Abstract
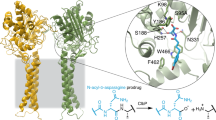
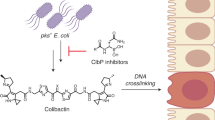
The clb gene cluster encodes the biosynthesis of metabolites known as precolibactins and colibactins. The clb pathway is found in gut commensal Escherichia coli, and clb metabolites are thought to initiate colorectal cancer via DNA crosslinking. Here we report confirmation of the structural assignment of the complex clb product precolibactin 886 via a biomimetic synthetic pathway. We show that an α-ketoimine linear precursor undergoes spontaneous cyclization to precolibactin 886 on HPLC purification. Studies of this α-ketoimine and the related α-dicarbonyl revealed that these compounds are unexpectedly susceptible to nucleophilic cleavage under mildly basic conditions. This cleavage pathway forms other known clb metabolites or biosynthetic intermediates and explains the difficulties in isolating fully mature biosynthetic products. This cleavage also accounts for a recently identified colibactin–adenine adduct. The colibactin peptidase ClbP deacylates synthetic precolibactin 886 to form a non-genotoxic pyridone, which suggests precolibactin 886 lies off the path of the major biosynthetic route.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All the relevant data are available from the authors and/or are included with the manuscript and Supplementary Information.
References
Nougayrède, J.-P. et al. Escherichia coli induces DNA double-strand breaks in eukaryotic cells. Science 313, 848–851 (2006).
Arthur, J. C. et al. Intestinal inflammation targets cancer-inducing activity of the microbiota. Science 338, 120–123 (2012).
Buc, E. et al. High prevalence of mucosa-associated E. coli producing cyclomodulin and genotoxin in colon cancer. PLoS ONE 8, e56964 (2013).
Trautman, E. P. & Crawford, J. M. Linking biosynthetic gene clusters to their metabolites via pathway-targeted molecular networking. Curr. Top. Med. Chem. 16, 1–12 (2015).
Balskus, E. P. Colibactin: understanding an elusive gut bacterial genotoxin. Nat. Prod. Rep. 32, 1534–1540 (2015).
Taieb, F., Petit, C., Nougayrede, J. P. & Oswald, E. The enterobacterial genotoxins: Cytolethal distending toxin and colibactin. EcoSal Plus https://doi.org/10.1128/ecosalplus.ESP-0008-2016 (2016).
Healy, A. R. & Herzon, S. B. Molecular basis of gut microbiome-associated colorectal cancer: a synthetic perspective. J. Am. Chem. Soc. 139, 14817–14824 (2017).
Faïs, T., Delmas, J., Barnich, N., Bonnet, R. & Dalmasso, G. Colibactin: more than a new bacterial toxin. Toxins 10, 151 (2018).
Dejea, C. M. et al. Patients with familial adenomatous polyposis harbor colonic biofilms containing tumorigenic bacteria. Science 359, 592–597 (2018).
Li, Z. R. et al. Divergent biosynthesis yields a cytotoxic aminomalonate-containing precolibactin. Nat. Chem. Biol. 12, 773–775 (2016).
Trautman, E. P., Healy, A. R., Shine, E. E., Herzon, S. B. & Crawford, J. M. Domain-targeted metabolomics delineates the heterocycle assembly steps of colibactin biosynthesis. J. Am. Chem. Soc. 139, 4195–4201 (2017).
Brachmann, A. O. et al. Colibactin biosynthesis and biological activity depend on the rare aminomalonyl polyketide precursor. Chem. Commun. 51, 13138–13141 (2015).
Zha, L., Wilson, M. R., Brotherton, C. A. & Balskus, E. P. Characterization of polyketide synthase machinery from the pks island facilitates isolation of a candidate precolibactin. ACS Chem. Biol. 11, 1287–1295 (2016).
Li, Z.-R. et al. Macrocyclic colibactin induces DNA double-strand breaks via copper-mediated oxidative cleavage. Preprint at bioRxiv https://doi.org/10.1101/530204 (2019).
Dubois, D. et al. ClbP is a prototype of a peptidase subgroup involved in biosynthesis of nonribosomal peptides. J. Biol. Chem. 286, 35562–35570 (2011).
Cougnoux, A. et al. Analysis of structure–function relationships in the colibactin-maturating enzyme ClbP. J. Mol. Biol. 424, 203–214 (2012).
Brotherton, C. A. & Balskus, E. P. A prodrug resistance mechanism is involved in colibactin biosynthesis and cytotoxicity. J. Am. Chem. Soc. 135, 3359–3362 (2013).
Bian, X. et al. In vivo evidence for a prodrug activation mechanism during colibactin maturation. Chembiochem 14, 1194–1197 (2013).
Vizcaino, M. I., Engel, P., Trautman, E. & Crawford, J. M. Comparative metabolomics and structural characterizations illuminate colibactin pathway-dependent small molecules. J. Am. Chem. Soc. 136, 9244–9247 (2014).
Guntaka, N. S., Healy, A. R., Crawford, J. M., Herzon, S. B. & Bruner, S. D. Structure and functional analysis of ClbQ, an unusual intermediate-releasing thioesterase from the colibactin biosynthetic pathway. ACS Chem. Biol. 12, 2598–2608 (2017).
