Abstract
Dominance hierarchy is a fundamental phenomenon in grouped animals and human beings, however, the underlying regulatory mechanisms remain elusive. Here, we report that an antisense long non-coding RNA (lncRNA) of synapsin II, named as AtLAS, plays a crucial role in the regulation of social hierarchy. AtLAS is decreased in the prefrontal cortical excitatory pyramidal neurons of dominant mice; consistently, silencing or overexpression of AtLAS increases or decreases the social rank, respectively. Mechanistically, we show that AtLAS regulates alternative polyadenylation of synapsin II gene and increases synapsin 2b (syn2b) expression. Syn2b reduces AMPA receptor (AMPAR)-mediated excitatory synaptic transmission through a direct binding with AMPAR at the postsynaptic site via its unique C-terminal sequence. Moreover, a peptide disrupting the binding of syn2b with AMPARs enhances the synaptic strength and social ranks. These findings reveal a novel role for lncRNA AtLAS and its target syn2b in the regulation of social behaviors by controlling postsynaptic AMPAR trafficking.
Similar content being viewed by others
Introduction
Social hierarchy is a naturally occurring and evolutionarily conserved phenomenon that serves as a guiding principle for the access to resources, such as food, mating opportunities and resting spots, and has profound influences on health and disease in both animals and humans.1,2,3 The establishment of social dominance relies on the well-organization and orchestration of multiple brain regions, including the medial prefrontal cortex (mPFC), hippocampus, amygdala and nucleus accumbens.4 Among them, the mPFC is of particular interest. A functional magnetic resonance imaging (fMRI) study in humans has validated that the dorsal medial prefrontal cortex (dmPFC) is tightly correlated with dominance-related behaviors.2 In the group-housed rodents, mPFC is recognized as a center for social dominance. Activities of the mPFC neurons are up-regulated in the dominant mice,5 and lesions in the mPFC lead to downward social ranking.6 Elevation of synaptic strength in the mPFC by overexpression of AMPA receptors (AMPARs)5 or by optogenetic activation reinforces the social dominance.7 Also, ablation of BDNF/TrkB signals in the inhibitory neurons results in social dominance, which can be reversed by optogenetic silencing of the excitatory neurons in the mPFC.8 Therefore, synaptic activity in the mPFC plays a critical role for social status.
A recent study shows that a prior history of winning is able to reinforce social dominance, known as the “winner effect”,7 indicating the critical role of previous experience in social ranks. It has been shown that epigenetic regulation is essential for transforming such winning experience (environmental stimulus) to the ascendance in social status (phenotype).9,10 However, the detailed molecular regulatory mechanisms underlying social hierarchy remain elusive.
Long non-coding RNAs (lncRNAs), a large and diverse class of RNA transcripts with the length over 200 nucleotides,11 are well known in epigenetic regulation.12,13,14 Due to the diverse genomic loci and the abundant functional modes, lncRNAs have been implicated in multiple biological processes including neurodevelopment, neuronal survival and neurodegeneration.12,15 In addition, alterations of specific lncRNA expression have been implicated in aberrant social behaviors. For instance, MSNP1AS, an antisense lncRNA of the moesin pseudogene 1 gene (MSNP1), is significantly up-regulated in the temporal cortex of autism,16 a well-known neurodevelopmental disorder characterized by impaired social interactions. Knockout of the brain cytoplasmic (BC1) RNA in mice reduces social interaction levels.17 Therefore, it is important to explore whether and how lncRNAs participate in the establishment of social hierarchy.
Results
Expression profiles of lncRNAs in the dmPFC of dominant mice
To investigate the involvement of lncRNAs in the formation of social hierarchy, we first differentiated the mice in social dominant and subordinate ones in the same cage by performing a series of social hierarchy-related behavioral tests including the tube test, visible burrow system and urine marking assays (Supplementary information, Fig. S1a–f) as previously reported.5,18,19 Then, an established lncRNA microarray analysis (Fig. 1a) was applied in RNAs isolated from the dmPFC of both dominant and subordinate groups of mice. We chose this brain region for RNA assay because previous studies in human and rodent have suggested the critical role of dmPFC in dominance hierarchy-related behaviors as described above.2 We found that while most of the known lncRNAs were detected with comparable levels (30,894 of 31,172 lncRNAs), several lncRNAs showed marked differences between groups. We found that in dominant mice, 181 lncRNAs were up-regulated whereas 97 lncRNAs were down-regulated (Supplementary information, Fig. S1a). We then screened out those transcripts that were located within protein-coding genes in the sense orientation to avoid mis-detection of primary mRNAs in the following quantitative polymerase chain reaction (qPCR) assays. After that, we selected 40 lncRNAs: 20 down-regulated and 20 up-regulated with log (fold changes) > 4 for validation (Supplementary information, Fig. S1a, g). Among these candidates, 19 transcripts were found to undergo consistent patterns of changes in all tested samples (Fig. 1b). The result of qPCR showed that AK013786 level was most dramatically altered among mice with different social status (Fig. 1b and Supplementary information, Fig. S1h), and the northern blot revealed that AK013786 was significantly reduced in the dominant mice (Fig. 1c). We then chose AK013786 for further studies. As the genomic location of AK013786 shows that it is an antisense long non-coding RNA of synapsin II, we hereafter refer to it as AtLAS.
a Volcano plot shows the fold change (FC) and significance of lncRNAs from the mPFC area of rank-1 (R1) compared to rank-4 (R4) mice. P values were determined by two-tailed t-test. FC > 4 and P < 0.01 are indicated by red color and dashed line; AK013786/AtLAS is highlighted by a bigger red circle. b Validation of deregulated lncRNAs in high- and low-ranked mice determined by qRT-PCR. Data are means (FC) ± SEM relative to R4 (n = 6). c Northern blot of AtLAS with total RNA from R1 and R4 mice mPFC tissues. 18S rRNA serves as a loading control. The data are representative of at least three independent experiments. d AtLAS expression detected in RACE experiments. Results were repeated in at least three independent experiments. e The distribution of AtLAS in different cell types of mPFC in immunofluorescence experiments using the antibodies for CamKII, GAD, PV, Iba1, GFAP or SST (green) and FISH signals for the AtLAS (red). Scale bar, 20 μm. f Quantification of AtLAS distribution in six types of neurons, n = 17–20 from five mice. g The relative expression profiles of AtLAS in different brain areas from dominant and subordinate mice detected in qRT-PCR. Ob, olfactory bulb; Str, striatum; Hp, Hippocampus, n = 5 mice. Two-tailed t-test, **P < 0.01. Data are presented as means ± SEM.
To get the full sequence information of AtLAS, we performed rapid amplification of cDNA ends (RACE) and found that AtLAS contains a poly-A tail in the 3′ terminus and the sequence of AtLAS is identical to that from the UCSC genome database (Fig. 1d and Supplementary information, Fig S2a, b). In order to verify that AtLAS is a lncRNA, we examined its coding probability in the Coding Potential Assessment Tool (CPAT)20 and Coding Potential Calculator (CPC).21 We found that like most known lncRNAs: Xist, Hotair and Sox2ot, AtLAS scores very low in both algorithms when compared to those known coding transcripts (Supplementary information, Fig. S2c). A further biochemical study confirmed that AtLAS has no coding capacity (Supplementary information, Fig. S2d). To understand the tissue and cellular localization of AtLAS, we performed fluorescence in situ hybridization (FISH) assays by using a specific RNA probe. We observed that AtLAS was localized to the nuclear and cytoplasmic regions (Supplementary information, Fig. S2e) and was distributed in the entire brain (Supplementary information, Fig. S2f, g). However, AtLAS was expressed mainly in the excitatory neurons (Fig. 1e, f) and the reduction of AtLAS in the dominant mice can only be detected in the dmPFC rather than other brain regions (Fig. 1g and Supplementary information, Fig. S2h).
AtLAS is both essential and sufficient for suppressing social dominance
Given the significant reduction of AtLAS in the mPFC of dominant mice, we hypothesized that AtLAS might play a key role in the formation of social hierarchy. To test this, we generated a Cre-recombinase dependent shRNA specific to AtLAS that was introduced by Adeno-associated virus 2/8 (AAV2/8-DIO-sh-AtLAS) according to the previous reports22,23 (Supplementary information, Fig. S3a). We injected the sh-AtLAS and its scrambled control virus (sh-Con) into the dmPFC regions of CamKII-Cre mice24 (Supplementary information, Fig. S3b). By using tube-test ranking, visible burrow system, urine-marking assay and warm spot test, we found that the mice with sh-AtLAS infection displayed higher ranks compared to mice with sh-Con infection (Fig. 2a–d and Supplementary information, Movie S1, 2). The established social hierarchy was stable and remained for at least 30 days (Supplementary information, Fig. S3c, d). To further determine the important role of AtLAS in social hierarchy, we delivered the AAV-packed full-length mouse AtLAS under a CamKII promoter into the dmPFC of dominant mice (Fig. 2e) which were then subjected to another round hierarchy tests. Interestingly, dominant mice with the AtLAS-virus infection were shifted downwardly in the rank while control mice did not have any rank shift (Fig. 2f, g; Supplementary information, Fig. S3e–h). These data strongly demonstrate that AtLAS is essential and sufficient for the formation of social hierarchy.
a Summary of the rank ratio for mice injected with sh-AtLAS or with the sh-Con in the tube tests (n = 18 for each group). χ2 test, linear-linear association, χ2 = 5.531, Ptrend = 0.019. b Representative picture of the urine-marking patterns of sh-Con- and sh-AtLAS-injected mice revealed by ultraviolet light (left). Contingency table shows the number of mice in each category. n = 42 pairs from eight cages (right). χ2 test, χ2 = 5.333, P = 0.004. Gray area represents for dominant mice in two tests, black for subordinate ones, and white for different results. c The correlation analysis for the ranks in the tube tests with average weight-change ranks in visible burrow system (VBS) tests for the mice with the injection of sh-AtLAS (sh-AS) or sh-Con virus. Linear regression, n = 8 cages, r = 0.813, P = 1.59 × 10–8. d Cumulative time spent in the warm spot for four grouped mice previously ranked in the tube tests (top). The correlate analysis for the time spent in the warm spot and the ranks in the tube tests after mice were injected with sh-AtLAS or sh-Con (bottom). Linear regression, n = 8 cages, r = 0.864, P = 1.87 × 10–8. e Schematic diagram illustrating the CaMKII:AtLAS viral construct (top). Representative confocal image shows infection of AtLAS-EGFP into the mPFC (bottom). Scale bars, 200 μm (top), 50 μm (bottom). f Experimental design (top) and example of rank-shift in the tube tests for four grouped mice before and after the injection of AtLAS or EGFP virus (bottom). g Summary graph for the rank-shift in the tube test for mice before and after AtLAS or EGFP injection (n = 8 cages). Wilcoxon-signed rank test, P = 1, 1, 1, 1 for EGFP, P = 0.102, 0.039, 0.023, 0.023 for AtLAS. Data are presented as means ± SEM.
