Abstract
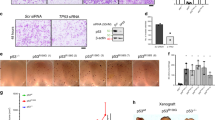
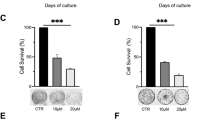
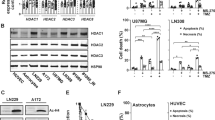
Various inhibitors of histone deacetylase (HDAC) activity can sensitize drug resistant cancer cells to chemotherapeutic agents. However, the mechanisms underlying such effects of distinct HDAC inhibitors (HDACi) remain poorly understood. Here we show that both the HDACi trichostatin A and valproic acid induced a sensitization of multidrug-resistant cancer cells to the topoisomerase II inhibitor etoposide/VP16. This effect was associated with increased acetylation of certain lysines on histones H3 and H4, including lysine 16 on histone H4 (H4K16). Overexpression of the histone acetyltransferase hMOF, known to target H4K16, was sufficient to mimic HDACi treatment on sensitization and H4K16 acetylation, and importantly, small-interfering RNA (siRNA)-mediated knockdown of hMOF abolished the HDACi-mediated sensitizing effects as well as the increase in H4K16 acetylation. Conversely, siRNA-mediated knockdown of the H4K16 deacetylase SIRT1 mimicked HDACi treatment whereas overexpression of SIRT1 abolished H4K16 acetylation and significantly reduced the sensitizing effects of HDACi. Interestingly, the effects of hMOF on H4K16 acetylation and sensitization to the topoisomerase II inhibitor could be directly counteracted by exogenous expression of increasing amounts of SIRT1 and vice versa. Our study results suggest that hMOF and SIRT1 activities are critical parameters in HDACi-mediated sensitization of multidrug-resistant cancer cells to topoisomerase II inhibitor and increased H4K16 acetylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Abbreviations
- EX-527:
-
6-chloro-2,3,4,9-tetrahydro-1H-carbazole-1-carboxamide;
- DAPI:
-
4′,6-diamidino-2-phenylindole
- VP16:
-
etoposide
- GFP:
-
green fluorescent protein
- HAT:
-
histone acetyltransferase
- HDAC:
-
histone deacetylase
- HDACi:
-
HDAC inhibitors
- H4K16:
-
lysine 16 on histone H4
- MOF:
-
males absent on the first
- NSCLC:
-
non-small-cell lung carcinoma
- siRNA:
-
small-interfering RNA
- TSA:
-
trichostatin A
- VPA:
-
valproic acid
References
Andersson T, Sodersten E, Duckworth JK, Cascante A, Fritz N, Sacchetti P et al. (2009). CXXC5 is a novel BMP4-regulated modulator of Wnt signaling in neural stem cells. J Biol Chem 284: 3672–3681.
Bolden JE, Peart MJ, Johnstone RW . (2006). Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov 5: 769–784.
Castedo M, Perfettini JL, Roumier T, Andreau K, Medema R, Kroemer G . (2004). Cell death by mitotic catastrophe: a molecular definition. Oncogene 23: 2825–2837.
Chen WY, Wang DH, Yen RC, Luo J, Gu W, Baylin SB . (2005). Tumor suppressor HIC1 directly regulates SIRT1 to modulate p53-dependent DNA-damage responses. Cell 123: 437–448.
Deng CX . (2009). SIRT1, is it a tumor promoter or tumor suppressor? Int J Biol Sci 5: 147–152.
Drummond DC, Noble CO, Kirpotin DB, Guo Z, Scott GK, Benz CC . (2005). Clinical development of histone deacetylase inhibitors as anticancer agents. Annu Rev Pharmacol Toxicol 45: 495–528.
Fraga MF, Ballestar E, Villar-Garea A, Boix-Chornet M, Espada J, Schotta G et al. (2005). Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nat Genet 37: 391–400.
Gallego MA, Joseph B, Hemstrom TH, Tamiji S, Mortier L, Kroemer G et al. (2004). Apoptosis-inducing factor determines the chemoresistance of non-small-cell lung carcinomas. Oncogene 23: 6282–6291.
Glozak MA, Sengupta N, Zhang X, Seto E . (2005). Acetylation and deacetylation of non-histone proteins. Gene 363: 15–23.
Hajji N, Wallenborg K, Vlachos P, Nyman U, Hermanson O, Joseph B . (2008). Combinatorial action of the HDAC inhibitor trichostatin A and etoposide induces caspase-mediated AIF-dependent apoptotic cell death in non-small cell lung carcinoma cells. Oncogene 27: 3134–3144.
