5× and 10× molar excess of LptC(ΔTM). Blots shown in a, b and c are representative of data from three biological replicates.
Extended Data Fig. 5 Comparison of previously published LptB2FG structures to the structure of V. cholerae LptB2FGC.
a, Top-down views of the P. aeruginosa LptB2FG structure (left, PDB ID 5X5Y)20, K. pneumoniae LptB2FG structure (centre, PDB ID 5l75)19 and our V. cholera LptB2FGC structure (right). b, Alignments of P. aeruginosa LptB2FG (left, dark blue) and K. pneumoniae LptB2FG (right, dark blue) with V. cholerae LptB2FGC (LptC, pink; LptF, green; LptG, teal; and LptB, grey).
Extended Data Fig. 6 LPS enters the cavity of LptB2FGC via LptG TM1, LptF TM5 and the LptC transmembrane helix in a nucleotide-independent manner.
a, Western blots from Fig. 3b are shown alongside blots from additional in vivo photocrosslinking experiments comparing LPS crosslinking in wild-type and LptB(E163Q) backgrounds. b, In vitro reconstitution (as in Extended Data Fig. 1b) of LptB2FGC, LptB2FGC(M19pBPA), LptB2FGC(G21pBPA) and LptB2FGC(F78pBPA) shows that the variants with amber codons incorporated in the LptC transmembrane helix release LPS to LptA(I36pBPA) as well as wild type. c, In vitro photocrosslinking of LptB2FGC pBPA variants reconstituted into proteoliposomes with LPS, as in Fig. 3d, comparing complexes containing LptB(E163Q) to those containing wild-type LptB, and the effects of AMPPNP relative to ATP. d–f, Additional crosslinking experiments with pBPA incorporated at other positions in the transmembrane helix of LptC, LptF TM1 and TM5, and LptG TM1 and TM5. g, h, Ribbon diagrams of E. cloacae LptB2FGC, with views of the two potential gates in the LptFGC transmembrane helices; LptG (teal), LptC (pink), and LptF (green). Residues of interest are shown as sticks. Blots for LptG(S30pBPA) are shown in Extended Data Fig. 7b. Blots shown in a–f are representative of data from three biological replicates.
Extended Data Fig. 7 In vivo photocrosslinking experiments show that LPS passes through the β-jellyroll domains of LptF and LptC, but not LptG.
a, Blots from Fig. 3c are shown for reference. b, In vivo photocrosslinking experiments performed analogously to those in Fig. 3b, c. The UV-photocrosslinkable amino acid pBPA was substituted at positions in the periplasmic β-jellyroll domains of LptC, LptF and LptG. Positions in the interior of the LptF and LptC can crosslink to LPS, whereas those facing outside the β-jellyroll do not. No crosslinks to LPS were observed at ten positions in the LptG β-jellyroll; by contrast, LptG(S30pBPA), in TM1 and facing the cavity of the transmembrane domain, can crosslink LPS. c, Ribbon diagram of the structure of the V. cholerae complex, showing the β-jellyroll domains of LptC, LptF, and LptG. Side chains of residues that crosslink LPS are shown in red, whereas residues that do not crosslink LPS are shown in blue. Blots shown in a and b are representative of data from three biological replicates.
Extended Data Fig. 8 Comparison of the V. cholerae and E. cloacae structures reveals two states of the gate in LptF.
a, An overlay of the V. cholerae (LptG, teal; LptF, green; and LptC, magenta) and E. cloacae (LptG, grey; LptF, orange; and LptC, pink) structures shows two conformations of the loop between the third and fourth β-strands of the LptF periplasmic domain. b, Magnified view of this loop and an adjacent loop from LptC in the V. cholerae structure. LptF residues 179–183 and LptC residues 71–74 are depicted as sticks on top of the cartoon. c, Magnified view of the same region in E. cloacae. The 2Fo − Fc omit maps in b and c are contoured at 1.0σ and carved 2.5 Å from the protein.