Loss of AtLAS differentially regulates syn2a and syn2b
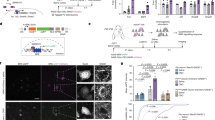
We next aimed to identify the direct targets for AtLAS in the formation of social hierarchy. As described above, AtLAS resides at the negative strand of chromosome 6, on the opposite strand to Synapsin II (Syn2) (Exon 11 of Syn2b and Intron 11 of Syn2a) (Fig. 3a and Supplementary information, Fig. S4a), indicating that it may regulate Syn2 directly via cis effects.25 Indeed, we found that in the mPFC of sh-AtLAS mice, both the protein and mRNA levels of synapsin IIb (syn2b) were decreased, while synapsin IIa levels (syn2a) increased (Fig. 3b, c). In contrast, in mice with AtLAS over-expression, opposite changes on syn2a and syn2b were detected (Supplementary information, Fig. S4b–d). By isolating the nuclear and cytoplasmic fractions with a reported protocol,26 we found that in the dominant mice the loss of AtLAS largely happened in the nuclear region rather than in the cytoplasmic compartment (Fig. 3d), supporting the hypothesis that AtLAS is involved in the events taking place in nuclei such as gene regulation. Given the specific genomic location of AtLAS, we proposed that AtLAS may control syn2a/b ratio by regulating the alternative polyadenylation (APA) of Syn2.
a The genomic view of mouse Syn2 and AtLAS. b, c The mRNA (b) and protein (c) levels of syn2a and syn2b expression after mice were injected with AAV virus carrying sh-AS or scrambled control (Con). n = 4 replicates/group. Two-way ANOVA, post-hoc, Bonferroni, **P < 0.01. Data are presented as means ± SEM. d Nuclear/cytosolic AtLAS ratio from the mPFC areas of R4 (red) and R1 (blue) mice (n = 5, 6). Two-tailed t-test, **P < 0.01. Data are presented as means ± SEM. e Sequence logo of CELF4 recognition motif 1 identified from MEME analysis of pre-syn2 iClip sequence read clusters. f IClip data indicate three binding peaks within the syn2b pre-mRNA 3′UTR and two conserved CELF4 binding motifs. g The HEK293T cells were transfected with pre-syn2 only or combined with AtLAS, then the cell lysates were immuno-precipitated with CELF4 antibody. The pellets were subjected to PCR assay. The results were repeated in 3 independent experiments. h RNA pull-down analysis shows that CELF4 is a pre-syn2 binding protein detected in silver-staining (left) and western blot (right). n = 3 independent experiments. i PCR validates the changes in the alternative splicing of Syn2 target-exons upon AtLAS or/and CELF4 transfection in HEK293T cell overexpressing pre-syn2 (n = 4 independent experiments). j REMSA experiment using recombinant proteins CELF4 or/and AtLAS combined with syn2b 3′UTR (including the alternative splicing site). k Generation of the two mutants of pre-syn2 (Mut1 and Mut2) by deleting the binding sites of AtLAS to CELF4 (shown in red). The specific sequences of AMO1 and AMO2 are also listed. l PCR validates the changes in the alternative splicing of Syn2 target-exons upon co-expression of CELF4 with wild-type (WT) or Mut1/2 pre-syn2 in HEK293T cells. (n = 4 independent experiments). m PCR validates the changes in the alternative splicing of Syn2 target-exons upon transfection of CELF4 with AMO1 or/and AMO2 together with WT pre-syn2 in HEK293T cells.
AtLAS regulates the alternative polyadenylation of synapsin II by CELF4
Alternative polyadenylation (APA) is an RNA-processing mechanism that generates distinct 3′termini on mRNAs and regulated by RNA-binding proteins (RBPs).27,28 By using high stringency level and conservation filter in RBPmap,29 74 RBPs were predicted to contain potential binding sites within exon 11 (the first alternative polyadenylation site) of Syn2 (Supplementary information, Table S1). We then examined the iCLIP data (Supplementary Information, Table S2) from mouse database of CLIPbp2 and found that only four proteins: CELF4 (CUGBP, ELAV-like family member 4), RNA-binding protein Fus, Srrm4 (Serine/arginine repetitive matrix protein 4) and TDP-43 (TAR DNA-binding protein 43)30 bind to Syn2. Furthermore, only CELF4 and TDP-43 have the binding peaks in exon 11 of Syn2 (Supplementary information, Fig. S4e) and we found two conserved binding motifs for CELF4 in the 3′-UTR region of syn2b (Fig. 3e, f; Supplementary information, Fig. S4f). We therefore chose CELF4 and TDP-43 for further biological experiments. DNA sequence from exon 11 to exon 12 of Syn2 (Chr 6: 115274196-115278334, mm10, in which the polyadenylation site was included, named pre-syn2) was cloned (Supplementary information, Fig. S4e), and it was transfected together with CELF4 or TDP-43 into the HEK293T cells. We found that only CELF4 transfection induced the up-regulation of syn2a and down-regulation of syn2b (Supplementary information, Fig. S4g). To confirm the direct binding of CELF4 with pre-syn2, we performed the RNA immunoprecipitation (RIP) with CELF4 antibody and mass spectrometry after RNA pull-down with a probe targeting pre-syn2. We found that CELF4 indeed physically interacted with pre-syn2 both in vivo and in vitro (Fig. 3g and Supplementary information, Fig. S4h). No significant difference in CELF4 expression was found from R1 and R4 animals brain areas (Supplementary information, Fig. S4i). Although there were a lot bands existing in the pellets precipitated with the pre-syn2 probe, we detected two sequences identical to the CELF4 protein, which was indicated at the band of 52 kDa (Fig. 3h and Supplementary information, Fig. S4j). In AtLAS-overexpressing cells, the binding affinity of CELF4 to pre-syn2 was reduced (Fig. 3g). Additionally, forced expression of CELF4 dramatically decreased the syn2b level (Fig. 3i), while silencing of CELF4 (Supplementary information, Fig. S4k) restored the syn2b level in the cells with sh-AtLAS transfection (Supplementary information, Fig. S4l). Moreover, we incubated the recombinant CELF4 protein with in-vitro transcribed syn2b 3′-UTR with or without AtLAS and then detected their binding affinity by electrophoretic mobility shift assay (EMSA) according to a well-established protocol.31 We found that CELF4 can only bound with single strand syn2b 3′-UTR but not the paired sense-antisense RNAs formed between the AtLAS and syn2b 3′-UTR (Fig. 3j). To determine a specific sequence in pre-syn2 that binds with CELF4, we generated two mutants of pre-syn2 by deleting the two predicted binding sites of CELF4 (Mut1 and Mut2) (Fig. 3k) and found that CELF4 overexpression did not alter the syn2a/b ratio in Mut1-expressing cells (Fig. 3l). We then synthesized two different antisense 2′-O-methoxyethyl modified oligonucleotides32,33 1 and 2 (AMO1 and AMO2) (Fig. 3k) to target two different regions within the deleted region of Mut1. We found that AMO2 disrupted the regulation of syn2a/b ratio by CELF4, indicating that the sequence of AMO2 (Chr 6: 115275158-115275182) was important for the binding of CELF4 with pre-syn2 (Fig. 3m). Collectively these findings suggest that CELF4 binds with the 3′-UTR of syn2b and retards the APA of syn2b, which in turn promotes the syn2a isoform expression. LncRNA AtLAS alters syn2a/2b ratio by the regulation on CELF4 binding to syn2b and the following APA event.