Hermanson O, Glass CK, Rosenfeld MG . (2002). Nuclear receptor coregulators: multiple modes of modification. Trends Endocrinol Metab 13: 55–60.
Huffman DM, Grizzle WE, Bamman MM, Kim JS, Eltoum IA, Elgavish A et al. (2007). SIRT1 is significantly elevated in mouse and human prostate cancer. Cancer Res 67: 6612–6618.
Ikura T, Ogryzko VV, Grigoriev M, Groisman R, Wang J, Horikoshi M et al. (2000). Involvement of the TIP60 histone acetylase complex in DNA repair and apoptosis. Cell 102: 463–473.
Jepsen K, Solum D, Zhou T, McEvilly RJ, Kim HJ, Glass CK et al. (2007). SMRT-mediated repression of an H3K27 demethylase in progression from neural stem cell to neuron. Nature 450: 415–419.
Jin Q, Yan T, Ge X, Sun C, Shi X, Zhai Q . (2007). Cytoplasm-localized SIRT1 enhances apoptosis. J Cell Physiol 213: 88–97.
Joseph B, Ekedahl J, Lewensohn R, Marchetti P, Formstecher P, Zhivotovsky B . (2001). Defective caspase-3 relocalization in non-small cell lung carcinoma. Oncogene 20: 2877–2888.
Joseph B, Marchetti P, Formstecher P, Kroemer G, Lewensohn R, Zhivotovsky B . (2002). Mitochondrial dysfunction is an essential step for killing of non-small cell lung carcinomas resistant to conventional treatment. Oncogene 21: 65–77.
Karagiannis TC, El-Osta A . (2006). Modulation of cellular radiation responses by histone deacetylase inhibitors. Oncogene 25: 3885–3893.
Keshelava N, Seeger RC, Groshen S, Reynolds CP . (1998). Drug resistance patterns of human neuroblastoma cell lines derived from patients at different phases of therapy. Cancer Res 58: 5396–5405.
Kyrylenko S, Kyrylenko O, Suuronen T, Salminen A . (2003). Differential regulation of the Sir2 histone deacetylase gene family by inhibitors of class I and II histone deacetylases. Cell Mol Life Sci 60: 1990–1997.
Liu T, Liu PY, Marshall GM . (2009). The critical role of the class III histone deacetylase SIRT1 in cancer. Cancer Res 69: 1702–1705.
Minucci S, Pelicci PG . (2006). Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nat Rev Cancer 6: 38–51.
Motta MC, Divecha N, Lemieux M, Kamel C, Chen D, Gu W et al. (2004). Mammalian SIRT1 represses forkhead transcription factors. Cell 116: 551–563.
Napper AD, Hixon J, McDonagh T, Keavey K, Pons JF, Barker J et al. (2005). Discovery of indoles as potent and selective inhibitors of the deacetylase SIRT1. J Med Chem 48: 8045–8054.
Neal KC, Pannuti A, Smith ER, Lucchesi JC . (2000). A new human member of the MYST family of histone acetyl transferases with high sequence similarity to Drosophila MOF. Biochim Biophys Acta 1490: 170–174.
Nyman U, Vlachos P, Cascante A, Hermanson O, Zhivotovsky B, Joseph B . (2009). Protein kinase C-dependent phosphorylation regulates the cell cycle-inhibitory function of the p73 carboxy terminus transactivation domain. Mol Cell Biol 29: 1814–1825.
Pfister S, Rea S, Taipale M, Mendrzyk F, Straub B, Ittrich C et al. (2008). The histone acetyltransferase hMOF is frequently downregulated in primary breast carcinoma and medulloblastoma and constitutes a biomarker for clinical outcome in medulloblastoma. Int J Cancer 122: 1207–1213.
Pruitt K, Zinn RL, Ohm JE, McGarvey KM, Kang SH, Watkins DN et al. (2006). Inhibition of SIRT1 reactivates silenced cancer genes without loss of promoter DNA hypermethylation. PLoS Genet 2: e40.
Schneider R, Grosschedl R . (2007). Dynamics and interplay of nuclear architecture, genome organization, and gene expression. Genes Dev 21: 3027–3043.
Shogren-Knaak M, Ishii H, Sun JM, Pazin MJ, Davie JR, Peterson CL . (2006). Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311: 844–847.