Extended Data Fig. 9 Complementation and purification of LptF cysteine mutants.
a, Table of results from assays that tested the ability of plasmids encoding E. coli LptF variants to complement a ΔlptF strain. b, To assess whether the LptF double-cysteine mutant was being expressed and translated properly, we measured LptF–Flag levels by anti-Flag western blot in whole-cell lysates from merodiploid E. coli strains containing either pBAD18-LptF-Flag or pBAD18-LptF(S157C/I234C)-Flag. c, Disc diffusion assays were carried out to assess the outer-membrane permeability (relative to wild type) of E. coli strains expressing only plasmid-encoded LptF variants. Values denote the diameter (in mm) of regions with no growth, and values in parentheses denote regions of inhibited growth. Discs were 6 mm in diameter. d, Disc diffusion assays were performed as in c, except that experiments were performed in merodiploid strains with wild-type chromosomal copies of lptFG. Alleles are listed as follows: plasmid-encoded lptFG alleles, a solidus and then the chromosome-encoded lptFG alleles. e, Coomassie-stained samples of purified LptB2FGC used in the LPS release assay: (1) LptF–Flag; (2) LptF(S157C)–Flag; (3) LptF(I234C)–Flag; and (4) LptF(S157C/I234C)–Flag. LptC contained a C-terminal thrombin-cleavable His7 tag used for nickel-affinity chromatography, which was cleaved before reconstitution into liposomes. f, The ATPase activity of E. coli LptB2FGC reconstituted into liposomes was assessed by measuring the rate of phosphate release over time. Data are mean ± s.d. of results from three technical replicates, each of two biological replicates, except two replicates that were performed with one LptB(E163Q)2FGC sample. Data in b and e are representative of data from three biological replicates. Data in c and d are mean ± s.d. calculated from three independent experiments.
Extended Data Fig. 10 Examples of small molecules bound in the crystal structures of V. cholerae and E. cloacae LptB2FGC, and unmodelled electron density in the V. cholerae LptB(E163Q)2FGC structure.
a, Two views of the one molecule of novobiocin bound at the interface between two non-crystallographically symmetric copies of E. cloacae LptB, with 2Fo − Fc simulated-annealing composite omit map contoured at 1.5σ and carved at 2.0 Å. b, Electron density for several ordered detergent molecules is present in the structure of V. cholerae LptB2FGC; here, a DDM molecule is shown bound to LptG with an Fo − Fc Polder map contoured at 2.5σ. c, Ordered PEG400 at the interface of V. cholerae LptB and LptG, with the 2Fo − Fc map contoured at 1.0σ and carved at 2.0 Å. d, View of the lumen between the transmembrane helices of LptC (purple), LptF (green) and LptG (teal), looking towards the inner membrane from the periplasm. The 2Fo − Fc map is contoured at 1.0σ (blue mesh) and the Fo − Fc map is contoured at 3.0σ (green and red meshes). e, View of the same region from within the membrane, with LptF helices in the foreground clipped away; the perspective is approximated by the teal arrow at the top right of d.
Supplementary information
Supplementary Figures
Supplementary Figure 1: Uncropped gels and blots for data shown in main-text and extended data figures. For each figure in the main text and extended data that contains western blots or Coomassie-stained gels, this supplementary figure shows the uncropped blot or gel along with a protein ladder.
Supplementary Table
Supplementary Table 1: X-ray data collection and structure refinement. A table of statistics from X-ray data collection alongside the statistics generated during structure refinement.
Supplementary Table
Supplementary Table 2: Primers used to generate plasmids and strains in this study. Primers used to generate new plasmids for protein expression or complementation assays, including primers for amplifying genes from genomic DNA.
Supplementary Table
Supplementary Table 3: Strains used in this study. Genotypes of E. coli stains used in this study.
Rights and permissions
About this article
Cite this article
Owens, T.W., Taylor, R.J., Pahil, K.S. et al. Structural basis of unidirectional export of lipopolysaccharide to the cell surface. Nature 567, 550–553 (2019). https://doi.org/10.1038/s41586-019-1039-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1039-0
This article is cited by
-
A new antibiotic traps lipopolysaccharide in its intermembrane transporter
Nature (2024)
-
Conformational Investigation of the Asymmetric Periplasmic Domains of E. coli LptB2FGC Using SDSL CW EPR Spectroscopy
Applied Magnetic Resonance (2024)
-
A novel antibiotic class targeting the lipopolysaccharide transporter
Nature (2024)
-
Alternating L4 loop architecture of the bacterial polysaccharide co-polymerase WzzE
Communications Biology (2023)
-
Lipopolysaccharide biosynthesis and traffic in the envelope of the pathogen Brucella abortus
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.