Syn2b but not syn2a participates in the social dominance
As syn2b is positively and syn2a is negatively regulated by AtLAS in vitro, we then queried their contribution to social hierarchy regulated by AtLAS. From qPCR and western blot results, we found that in dominant mice the mRNA and protein levels of syn2b were significantly decreased, whereas opposite changes were detected for syn2a. Also, these changes were observed in mPFC but not other brain regions (Fig. 4a, b; Supplementary information, Fig. S5a–c). In comparison, no apparent difference was found in synapsin1 (syn1), another important member of the synapsin family (Supplementary information, Fig. S5d). Synapsin3 (syn3) was not examined because it mainly exists during the developmental stage.34 In the dominant mice, forced expression of syn2b (Fig. 4c–e; Supplementary information, Fig. S5e–h and Movie S3) but not silenced syn2a (Supplementary information, Fig. S5i–l) in the dmPFC induced a downward shift in the rank. In the subordinate mice, silencing syn2 elevated the rank35 (Supplementary information, Fig. S6a–c), but overexpression of syn2a had no effect (Supplementary information, Fig. S6d, e).
a, b The protein levels of syn2 isoforms from the mPFC of R4 and R1 mice were detected in western blot (a) and their quantification in (b) (n = 3). Two-way ANOVA, post-hoc, Bonferroni, **P < 0.01. Data are presented as means ± SEM. c Schematic diagram shows the syn2b-mCherry virus construct (top) and representative images for virus infection (bottom) in the dmPFC. Scale bars, 200 μm (top), 50 μm (bottom). d, e Rank-shift of grouped mice in the tube tests before and 30 days after syn2b-mCherry (Syn2b) or mCherry control virus (mCherry) injection (n = 8 cages for each group). R1 and R2 mice were randomly selected for the syn2b-virus injection, and other mice for mCherry control virus injection. Wilcoxon matched-pairs signed rank test, P = 1, 1, 1 for mCherry, P = 0.02, 0.007, 0.007 for syn2b. f The diagram shows the design of syn2b-KD mice by using CRISPR-Cas9 system to delete its 3′UTR with two specific sgRNAs. g Analysis of the effect of syn2b-KD on the expression of syn2 isoforms. Data are representative of three independent experiments. +/−, heterozygous; −/−, homozygous mice. Two-way ANOVA, post-hoc, Bonferroni, **P < 0.01. h Summary of rank ratio in the tube tests for mice from the same cage (2 of C57 and 2 of syn2b-KD mice). n = 8 cages for each test; χ2 test, linear-linear association, χ2 = 4.747, Ptrend = 0.029. i The correlation analysis for the ranks in the tube test with the ranks in the warm spot test for C57 and syn2b-KD mice. Linear regression, n = 8 cages, r = 0.90, P = 2.42 × 10−12.
To confirm the role of syn2b in social hierarchy, we generated syn2b-knockdown mice (syn2b-KD) by deletion of the 3′-UTR region of syn2b with CRISPR/Cas9 strategy36 (Fig. 4f; Supplementary information, Fig. S6f). Syn2b-KD mice had reduced syn2b and higher syn2a expression in the mPFC (Fig. 4g), and had comparable brain volume and body sizes to the WT littermates (Supplementary information, Fig. S6g, h). In the rotarod test, open field test and elevated plus maze task, the syn2b-KD mice showed similar performances to C57 mice (Supplementary information, Fig. S6i–l), suggesting the normal development and locomotor ability in those mice. However, the syn2b-KD mice displayed higher hierarchy relative to the WT littermates (Fig. 4h, i). These results strongly indicate a critical role of syn2b down-regulation in mediating the formation of social dominance.
Loss of AtLAS/syn2b pathway reinforces the AMPAR membrane trafficking
It has been reported that dominant mice had a larger amplitude in miniature excitatory synaptic currents (mEPSCs), which are mediated by the a-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) subtype of glutamate receptors, compared to subordinate mice in layer V pyramidal neurons of the mPFC.5 The mPFC may function as a central regulator in linking its activity to the downstream brain areas, including basolateral nucleus of the amygdala (BLA),2,37 hypothalamus38 and periaqueductal gray (PAG).39,40 Thus, the excitatory synaptic strength in layer V pyramidal neurons which as the main output neurons of mPFC plays an important role in the social dominance.5 Given the reduction of AtLAS occurs mainly in the excitatory neurons in the dmPFC, we explored alterations of synaptic activities by whole-cell patch clamp in the dmPFC slices (Fig. 5a). We found that compared to the control mice, the sh-AtLAS mice displayed larger amplitudes in miniature excitatory synaptic currents (mEPSCs) (Fig. 5b–d; Supplementary information, Fig. S7a) but comparable miniature inhibitory synaptic currents (mIPSCs) (Supplementary information, Fig. S7b). Moreover, sh-AtLAS neurons had higher ratio of AMPAR-/NMDAR-mediated currents (Fig. 5e, Supplementary information, Fig. S7c) but not paired-pulse facilitation ratio (Supplementary information, Fig. S7d, e), which were observed in the R1 vs R4 mice as previously reported.5 Consistently, overexpression of AtLAS (Fig. 5f–h; Supplementary information, Fig. S7f) or syn2b (Fig. 5i–k; Supplementary information, Fig. S7g) led to a reduction in mEPSC amplitudes in the dmPFC neurons of the dominant WT mice. These results indicated the involvement of postsynaptic AMPARs in AtLAS/syn2b-dependent regulation of synaptic efficacy. In addition, we expressed sh-AtLAS in vivo in the hippocampal CA1 region of juvenile mice, and recorded whole-cell currents from individual pyramidal neurons in acute slices 20–36 days after the injection.41 We found that the mEPSC amplitudes were increased in neurons expressing sh-AtLAS (Fig. 5l, m). These findings suggest that changes in postsynaptic AMPARs may underlie the AtLAS/syn2b-induced alterations in synaptic efficacy.
a A representative neuron image in the patch clamp. Scale bar, 10 μm. b–d Representative traces of mEPSCs (b) from mPFC neurons infected with sh-Con or sh-AtLAS virus. The mean mEPSC amplitudes (c) and representative cumulative distribution of mEPSC amplitudes (d) were analyzed. n = 30 neurons from five mice for each group. Two-tailed t-test, **P < 0.01. Data are presented as means ± SEM. e The AMPA to NMDA current ratio was altered in the mPFC neurons from mice infected with sh-AtLAS. Representative traces from whole-cell voltage-clamp experiments shows NMDAR- and AMPAR-mediated currents recorded in a mPFC pyramidal neuron from mice infected with sh-Con (red) or sh-AtLAS (blue). Two-tailed t-test, **P < 0.01. f–h Representative traces of mEPSCs (f) from mPFC neurons infected with full-length AtLAS or control (EGFP). The mean mEPSC amplitudes (g) and representative cumulative distribution of mEPSC amplitudes (h) were analyzed. n = 25 neurons from four mice for each group, means ± SEM, two-tailed t-test, **P < 0.01. i–k Representative traces of mEPSCs (i) from mPFC neurons infected with coding sequence of syn2b or control (mCherry). The mean mEPSCs amplitudes (j) and representative cumulative distribution of mEPSC amplitudes (k) were analyzed. n = 27 neurons from five mice for each group. Means ± SEM, two-tailed t-test, **P < 0.01. l, m Representative EGFP expression (l) of the sparse lenti-virus infection in hippocampal CA1 slices after the delivery of sh-Con or sh-AtLAS constructs, the representative traces and quantitative analysis of mEPSC are shown in (m). Scale bar, 20 μm. n = 32 neurons from five mice for each group. Two-tailed t-test, **P < 0.01.
Syn2b physically associates with AMPARs via its C-terminus
We then queried how syn2b, a previously known presynaptic protein,42 could regulate the synaptic function via postsynaptic AMPARs. By purifying different fractions of synaptic components, we detected that syn2a and syn2b were localized at both the presynaptic and postsynaptic compartments (Fig. 6a–c), which is in line with several previous proteomic studies.43,44,45 Although the distinct roles for syn2a vs syn2b have not been well studied, syn2a, but not syn2b, was shown to play a fundamental role in synaptic vesicle mobilization and reserve pool regulation in presynaptic nerve terminals.46 We hypothesized that loss of syn2b may regulate the social hierarchy via postsynaptic mechanisms, independent of syn2a. Indeed, only syn2b was found to have the physical association with GluA1 and GluA2 subunits of AMPARs (Fig. 6d–f). Both subunits were increased in the membrane fraction of the dmPFC in sh-AtLAS mice (Fig. 6g) and in dominant WT mice (Fig. 6h), but not NR2A, NR2B subunits (Supplementary information, Fig. S7h). To identify the specific site responsible for the interaction of syn2b-AMPAR, we transfected the common sequence of synapsin II (c-syn2, aa 1–458 of syn2a and syn2b) (Supplementary information, Fig. S7i, j), the coding sequence of syn2b (full length: aa 1–479) (Fig. 6i, j), or syn2a (full length: aa 1–586) (Supplementary information, Fig. S7k–l) with different subunits of AMPARs (GluA1, GluA2) into HEK293 cells. We found that only syn2b was able to interact with GluA1 and GluA2 (Fig. 6i, j; Supplementary information, Fig. S7i–l). The identical sequence of syn2a and syn2b is about 85.62%, indicating a key role of the specific C-terminal sequence (the unique 21aa sequence) of syn2b in mediating the binding with AMPARs. We then attempted to identify the key residues in syn2b critical for the binding with AMPARs. To this end, we generated mutations at two sites in syn2b (Q461R, D465H) by referring to the two SNPs (rs751232089 and rs2289706 from dbSNP database: https://www.ncbi.nlm.nih.gov/snp) in human syn2b genes (Supplementary information, Fig. S7m). However, the two mutations failed to disrupt the binding of syn2b with GluA1/A2 (Supplementary information, Fig. S7n, o). More efforts will be made in future to reveal the details of how syn2b interacts with AMPARs.
a, b The different synaptic components were extracted from the mPFC and then subjected to immunoblot (a) using the antibodies as indicated and the quantitative analysis (b, n = 5). Synaptophysin (sypt) was used as a presynaptic marker, and PSD-95 as a postsynaptic marker. β-actin serves as the loading control. c The PFC primary neurons were cultured to DIV12 and immunolabeled with syn2 and drebrin, a postsynaptic marker. Scale bar, 3 μm. d–f Immunoblot shows the interactions between AMPAR subunits (GluA1 and GluA2) and syn2b in the mPFC homogenates detected by co-IP. n = 3 independent experiments. g Levels of cell-surface GluA1/2/4 from the mPFC areas of mice injected with sh-Con or sh-AtLAS virus (top) were quantitatively analyzed (bottom). Means ± SEM, two-tailed t-test, **P < 0.01. h Western blot assays indicate the surface expression of GluA1, GluA2 and GluA4 in the mPFC of R4 and R1 mice. Means ± SEM, two-tailed t-test, **P < 0.01. i, j Co-IP shows the interactions between EGFP-syn2b and GluA1 (i) or GluA2 (j) in HEK293T cells 48 h after the co-transfection. These data were repeated in at least three independent experiments.