Singh NP, McCoy MT, Tice RR, Schneider EL . (1988). A simple technique for quantitation of low levels of DNA damage in individual cells. Exp Cell Res 175: 184–191.
Smith ER, Cayrou C, Huang R, Lane WS, Cote J, Lucchesi JC . (2005). A human protein complex homologous to the Drosophila MSL complex is responsible for the majority of histone H4 acetylation at lysine 16. Mol Cell Biol 25: 9175–9188.
Solomon JM, Pasupuleti R, Xu L, McDonagh T, Curtis R, DiStefano PS et al. (2006). Inhibition of SIRT1 catalytic activity increases p53 acetylation but does not alter cell survival following DNA damage. Mol Cell Biol 26: 28–38.
Stunkel W, Peh BK, Tan YC, Nayagam VM, Wang X, Salto-Tellez M et al. (2007). Function of the SIRT1 protein deacetylase in cancer. Biotechnol J 2: 1360–1368.
Taipale M, Rea S, Richter K, Vilar A, Lichter P, Imhof A et al. (2005). hMOF histone acetyltransferase is required for histone H4 lysine 16 acetylation in mammalian cells. Mol Cell Biol 25: 6798–6810.
Vakifahmetoglu H, Olsson M, Tamm C, Heidari N, Orrenius S, Zhivotovsky B . (2008). DNA damage induces two distinct modes of cell death in ovarian carcinomas. Cell Death Differ 15: 555–566.
Vaquero A, Scher M, Lee D, Erdjument-Bromage H, Tempst P, Reinberg D . (2004). Human SirT1 interacts with histone H1 and promotes formation of facultative heterochromatin. Mol Cell 16: 93–105.
Vaquero A, Scher MB, Lee DH, Sutton A, Cheng HL, Alt FW et al. (2006). SirT2 is a histone deacetylase with preference for histone H4 Lys 16 during mitosis. Genes Dev 20: 1256–1261.
Vaquero A, Sternglanz R, Reinberg D . (2007). NAD+-dependent deacetylation of H4 lysine 16 by class III HDACs. Oncogene 26: 5505–5520.
Vaziri H, Dessain SK, Ng Eaton E, Imai SI, Frye RA, Pandita TK et al. (2001). hSIR2(SIRT1) functions as an NAD-dependent p53 deacetylase. Cell 107: 149–159.
Villar-Garea A, Esteller M . (2004). Histone deacetylase inhibitors: understanding a new wave of anticancer agents. Int J Cancer 112: 171–178.
Yuan Z, Zhang X, Sengupta N, Lane WS, Seto E . (2007). SIRT1 regulates the function of the Nijmegen breakage syndrome protein. Mol Cell 27: 149–162.
Acknowledgements
We thank RG Roeder (The Rockefeller University), L Guarente (Massachusetts Institute of Technology) and M Henriksson (Karolinska Institutet) for providing us with hMOF DNA construct, SIRT1 DNA construct and SK-N-BE(2) cell line, respectively, and S Orrenius, T Perlmann and members of Hermanson and Joseph's labs for helpful comments. This work was supported by the Swedish Research Council, the Swedish Cancer Society, the Åke Wiberg Foundation, the Swedish Medical Society, the Karolinska Institutet Foundations (KI Cancer) (BJ and OH), SSF (CEDB & OBOE) and the Swedish Children's Cancer Foundation (OH).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Hajji, N., Wallenborg, K., Vlachos, P. et al. Opposing effects of hMOF and SIRT1 on H4K16 acetylation and the sensitivity to the topoisomerase II inhibitor etoposide. Oncogene 29, 2192–2204 (2010). https://doi.org/10.1038/onc.2009.505
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.505
Keywords
This article is cited by
-
The histone acetyltransferase Mof regulates Runx2 and Osterix for osteoblast differentiation
Cell and Tissue Research (2023)
-
Chidamide epigenetically represses autophagy and exerts cooperative antimyeloma activity with bortezomib
Cell Death & Disease (2020)
-
JMJD6 modulates DNA damage response through downregulating H4K16ac independently of its enzymatic activity
Cell Death & Differentiation (2020)
-
Tricho-rhino-phalangeal syndrome 1 protein functions as a scaffold required for ubiquitin-specific protease 4-directed histone deacetylase 2 de-ubiquitination and tumor growth
Breast Cancer Research (2018)
-
The return of the nucleus: transcriptional and epigenetic control of autophagy
Nature Reviews Molecular Cell Biology (2014)