Patonay, T. & Hoffman, R. V. Base-promoted reactions of α-azido ketones with aldehydes and ketones: a novel entry to α-azido-β-hydroxy ketones and 2,5-dihydro-5-hydroxyoxazoles. J. Org. Chem. 60, 2368–2377 (1995).
Healy, A. R., Nikolayevskiy, H., Patel, J. R., Crawford, J. M. & Herzon, S. B. A mechanistic model for colibactin-induced genotoxicity. J. Am. Chem. Soc. 138, 15563–15570 (2016).
Shine, E. E. et al. Model colibactins exhibit human cell genotoxicity in the absence of host bacteria. ACS Chem. Biol. 13, 3286–3293 (2018).
Xue, M., Shine, E., Wang, W., Crawford, J. M. & Herzon, S. B. Characterization of natural colibactin–nucleobase adducts by tandem mass spectrometry and isotopic labeling. Support for DNA alkylation by cyclopropane ring opening. Biochemistry 57, 6391–6394 (2018).
Wilson, M. R. et al. The human gut bacterial genotoxin colibactin alkylates DNA. Science 363, eaar7785 (2019).
Healy, A. R., Vizcaino, M. I., Crawford, J. M. & Herzon, S. B. Convergent and modular synthesis of candidate precolibactins. Structural revision of precolibactin A. J. Am. Chem. Soc. 138, 5426–5432 (2016).
Seashore-Ludlow, B., Torssell, S. & Somfai, P. Addition of azomethine ylides to aldehydes: mechanistic dichotomy of differentially substituted α-imino esters. Eur. J. Org. Chem. 2010, 3927–3933 (2010).
Lou, S., Ramirez, A. & Conlon, D. A. Catalytic syn-selective direct aldol reactions of benzophenone glycine imine with aromatic, heteroaromatic and aliphatic aldehydes. Adv. Synth. Catal. 357, 28–34 (2015).
Alper, P. B., Hung, S.-C. & Wong, C.-H. Metal catalyzed diazo transfer for the synthesis of azides from amines. Tetrahedron Lett. 37, 6029–6032 (1996).
Nyffeler, P. T., Liang, C.-H., Koeller, K. M. & Wong, C.-H. The chemistry of amine–azide interconversion: catalytic diazotransfer and regioselective azide reduction. J. Am. Chem. Soc. 124, 10773–10778 (2002).
Goddard-Borger, E. D. & Stick, R. V. An efficient, inexpensive, and shelf-stable diazotransfer reagent: imidazole-1-sulfonyl azide hydrochloride. Org. Lett. 9, 3797–3800 (2007).
Edwards, O. E. & Purushothaman, K. K. Some reactions of alicyclic α-azidoketones. Can. J. Chem. 42, 712–716 (1964).
Faiz, S. et al. Synthesis and consecutive reactions of alpha-azido ketones: a review. Molecules 20, 14699–14745 (2015).
Dess, D. B. & Martin, J. C. A useful 12-I-5 triacetoxyperiodinane (the Dess–Martin periodinane) for the selective oxidation of primary or secondary alcohols and a variety of related 12-I-5 species. J. Am. Chem. Soc. 113, 7277–7287 (1991).
Vizcaino, M. I. & Crawford, J. M. The colibactin warhead crosslinks DNA. Nat. Chem. 7, 411–417 (2015).
Li, Z. R. et al. Critical intermediates reveal new biosynthetic events in the enigmatic colibactin pathway. Chembiochem 16, 1715–1719 (2015).
White, M. J. & Leeper, F. J. Kinetics of the thiazolium ion-catalyzed benzoin condensation. J. Org. Chem. 66, 5124–5131 (2001).
Kim, T. & Spiegel, D. A. The unique reactivity of N-phenacyl-derived thiazolium salts toward α-dicarbonyl compounds. Rejuvenation Res 16, 43–50 (2012).
Acknowledgements
We thank D. Spiegel for helpful discussions regarding the mechanism of the hydrolytic cleavage of 2. Financial support from the National Institutes of Health (R01GM110506 to S.B.H., 1DP2-CA186575 to J.M.C. and R01CA215553 to S.B.H. and J.M.C.), the Chemistry Biology Interface Training Program (T32GM067543 to K.M.W.), the Charles H. Revson foundation (postdoctoral fellowship to A.R.H.) and Yale University is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
A.R.H., K.M.W. and N.R.L. carried out the synthetic experiments. C.S.K. conducted the purification of precolibactin 886 (1) and ClbP deacylation studies. S.B.H. and J.M.C. oversaw the project. S.B.H. wrote the manuscript. All the authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8, Supplementary Tables 1 and 2, Supplementary synthetic procedures and characterization data (1H, 13C NMR, IR, HRMS) for all new compounds.
Rights and permissions
About this article
Cite this article
Healy, A.R., Wernke, K.M., Kim, C.S. et al. Synthesis and reactivity of precolibactin 886. Nat. Chem. 11, 890–898 (2019). https://doi.org/10.1038/s41557-019-0338-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-019-0338-2