Disrupting the syn2b-AMPAR binding enhances synaptic efficacy and up-shifts social hierarchy
To determine the significance of syn2b interaction with AMPARs, we designed a membrane-permeable peptide (P-2B) by fusing the TAT sequence47,48 to the C-terminus of syn2b, and confirmed that P-2B but not control scramble peptide (S-2B) was able to disrupt the binding of syn2b to AMPARs (Fig. 7a–c). Importantly, incubation of cultured prefrontal cortex neurons with P-2B resulted in the increase in AMPAR expression at the plasma membrane (Fig. 7d) and the larger amplitude of mEPSCs (Fig. 7e–h) compared to the S-2B treatment. Furthermore, mice intraperitoneally injected with P-2B for consecutive 10 days displayed higher ranks than animals receiving scrambled control peptide (Fig. 7i). Thus, loss of syn2b-AMPAR interaction enhanced AMPARs membrane trafficking at the postsynaptic sites, which in turn led to the establishment of social dominance.
a A diagram illustrates that P-2B disrupts the binding of syn2b to AMPAR and prompts GluA1/GluA2 presence at synapses. b, c Co-IP shows the interactions between GluAs with syn2b after cells treated with P-2B (P) or S-2B (S). The HEK293T cells were transfected with GluA1 or GluA2 and EGFP-syn2b. 12 h post transfection, the cells were treated with S-2B or P-2B (10 μM) every 12 h for three times, then co-IP was performed using these cell lysates. d The mice were injected with P-2B or S-2B at the dose of 10 mg/kg for 10 days (intraperitoneal). The membrane fraction of mPFC homogenates were purified for western blot analysis by using the antibodies indicated. Representative immunoblots (right) and quantification (left). n = 4 for each group. Two-tailed t-test, *P < 0.05; **P < 0.01. e–h Relative representative traces (e), summary of the relative mean and representative cumulative distribution of amplitudes (f, g), and mean frequencies (h) of mEPSC from mPFC neurons treated with P-2B or S-2B. n = 30 neurons from six mice for each group. Two-tailed t-test, **P < 0.01. i Rank percentage in the tube test after mice treated with P-2B or S-2B for 10 days (10 mg/kg, intraperitoneal injection, n = 8 cages for each group). χ2 test, linear-linear association χ2 = 4.747, Ptrend = 0.029.
Discussion
Social dominance is a ubiquitous behavioral feature in both humans and other species. It has been well recognized that the mPFC serves as the central regulatory circuitry in the neural network.40 Two populations of neurons in mPFC were found to innervate the mediobasal hypothalamus (MBH) and the lateral hypothalamus (LH) separately. Stimulation of mPFC terminals in MBH increased bite counts in conflicts while stimulation of mPFC terminals in LH resulted in violent bites.38 These “quantitative and qualitative” aspects of aggressive behavior mediated by the mPFC-hypothalamus system are important for establishing social dominance. In addition, activation of mPFC-dPAG (dorsal periaqueductal gray) neural circuit by photo-stimulation of dopamine terminals or using electrochemical approaches could drive place avoidance and defensive behaviors,39 both of which are important for social status. While the underlying cellular and molecular mechanisms remain elusive, recent studies suggest that both the intrinsic and extrinsic factors,49 acting at genetic50 and epigenetic51,52 levels, play important roles in the regulation of social dominant behaviors and social status determination. Previous reports have revealed that genetic deletion of some critical molecules, including the sodium/potassium-transporting ATPase subunit alpha-3 (Atp1a3),53 progranulin54 and vasopressin 1b receptor,55 leads to shifting of social rank, suggesting the involvement of genetic factors in regulating social hierarchy. In line with this, global increases in DNA methylation has been shown to cause marked elevation in social hierarchy in African cichlid fish Astatotilapia burtoni,51 indicating the critical role of epigenetic regulation in the determination of social ranks. LncRNAs acting as key epigenetic regulators have significant influence on learning and memory, adaptive behavior in mice,56,57 but their roles in social hierarchy are not clear. In the present study, we found that loss of AtLAS in the excitatory neurons in mPFC is both required and sufficient for the establishment of social dominance in grouped mice. Loss of AtLAS in the CamKII-positive neurons of the mPFC resulted in an increase in synaptic strength of the excitatory neurons. As higher synaptic activity was observed in the mPFC neurons of dominant mice, the loss of AtLAS and the consequential hyperactivity of those neurons may underlie the higher social ranks for these mice. While the majority (~75%) of AtLAS-positive cells were CamKII-positive, we found that AtLAS was also present in other cell types. Thus, in future studies it will be important to explore the potential involvement of other AtLAS-positive cell types in establishing social hierarchy by employing cell-type-specific knockdown systems.
Further, we explored the potential downstream effectors for AtLAS and found that the suppression of AtLAS led to the inhibition of syn2b 3′UTR adenylation, which further resulted in the decrease of syn2b with an increase of syn2a by blocking the binding of pre-syn2b mRNA with CELF4, a protein important in co-transcriptional and post-transcriptional of RNA processing.58 We found two conserved CELF4 binding motifs (a U/G-rich motif and a U/C-rich motif59) that localized in the 3′UTR of syn2b, which was consistent with a previous report that CELF4 preferred to bind mRNAs in the 3′UTR.60 CELF4 is expressed predominantly in the excitatory neurons of the hippocampus and the cerebral cortex60 and most of its targets are highly involved in synaptic functions.60 In this study, we showed that the binding of CELF4 with syn2 pre-mRNA retarded the polyadenylation in the exon 11 of syn2b, which in turn promoted syn2a expression. In the presence of AtLAS, the formation of RNA-RNA duplex between AtLAS and syn2 pre-mRNA precluded the binding of CELF4 with syn2 pre-mRNA, facilitating the production of syn2b isoform. Whether CELF4 participates in the APA has not yet been validated, but several reports suggested this possibility. Muscleblind-like (MBNL) protein, a splicing factor sharing the same binding motif with CELF4, regulated the APA at developmental stages.61 CELF1, another important member of CELF family, directly regulates APA during T cell activation.62
We also found that the reduction of syn2b led to social dominance. This finding was consistent with previous studies showing that syn2b was a functional marker of submissive behaviors.63 Moreover, we demonstrated that syn2b binds with the GluA1 and GluA2 subunits of AMPARs at postsynaptic compartmenst and restrains the membrane insertion of functional AMPARs at excitatory synapses.41 Our findings on postsynaptic syn2 are consistent with several previous reports of proteomic studies.43,44,45 As a widely recognized presynaptic protein,42 the indispensable role for syn2b, but not syn2a, in the regulation of postsynaptic receptor trafficking reported here sheds new light in our understanding of the differential functions of these two isoforms. Syn2a and syn2b share over 85% homology (domain A, B, C, G), and the distinct sequence in the syn2b C-terminus determines its binding with AMPARs. These findings reveal the critical role of domain I,64 the unique domain in syn2b of synapsin family, in mediating AMPAR membrane insertion. It is valuable to further explore the detailed regulatory mechanisms in domain I-mediated syn2b postsynaptic localization and AMPAR interaction in the future. AMPAR trafficking, which is implicated in neuronal circuit formation and behavioral modification,65,66 is expected to play an important role in the regulation of transmission strength of the efferent synapses from the mPFC to other brain regions. It is known that the membrane insertion or internalization of AMPARs is tightly correlated with synaptic strength, synaptic plasticity, and the related behaviors.66 In the mPFC, upregulation of AMPAR by injection of GluA4-expressing virus5 or isolation-induced decrease of AMPARs in neonatal mice67 led to changes in the social ranks in mice. It has also been reported that phosphorylation of AMPARs by stress or fluoxetine,68 and the allosteric modulation of AMPARs69 are crucial for the social dominance.
Finally, we generated a membrane permeable peptide (P-2B) that contains the C-terminus of syn2b to disrupt the direct binding of syn2b with AMPARs. We found that application of P-2B led to a marked increase in the membrane expression of AMPARs and the strength of excitatory synaptic activity, as well as in the social hierarchy ranks. As the weakened synaptic activity in the mPFC is tightly correlated with many other neuropsychiatric disorders in animals and humans,70,71,72 the peptide used in this study may serve as a potential strategy for enhancing the related synaptic transmission deficiencies in these disorders. We found that specific alteration in AtLAS expression and the subsequent AtLAS-syn2b-AMPAR axis in the mPFC regulated social hierarchy. While P-2B could specifically disrupt syn2b-AMPAR interaction, it is still an open question whether it can selectively target the mPFC via vein injection in mice. However, as we demonstrated here, i.p. injection of P-2B improved animals performance in the social dominance tests, which leads us to propose that P-2B may be able to reinforce synaptic activity in many brain regions, including those that directly project to the mPFC.7
In summary, we find that loss of lncRNA AtLAS decreases the syn2b expression by promoting the CELF4 binding and inhibiting the polyadenylation in the social dominant animals. Decreased association of syn2b with AMPAR via its C-terminus promotes AMPARs membrane expression and enhances synaptic strength in the dmPFC, leading to the superior social hierarchy ranks (Fig. 8).
In subordinate mice, AtLAS binds with the pre-mRNA of syn2 and regulates the syn2a/b ratio by inhibiting CELF4-mediated alternative splicing, which leads to the APA and the increased expression of syn2b. Syn2b binds with AMPAR via its unique C-terminus to restrain the membrane insertion of AMPAR, the weakened synaptic strength in the mPFC renders social submissive roles in these animals. Conversely, in dominant mice, loss of AtLAS and the reduction of syn2b pose less inhibition on AMPAR membrane expression, and then enhance the synaptic strength in the mPFC and social ranks. The effect can also be achieved by the application of P-2B to disturb syn2b/AMPAR interaction.
Materials and methods
Mice
The syn2b 3′UTR-Knockout (syn2b-KD) mouse, including heterozygous (syn2b+/−) and homozygous (syn2b−/−) mouse was generated using CRISPR-Cas9 system (Bioray Laboratories Inc., Shanghai, China). Two sgRNAs, GTAAAAACGAAGGGTACACGGGG and ACCCAGCCATGTCACCGCAAAGG were designed to specifically knock out the sequence of syn2b but not syn2a. Genotyping for syn2b-KD mice was performed by multiplex PCR using a pair of F (5′-acagtctggtgcagttcga-3′) and R (5′-ctaagtgccgtcatctgtg-3′) primers. The genetic background of mutant mice is C57BL/6.
All animal experiments were carried out according to the “Policies on the Use of Animals and Humans in Neuroscience Research” revised and approved by the Society for Neuroscience in 1995. All the animals were housed (four per cage) in an air-conditioned room (22 ± 2 °C, 12-h light and 12-h dark cycle) with food and water ad libitum, bred in the Experimental Animal Central of Tongji Medical College, Huazhong University of Science and Technology. The behavior tests were performed in their active hours. Prior to testing, mice were acclimated to the test room for at least 0.5 h. Adult male C57BL/6 mice aged 10–12 weeks and weighing 22–28 g were used in experiments.
Microarray-based lncRNA expression profiling
To obtain a genome-wide view of lncRNA expression in social hierarchy, mPFC samples isolated from mouse brain of rank1 and rank4 identified by tube-test ranking, visible burrow system and urine marking assays. These samples were subjected to global lncRNA profiling using the Arraystar Mouse LncRNA Microarray v3.0 which allows a simultaneous detection of 35,923 lncRNAs and 24,881 coding transcripts. Agilent Microarray Scanner (Agilent p/n G2565BA) and Agilent Feature Extraction (v11.0.1.1) were used to scan and analyze acquired array images. Using GeneSpring GX v12.1 software package (Agilent Technologies) for quantile-normalization and subsequent data processing. Detailed processing program was conducted as previously described73,74). Differentially expressed lncRNAs were identified through filtering on “Fold Change” and there were 278 lncRNA found to have differentially expressed transcripts (FC > 4.0, P < 0.01). Then we excluded lncRNAs that were annotated as “sense overlap” and selected top deregulated lncRNAs (20 transcripts with the highest and 20 with the lowest fold change) for further analysis.
RACE of full-length AtLAS
RNA extraction from R1 and R4 mouse brains was prepared using TRIzol reagent following the manufacturer’s instructions. The synthesis of 5′ and 3′ RACE cDNA libraries were performed following the manufacturer’s instructions (SMARTer RACE 5′/3′ Kit, Takara) with a modified 3′ RACE reverse transcription primer. RACE amplification products were analyzed with agarose gel electrophoresis and Sanger sequencing. Primers designed for RACE and PCR are presented in Supplementary information, Table S3.
Isolation of nuclear and cytoplasmic RNAs
mPFC brain tissues from the R1 and R4 mice were washed with ice-cold PBS twice. After centrifugation at 1000× g for 5 min, supernatants were removed. Cytoplasmic and nuclear RNA were isolated with a cytoplasmic and nuclear RNA isolation kit (Norgen) following the manufacturer’s protocol.26
REMSA
The assay was performed according to the previously described protocol.75 In brief, RNA was labeled with desthiobiotinylation using the Pierce RNA 3′ End Desthiobiotinylation Kit (Thermo). His-tagged CELF4 were purified with His-tag Protein Purification Kit (Beyotime). RNA was denatured at 95 °C for 1 min and then placed on ice before use. RNA and CELF4 mixed together in RNA-protein binding buffer (25 mM Tris-HCl, pH 7.5, 1 mM MgCl2, 100 mM KCl, 1 mM dithiothreitol), and incubated at room temperature for 30 min. Reactions were stopped with loading buffer and the final products were loaded into 4% polyacrylamide gel in 0.5× TBE. The RNA and protein were transferred to nylon membrane. After the UV-light crosslinking, the membranes were blocked and detected by chemiluminescence.
RNA isolation and qRT-PCR assay
Total RNA was isolated from cultured cells or brain tissues, samples were extracted by TRIzol reagent following the manufacturer’s instruction. 1 µg of total RNA used to synthesize cDNA by Reverse Transcription Kit (Toyobo life science). The qRT-PCR with SYBR Green PCR Master Mix (Takara) was implemented on CFX96 Real-Time PCR Detection System (Bio-Rad). Relative gene expression was analyzed by 2−ΔΔCt normalized to GAPDH. The primers designed are listed in Supplementary Information, Table S3.
RNA pull-down assay
Cell fractionations of nuclear and cytoplasmic components were isolated from treated 293T cells or from mPFC tissues, the cytoplasmic fraction or nuclear pellets were resuspended in lysis buffer (25 mM Tris, pH 7.4, 150 mM KCl, 5 mM EDTA, 5 mM MgCl2, 1% NP-40, 0.5 mM DTT with a protease and RNAse inhibitor). One volume of lysis buffer was added with 0.1 μg/μL tRNA and biotinylated RNA. The mixture was incubated at 37 °C for 15 min, at room temperature for 15 min and at 4 °C for 6 h to overnight. Then streptavidin beads were added to the mix and incubated for 1h at 4 °C. The beads were washed 5 times (10 min each) with 1 ml of the lysis buffer on a magnetic rack. The beads were boiled with SDS buffer for western blot assays and mass spectrometric analysis.
RNA immunoprecipitation
RNA immunoprecipitation was performed as previously described.76 Cells were lysed with RIP buffer (100 mM KCl, 5 mM MgCl2,10 mM HEPES pH 7.0, 0.5% NP-40, and 1 mM Dithiothrectol) with RNase and protease inhibitors for 30 min on ice. Cell debris was removed by centrifugation at 4 °C. Cell lysate pre-bound with protein G dynal beads before respective antibodies were added for 2 h at RT. 3 µg of anti-CELF4 (Santa) antibody was used in immunoprecipitation to pull down the associated RNAs. Co-immunoprecipitated RNAs were extracted using TRIzol reagent (Invitrogen), and cDNA was synthesized with the ReverTra RT reagent Kit (Toyobo, FSQ-101) and then analyzed by qRT-PCR. The primers used in RIP-qPCR analysis are listed in Supplementary information, Table S3 and the amount of immunoprecipitated RNAs was compared to the input.
Northern blot
Single-stranded RNA probe was generated using DIG Northern Starter Kit (Roche) according to the manufacturer’s protocol. For agarose gel electrophoresis, the RNA was mixed with two volume of denaturing loading buffer (20 mM MOPS, pH 7.0, 5 mM sodium acetate, 2 mM EDTA, 50% formamide, 6% formaldehyde and 10% glycerol) and samples were denatured at 65 °C for 10 min and then immediately placed on ice. Denatured RNA samples were loaded onto 2% agarose MOPS (20 mM MOPS, pH 7.0, 5 mM sodium acetate and 2 mM EDTA) gels containing 2% formaldehyde. RNA was transferred to positively charged nylon membranes (Roche) by capillary electrophoresis. RNA was fixed to the membrane by UV cross-linking. Blots were hybridized overnight at 60 °C with a DIG-labeled antisense AtLAS RNA probe according to the manufacturer's instructions.
FISH
Primary culture cells were used for RNA FISH analysis. RNA-FISH assays were performed according to the manufacture’s protocol provided by Stellaris FISH Probes (RiboBio) as described.77 Briefly, after the fixation in 4% paraformaldehyde cells were permeabilized in 1× PBS with 0.5% Triton X-100 (Sigma) for 5 min at 4 °C. Cells were blocked by adding pre-hybridization buffer at 37 °C for 30 min. Hybridization was carried out with a FISH probe in a moist chamber at 37 °C away from light more than 10 h using Ribo™ Fluorescent In Situ Hybridization Kit (RiboBio). Cells were washed three times with Wash Buffer I (4× SSC with 0.1% Tween-20, Sigma), then with Wash Buffer II (2× SSC, Sigma) and Wash Buffer III (1× SSC, Sigma) at 42 °C in the dark for 5 min. After the final rinse, cells were stained with DAPI in the dark for 10 min. AtLAS FISH probes were designed and synthesized by RiboBio Co., Ltd. Mouse 18S FISH probes (RiboBio) were used as the cytoplasmic controls. All images were obtained with a confocal microscope (ZEISS, LSM 780). FISH on mouse brain slice (10 μm) was carried out according to the previously reported protocol.78
Plasmids and viruses
GluA1, GluA2 and GluA4 plasmids were kindly provided by Dr Yu Tian Wang (University of British Columbia, Vancouver, British Columbia, Canada). Adeno-associated viruses (AAVs) for Syn2b, Syn2a, AtLAS and short hairpin RNA for AtLAS, CELF4 were purchased from Obio technology (Shanghai, China). Lentivirus for short hairpin RNA of Syn2, Syn2a and AtLAS (for CA1 injection) were purchased from Genechem (Shanghai, China). AMOs for the CELF4 was synthesized by Umibio Co. Ltd. (Shanghai, China).
Stereotaxic surgeries
Syn2b-KD or CamKII-Cre or WT mice (about 6- or 12-week old) were anesthetized with a mixture of ketamine (100 mg/kg) and dexmedetomidine (0.5 mg/kg). The mouse brain was mounted by stereotaxic apparatus (RWD life science). Viral injections were targeted mPFC area by using coordinates according to the Paxinos and Franklin mouse brain atlas.79 The stereotaxic coordinates7 were AP, +2.43 mm; ML, ±0.28 mm; DV, −1.81 mm, angled 14° toward the midline in the coronal plane. For electrophysiology recording experiments or behavioral tests, packaged virus in volumes 500 nL was injected bilaterally into the mPFC. Before behavioral tests, mice were allowed 3–4 weeks recovery from the virus injections. Injection sites were confirmed at the end of the behavior experiments and incorrect injection placement were excluded from data analysis.
Behavioral assays
Tube test
We followed the previously reported protocol.5,7 In brief, before the tube test was performed, 4 mice with similar age and body weights were placed together in a cage for at least 4 weeks. Following training, paired mice encounter every other mouse of the group using a round robin test. A transparent plexiglas tube of 30 cm length and 3 cm inner diameter was used for training and tests. During training, each mouse was given ten trials to move forward out of a plexiglas tube on each side for 2 consecutive days. The tube was cleaned with 75% ethanol solution between each trial to remove odor, urine or feces. During the test trials, two mice were held by the tail and released simultaneously at the opposite ends of the tube when they reached the middle of the tube. The mouse that retreated to the end of tube as the “loser” scored 0, and the other as the “winner” scored 1. Each mouse was ranked by their cumulative scores that could vary from 0–3. Only mice with stable ranks (at the same rank position for over 3 or 6 days) were used for further tests.
Visible burrow system (VBS)
The test was performed as previously described,5 which were adjusted from Arakawa et al.19 The apparatus consists of a rectangular arena (61 cm × 61 cm) with a series of transparent tunnels and three opaque chambers that was illuminated with a 12:12 h light/dark cycle. Food and water are relatively difficult to retrieve by the narrow ramp, and only one animal is allowed to access the resources at a time. During the test, four mice in one cage were housed in a VBS for 7 successive days. Weight change of each mouse was calculated for rank order in the group.
Territory urine marking assay
By a round-robin design, four male mice of a cage were tested pairwise for 3 days (2 pairs /day). In a two-chamber cage (26 cm × 21 cm × 26 cm), mice were placed in pairs at the opposite sides which were separated by a perforated plexiglas. Sheets of filter paper were arranged below each side of cage to collect urine deposited from mice. After two hours, mice were returned to their homecage and the filter papers were analyzed by a UV light source. The number and/or size of the urine marks as well as distance from the divider were analyzed blindly for rank order.
Warm spot test
The test was performed as described previously.18 In brief, a cage of grouped mice were taken into a cold cage (28 cm × 20 cm) for 30 min which floor was cooled down to 0 °C. Then they were transferred to the testing cold cage for 20 min where a nest with the diameter of 5 cm heated by a underneath coil at 34 °C in one corner. Behaviors were video-taped and the occupation time of the warm nest were respectively analyzed.
Open field test and rotarod test
Each individual mouse was placed in the open field arena (60 × 60 × 60 cm) in a fixed corner and allowed to move freely for 10 min. The behaviors were monitored by photo-beam detectors. The data were collected via a computer and were analyzed using the MED associates’ Activity Monitor Data Analysis software. Between sessions, the maze was cleaned with 75% ethanol and dried with paper towels. Time spent in center square and total distances traveled in all the areas were analyzed. Rotarods were used to measure the motor coordination of mice. The rotarod (AccuScan Instruments) measures the ability of the mouse to maintain balance on a motor-driven, rotating rod. Thus, the fore and hind limb motor coordination and balance can be analyzed. C57BL/6J or syn2b-KD mice were tested on the rotarod at 3 months of age.80
Elevated plus maze
The elevated plus maze contains two open arms, two closed arms, and a center. Two closed arms are 30.5 cm above the ground. At the start of the test, mice were placed in the center and allowed to explore the apparatus for 5 min. The movement traces of mice were analyzed by tracking software to measure the time spent in the openarms.
Electrophysiological recordings
Mice were sacrificed by isoflurane anesthesia followed by decapitation and the brain was quickly dissected into ice-cold ACSF (in mM: 3 KCl, 1.25 NaH2PO4, 26 NaHCO3, 7 MgCl2, 212 sucrose and 10 glucose) saturated with carbogen gas (95% oxygen, 5% carbon dioxide). 300 μm coronal brain slices containing mPFC were prepared from Vibratome. Slices were recovered in a holding chamber containing ACSF saturated with carbogen gas at 32 °C for 30 min and then at room temperature for 1 h before being transferred to the recording chamber. Recording was made from layer V pyramidal neurons of the mPFC from Bregma: +1.7 to +2.2 mm coronal slices. Brain slices were made and recorded from one group of mice within a day.
mEPSC/mIPSC recording and analysis
For mEPSC recordings, neurons were held at −70 mV in voltage-clamp mode and were recorded using an internal solution containing 140 mM potassium gluconate, 10 mM HEPES, 0.2 mM EGTA, 2 mM NaCl, 2 mM MgATP and 0.3 mM NaGTP, and an external solution containing 10 µM bicuculline, 1 µM tetrodotoxin (TTX) and 50 µM APV.5,81 Events were filtered at 2 kHz, and were acquired at 10 kHz (Molecular Devices, Sunnyvale, CA, USA) in conjunction with pClamp 10.2 software. Currents were recorded in 10 s epochs for a total duration of at least 200 s per recording. mEPSC events were analyzed using MiniAnalysis (Synaptosoft, Decatur, GA). mEPSC events detection criteria included an amplitude greater than 5 pA, a minimum rise rate of 0.3 pA/ms, and a decay time constant between 1–12 ms. Cells with less than 75 mEPSC events were excluded from averaged mEPSC waveform analyses.
Recordings of mIPSC82 were made in the presence of 1 µM TTX and 10 µM CNQX. Recordings of mIPSCs were made using a KCl–rich intracellular solution containing the following (in mM): 118 KCl, 9 EGTA, 10 HEPES, 4 MgCl2, 1 CaCl2, 4 NaATP, 0.3 NaGTP and 14 creatinine phosphate (pH 7.2 with KOH).
AMPAR EPSCs were recorded with cells held at −70 mV. We measured the rectification of AMPAR currents by evoking a pharmacologically isolated AMPAR EPSC at holding potentials ranging from −80 to +60 mV. For each cell at each stimulating intensity tested, six consecutive EPSCs were recorded and the peak amplitudes were averaged. NMDAR EPSCs were recorded at +60 mV in order to remove the voltage-dependent Mg2+ block in the presence of 10 µM CNQX and the stimulation protocol was the same to the one evoking AMPAR EPSCs. Current-voltage relationships for AMPAR and NMDAR EPSCs were performed and AMPA/NMDA ratio was calculated. Data were analyzed with Clampfit 10.0 and Sigmaplot 12.5 software. Recordings were performed by the experimenters who were kept blind to the group ranks.
Paired pulse paradigm
An interpulse interval of 50 ms was used in two stimuli of a paired pulse paradigm. Five pairs of stimuli were delivered with an interval of 5 s between each pair and the peak amplitudes of both EPSCs were measured. The paired pulse ratio (PPR) was measured by the peak amplitude ratios of the first and second responses.
Crude synaptosomal preparation (S2, P2, PSD)
mPFC samples were dissected from mice, then synaptosomes were prepared as described previously.83,84,85 Briefly, brain samples were homogenized in ice-cold HEPES-buffered sucrose (0.32 M sucrose, 4 mM HEPES, pH 7.4) with freshly added protease inhibitor cocktail tablets (Roche), then centrifuged at 1000× g for 10 min to yield the nuclear enriched pellet and the S1 fraction. The S1 fraction was centrifuged (12,000× g, 20 min) to obtain supernatant (S2, microsomes and cytosol) and pellet (P2, crude synaptosomal membranes) fractions. The pellet was resuspended in HEPES buffer (4 mM HEPES, 1 mM EDTA, pH 7.4), then centrifuged at 12,000× g for 20 min. Resuspension and centrifugation were repeated once. The resulting pellet was resuspended in buffer A (20 mM HEPES, 100 mM NaCl, 0.5% Triton, pH 7.2) and rotated slowly following the centrifugation at 12,000× g for 20 min. The supernatant contained non-PSD (peri-/extrasynaptic and presynaptic, PeP) membranes. The pellet was resuspended in buffer B (20 mM HEPES, 0.15 mM NaCl, 1% Triton-X, 1% deoxycholic acid, 1% SDS, 1 mM DTT, pH 7.5), and was gently rotated (1 h) and centrifuged (10,000× g, 15 min). The final supernatant containing PSD fraction was collected. Purified samples were stored at −80 °C until use.
Co-immunoprecipitation (Co-IP), protein isolation and western blot assays
In brief, cells were collected and lysed in RIPA buffer (Beyotime, P0013). After quantification using the BCA protein assay kit (ThermoFisher, #23225), 1 mg of total extracted protein was incubated with 2 µg of antibodies overnight. Normal rabbit IgG used as a negative IP control. Then the mixtures were incubated with protein A/G agarose beads for another 4 h, washed at least 4 times, and were boiled for 10 min in SDS sample buffer (Bio-Rad, #161-0737). Cell lysates for an input control were also treated with the equal volume of SDS buffer.
For Membrane Extraction, membrane and cytoplasmic protein fractions of cultured cells or brain tissues were obtained with a Mem-PER Plus Membrane Protein Extraction Kit (Pierce Protein Biology) according to the manufacturer’s protocol.86 The purified membrane proteins were resolved in SDS-PAGE under denaturing conditions and were transferred onto PVDF membranes (Bio-Rad, #162-0177). The membranes were blocked in 1× PBST-5% non-fat milk for 2 h and incubated overnight with primary antibodies at 4 °C, followed by the incubation with secondary antibodies for 1 h. Specific bands were visualized with Odyssey infrared imaging systems (Gene Company Limited). The intensity of the bands were quantified using the gel quantification plugin on ImageJ software. Absolute values were normalized to the intensity of loading control (GAPDH/β-actin/cadherin) bands from the same samples.
Administration of peptides
The mice were intraperitoneally injected with 10 mg/kg TAT-P-2B (TAT-CLQYILDCNGIAVGPKQVQAS) or the scrambled control peptide TAT-S-2B (TAT-CLYASQDLIQVKAGIPGQNCV) for 10 days. The peptides with 95.79% purity were synthesized by ChinaPeptides (Shanghai, China). The TAT-P-2B or TAT-S-2B were numbered, and the animal handlers were kept blind to the injected peptide kinds in all experiments.87
Quantification and statistical analysis
Statistical analyses (listed in Supplementary Information, Table S4) were performed using GraphPad Prism (GraphPad Software, Inc., CA, USA) and SPSS version 16.0 (SPSS Inc., Chicago, IL). Data are presented as means ± SD. Student’s t-test and one-way measures ANOVAs followed by Bonferroni post-hoc tests were used to make the single-variable comparisons. The χ2 test (Fisher’s exact test, linear-linear association) and Wilcoxon matched-pairs signed rank test were used to compare rank classes and the change in two groups. The Pearson’s product-moment correlation coefficient, the Spearman’s rank correlation coefficient or two-way ANOVA (post-hoc, Bonferroni) was used to analyze correlation between two variables. P < 0.05 was taken to indicate statistical significance, *P < 0.05 and **P < 0.01.
Change history
11 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41422-021-00492-y
References
Sapolsky, R. M. The influence of social hierarchy on primate health. Science 308, 648–652 (2005).
Zink, C. F. et al. Know your place: neural processing of social hierarchy in humans. Neuron 58, 273–283 (2008).
Nagy, M., Akos, Z., Biro, D. & Vicsek, T. Hierarchical group dynamics in pigeon flocks. Nature 464, 890–893 (2010).
Singer, T. The past, present and future of social neuroscience: a European perspective. NeuroImage 61, 437–449 (2012).
Wang, F. et al. Bidirectional control of social hierarchy by synaptic efficacy in medial prefrontal cortex. Science 334, 693–697 (2011).
Holson, R. R. Mesial prefrontal cortical lesions and timidity in rats. III. Behavior in a semi-natural environment. Physiol. Behav. 37, 239–247 (1986).
Zhou, T. et al. History of winning remodels thalamo-PFC circuit to reinforce social dominance. Science 357, 162–168 (2017).
Tan, S. et al. Postnatal TrkB ablation in corticolimbic interneurons induces social dominance in male mice. Proc. Natl. Acad. Sci. USA 115, E9909–E9915 (2018).
Dias, B. G., Maddox, S., Klengel, T. & Ressler, K. J. Epigenetic mechanisms underlying learning and the inheritance of learned behaviors. Trends Neurosci. 38, 96–107 (2015).
Bludau, A., Royer, M., Meister, G., Neumann, I. D. & Menon, R. Epigenetic regulation of the social brain. Trends Neurosci. 7, 471–484 (2019).
Wilusz, J. E., Sunwoo, H. & Spector, D. L. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev. 23, 1494–1504 (2009).
Ng, S. Y., Lin, L., Soh, B. S. & Stanton, L. W. Long noncoding RNAs in development and disease of the central nervous system. Trends Genet. 29, 461–468 (2013).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Lee, J. T. Epigenetic regulation by long noncoding RNAs. Science 338, 1435–1439 (2012).
Briggs, J. A., Wolvetang, E. J., Mattick, J. S., Rinn, J. L. & Barry, G. Mechanisms of long non-coding RNAs in mammalian nervous system development, plasticity, disease, and evolution. Neuron 88, 861–877 (2015).
Wilkinson, B. & Campbell, D. B. Contribution of long noncoding RNAs to autism spectrum disorder risk. Int. Rev. Neurobiol. 113, 35–59 (2013).
Briz, V. et al. The non-coding RNA BC1 regulates experience-dependent structural plasticity and learning. Nat. Commun. 8, 293 (2017).
Drickamer, L. C. Urine marking and social dominance in male house mice (Mus musculus domesticus). Behavioural Process. 53, 113–120 (2001).
Arakawa, H., Blanchard, D. C. & Blanchard, R. J. Colony formation of C57BL/6J mice in visible burrow system: identification of eusocial behaviors in a background strain for genetic animal models of autism. Behavioural Brain Res. 176, 27–39 (2007).
Wang, L. et al. CPAT: coding-potential assessment tool using an alignment-free logistic regression model. Nucleic Acids Res. 41, e74 (2013).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 35, W345–W349 (2007).
Hitz, C., Wurst, W. & Kuhn, R. Conditional brain-specific knockdown of MAPK using Cre/loxP regulated RNA interference. Nucleic Acids Res. 35, e90 (2007).
Xu, H. et al. A disinhibitory microcircuit mediates conditioned social fear in the prefrontal cortex. Neuron 102, 668–682 e665 (2019).
Paddison, P. J. et al. Cloning of short hairpin RNAs for gene knockdown in mammalian cells. Nat. Methods 1, 163–167 (2004).
Mattick, J. S. & Gagen, M. J. The evolution of controlled multitasked gene networks: the role of introns and other noncoding RNAs in the development of complex organisms. Mol. Biol. Evol. 18, 1611–1630 (2001).
Alarcon, C. R. et al. HNRNPA2B1 Is a Mediator of m(6)A-Dependent Nuclear RNA Processing Events. Cell 162, 1299–1308 (2015).
Tian, B. & Manley, J. L. Alternative polyadenylation of mRNA precursors. Nat. Rev. Mol. Cell Biol. 18, 18–30 (2017).
Chen, M. & Manley, J. L. Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat. Rev. Mol. Cell Biol. 10, 741–754 (2009).
Paz, I., Kosti, I., Ares, M. Jr., Cline, M. & Mandel-Gutfreund, Y. RBPmap: a web server for mapping binding sites of RNA-binding proteins. Nucleic Acids Res. 42, W361–W367 (2014).
Yang, Y. C. et al. CLIPdb: a CLIP-seq database for protein-RNA interactions. BMC Genom. 16, 51 (2015).
Rio, D. C. Electrophoretic mobility shift assays for RNA-protein complexes. Cold Spring Harb. Protoc. 2014, 435–440 (2014).
Bauman, J. A., Li, S. D., Yang, A., Huang, L. & Kole, R. Anti-tumor activity of splice-switching oligonucleotides. Nucleic Acids Res. 38, 8348–8356 (2010).
Wang, F. et al. SPSB1-mediated HnRNP A1 ubiquitylation regulates alternative splicing and cell migration in EGF signaling. Cell Res. 27, 540–558 (2017).
Ferreira, A. et al. Developmental expression, subcellular localization, and role in axon formation. J. Neurosci. 20, 3736–3744 (2000).
Michetti, C. et al. The knockout of synapsin II in mice impairs social behavior and functional connectivity generating an ASD-like phenotype. Cereb. Cortex 27, 5014–5023 (2017).
Schwartz, M. L., Bruce, J., Shneidman, P. S. & Schlaepfer, W. W. Deletion of 3'-untranslated region alters the level of mRNA expression of a neurofilament light subunit transgene. J. Biol. Chem. 270, 26364–26369 (1995).
Senn, V. et al. Long-range connectivity defines behavioral specificity of amygdala neurons. Neuron 81, 428–437 (2014).
Biro, L. et al. Task division within the prefrontal cortex: distinct neuron populations selectively control different aspects of aggressive behavior via the hypothalamus. J. Neurosci. 38, 4065–4075 (2018).
Vander Weele, C. M. et al. Dopamine enhances signal-to-noise ratio in cortical-brainstem encoding of aversive stimuli. Nature 563, 397–401 (2018).
Wang, F., Kessels, H. W. & Hu, H. The mouse that roared: neural mechanisms of social hierarchy. Trends Neurosci. 37, 674–682 (2014).
Wu, D. et al. Postsynaptic synaptotagmins mediate AMPA receptor exocytosis during LTP. Nature 544, 316–321 (2017).
Han, H. Q., Nichols, R. A., Rubin, M. R., Bahler, M. & Greengard, P. Induction of formation of presynaptic terminals in neuroblastoma cells by synapsin IIb. Nature 349, 697–700 (1991).
Dejanovic, B. et al. Changes in the synaptic proteome in tauopathy and rescue of tau-induced synapse loss by C1q antibodies. Neuron 100, 1322–1336 e1327 (2018).
Peng, J. et al. Semiquantitative proteomic analysis of rat forebrain postsynaptic density fractions by mass spectrometry. J. Biol. Chem. 279, 21003–21011 (2004).
Trinidad, J. C., Thalhammer, A., Specht, C. G., Schoepfer, R. & Burlingame, A. L. Phosphorylation state of postsynaptic density proteins. J. Neurochem. 92, 1306–1316 (2005).
Gitler, D., Cheng, Q., Greengard, P. & Augustine, G. J. Synapsin IIa controls the reserve pool of glutamatergic synaptic vesicles. J. Neurosci. 28, 10835–10843 (2008).
Vives, E., Brodin, P. & Lebleu, B. A truncated HIV-1 Tat protein basic domain rapidly translocates through the plasma membrane and accumulates in the cell nucleus. J. Biol. Chem. 272, 16010–16017 (1997).
Fan, X., Jin, W. Y., Lu, J., Wang, J. & Wang, Y. T. Rapid and reversible knockdown of endogenous proteins by peptide-directed lysosomal degradation. Nat. Neurosci. 17, 471–480 (2014).
Zhou, T., Sandi, C. & Hu, H. Advances in understanding neural mechanisms of social dominance. Curr. Opin. Neurobiol. 49, 99–107 (2018).
Kooij, M. & Sandi, C. The genetics of social hierarchies. Curr. Opin. Behav. Sci. 2, 52–57 (2015).
Lenkov, K., Lee, M. H., Lenkov, O. D., Swafford, A. & Fernald, R. D. Epigenetic DNA methylation linked to social dominance. PLoS One 10, e0144750 (2015).
Cummins, D. D. How the social environment shaped the evolution of mind. Synthese 122, 3–28 (2000).
Sugimoto, H., Ikeda, K. & Kawakami, K. Atp1a3-deficient heterozygous mice show lower rank in the hierarchy and altered social behavior. Genes Brain Behav. 17, e12435 (2018).
Arrant, A. E., Filiano, A. J., Warmus, B. A., Hall, A. M. & Roberson, E. D. Progranulin haploinsufficiency causes biphasic social dominance abnormalities in the tube test. Genes Brain Behav. 15, 588–603 (2016).
Caldwell, H. K., Dike, O. E., Stevenson, E. L., Storck, K. & Young, W. S. 3rd Social dominance in male vasopressin 1b receptor knockout mice. Hormones Behav. 58, 257–263 (2010).
Kocerha, J., Dwivedi, Y. & Brennand, K. J. Noncoding RNAs and neurobehavioral mechanisms in psychiatric disease. Mol. Psychiatry 20, 677–684 (2015).
Li, D. et al. Activity dependent LoNA regulates translation by coordinating rRNA transcription and methylation. Nat. Commun. 9, 1726 (2018).
Dasgupta, T. & Ladd, A. N. The importance of CELF control: molecular and biological roles of the CUG-BP, Elav-like family of RNA-binding proteins. Wiley Interdiscip. Rev. RNA 3, 104–121 (2012).
Ray, D. et al. A compendium of RNA-binding motifs for decoding gene regulation. Nature 499, 172–177 (2013).
Wagnon, J. L. et al. CELF4 regulates translation and local abundance of a vast set of mRNAs, including genes associated with regulation of synaptic function. PLoS Genet. 8, e1003067 (2012).
Batra, R. et al. Loss of MBNL leads to disruption of developmentally regulated alternative polyadenylation in RNA-mediated disease. Mol. Cell 56, 311–322 (2014).
Beisang, D., Reilly, C. & Bohjanen, P. R. Alternative polyadenylation regulates CELF1/CUGBP1 target transcripts following T cell activation. Gene 550, 93–100 (2014).
Nesher, E. et al. Synapsin IIb as a functional marker of submissive behavior. Sci. Rep. 5, 10287 (2015).
Monaldi, I. et al. The highly conserved synapsin domain E mediates synapsin dimerization and phospholipid vesicle clustering. Biochemical J. 426, 55–64 (2010).
Van den Oever, M. C. et al. Prefrontal cortex AMPA receptor plasticity is crucial for cue-induced relapse to heroin-seeking. Nat. Neurosci. 11, 1053–1058 (2008).
Kessels, H. W. & Malinow, R. Synaptic AMPA receptor plasticity and behavior. Neuron 61, 340–350 (2009).
Tada, H. et al. Neonatal isolation augments social dominance by altering actin dynamics in the medial prefrontal cortex. Proc. Natl. Acad. Sci. USA 113, E7097–E7105 (2016).
Park, M. J., Seo, B. A., Lee, B., Shin, H. S. & Kang, M. G. Stress-induced changes in social dominance are scaled by AMPA-type glutamate receptor phosphorylation in the medial prefrontal cortex. Sci. Rep. 8, 15008 (2018).
Tatsukawa, T. et al. Scn2a haploinsufficient mice display a spectrum of phenotypes affecting anxiety, sociability, memory flexibility and ampakine CX516 rescues their hyperactivity. Mol. Autism 10, 15 (2019).
Subramaniam, K. et al. Computerized cognitive training restores neural activity within the reality monitoring network in schizophrenia. Neuron 73, 842–853 (2012).
Cho, J. H., Deisseroth, K. & Bolshakov, V. Y. Synaptic encoding of fear extinction in mPFC-amygdala circuits. Neuron 80, 1491–1507 (2013).
Hadar, R. et al. Early neuromodulation prevents the development of brain and behavioral abnormalities in a rodent model of schizophrenia. Mol. Psychiatry 23, 943–951 (2018).
Viereck, J. et al. Long noncoding RNA chast promotes cardiac remodeling. Sci. Transl. Med. 8, 326ra322 (2016).
Shi, Y. & Shang, J. Long noncoding RNA expression profiling using arraystar LncRNA microarrays. Methods Mol. Biol. 1402, 43–61 (2016).
Smirnov, A., Wang, C., Drewry, L. L. & Vogel, J. Molecular mechanism of mRNA repression in trans by a ProQ-dependent small RNA. EMBO J. 36, 1029–1045 (2017).
Ng, S. Y., Johnson, R. & Stanton, L. W. Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. EMBO J. 31, 522–533 (2012).
Guo, X. et al. A Linc1405/Eomes complex promotes cardiac mesoderm specification and cardiogenesis. Cell Stem Cell 22, 893–908 (2018).
Baleriola J., Jean Y., Troy C., Hengst U. Detection of axonally localized mRNAs in brain sections using high-resolution in situ hybridization. J Vis. Exp. 100, e52799 (2015).
Paxinos G., and Franklin, K. B. J. The mouse brain in stereotaxic coordinates. 3rd edn (Academic Press, New York, 2008).
Li, X. et al. Serotonin receptor 2c-expressing cells in the ventral CA1 control attention via innervation of the Edinger-Westphal nucleus. Nat. Neurosci. 21, 1239–1250 (2018).
Yang, X. et al. A novel mechanism of memory loss in Alzheimer's disease mice via the degeneration of entorhinal-CA1 synapses. Mol. Psychiatry 23, 199–210 (2018).
Williams, C. et al. Coactivation of multiple tightly coupled calcium channels triggers spontaneous release of GABA. Nat. Neurosci. 15, 1195–1197 (2012).
He, S., Ma, J., Liu, N. & Yu, X. Early enriched environment promotes neonatal GABAergic neurotransmission and accelerates synapse maturation. J. Neurosci. 30, 7910–7916 (2010).
Sarkar, A. & Kabbaj, M. Sex differences in effects of ketamine on behavior, spine density, and synaptic proteins in socially isolated rats. Biol. Psychiatry 80, 448–456 (2016).
Pei, L. et al. A novel mechanism of spine damages in stroke via DAPK1 and Tau. Cereb. Cortex 25, 4559–4571 (2015).
Phuchareon, J., McCormick, F., Eisele, D. W. & Tetsu, O. EGFR inhibition evokes innate drug resistance in lung cancer cells by preventing Akt activity and thus inactivating Ets-1 function. Proc. Natl. Acad. Sci. USA 112, E3855–E3863 (2015).
Su, Y. et al. MicroRNA-26a/death-associated protein kinase 1 signaling induces synucleinopathy and dopaminergic neuron degeneration in Parkinsonʼs disease. Biol. Psychiatry 85, 769–781 (2019).
Acknowledgements
We thank Dr Yun-Yun Han at Dept. Neurobiology of HUST, Dr Yu-Tian Wang at UBC, Dr Ling-Ling Chen at CEMCS, CAS of China, Dr Qiang Liu at UTSC for their constructive comments; Dr Yu-Tian Wang at UBC for the GluA1/2/4 plasmids. This study is supported partially by the National Natural Science Foundation of China (81871108, 81829002, 81961128005, 81761138043, 91632114, 81771150, 31571039, 31721002), Top-Notch Young Talents Program of China of 2014, and Academic Frontier Youth Team of Huazhong University of Science and Technology to L.Q.Z.
Author information
Authors and Affiliations
Contributions
L.Q.Z. and D.L. initiated and designed the study. L.Q.Z., D.L., Y.L, and H.Y.M. supervised the study. M.M., W.X., and F.H. performed the molecular biological experiments and animal experiments. M.F.D. and X.H. performed the electrophysiological recording. M.M., W.X., F.H., J.G.C, Y.L., D.L. and L.Q.Z. analyzed the data. L.Q.Z. and H.Y.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The peptide used to block the binding of syn2b with AMPAR has been submitted to the Patent Office of the Peopleʼs Republic of China by L.Q.Z., D.L., W.X., M.M., X.H. (Application No. 201811638624.0). No other conflicts of interests are declared for all authors.
Supplementary information
Rights and permissions
About this article
Cite this article
Ma, M., Xiong, W., Hu, F. et al. A novel pathway regulates social hierarchy via lncRNA AtLAS and postsynaptic synapsin IIb. Cell Res 30, 105–118 (2020). https://doi.org/10.1038/s41422-020-0273-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-020-0273-1
This article is cited by
-
Cortical astrocytes modulate dominance behavior in male mice by regulating synaptic excitatory and inhibitory balance
Nature Neuroscience (2023)
-
Correlated evolution of social organization and lifespan in mammals
Nature Communications (2023)
-
Techniques for investigating lncRNA transcript functions in neurodevelopment
Molecular Psychiatry (2023)
-
Multiomic profiling of the acute stress response in the mouse hippocampus
Nature Communications (2022)
-
miR-135a-5p mediates memory and synaptic impairments via the Rock2/Adducin1 signaling pathway in a mouse model of Alzheimer’s disease
Nature Communications (2021)