Abstract
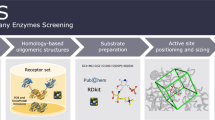
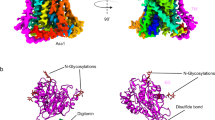
Cysteine is ligated to tRNACys by cysteinyl-tRNA synthetase in most organisms. However, in methanogenic archaea lacking cysteinyl-tRNA synthetase, O-phosphoserine is ligated to tRNACys by O-phosphoseryl–tRNA synthetase (SepRS), and the phosphoseryl-tRNACys is converted to cysteinyl-tRNACys. In this study, we determined the crystal structure of the SepRS tetramer in complex with tRNACys and O-phosphoserine at 2.6-Å resolution. The catalytic domain of SepRS recognizes the negatively charged side chain of O-phosphoserine at a noncanonical site, using the dipole moment of a conserved α-helix. The unique C-terminal domain specifically recognizes the anticodon GCA of tRNACys. On the basis of the structure, we engineered SepRS to recognize tRNACys mutants with the anticodons UCA and CUA and clarified the anticodon recognition mechanism by crystallography. The mutant SepRS-tRNA pairs may be useful for translational incorporation of O-phosphoserine into proteins in response to the stop codons UGA and UAG.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ibba, M. & Soll, D. Aminoacyl-tRNA synthesis. Annu. Rev. Biochem. 69, 617–650 (2000).
Bult, C.J. et al. Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 273, 1058–1073 (1996).
Sauerwald, A. et al. RNA-dependent cysteine biosynthesis in archaea. Science 307, 1969–1972 (2005).
O'Donoghue, P., Sethi, A., Woese, C.R. & Luthey-Schulten, Z.A. The evolutionary history of Cys-tRNACys formation. Proc. Natl. Acad. Sci. USA 102, 19003–19008 (2005).
Xie, J. & Schultz, P.G. A chemical toolkit for proteins–an expanded genetic code. Nat. Rev. Mol. Cell Biol. 7, 775–782 (2006).
Mosyak, L., Reshetnikova, L., Goldgur, Y., Delarue, M. & Safro, M.G. Structure of phenylalanyl-tRNA synthetase from Thermus thermophilus. Nat. Struct. Biol. 2, 537–547 (1995).
Sussman, J.L., Holbrook, S.R., Warrant, R.W., Church, G.M. & Kim, S.H. Crystal structure of yeast phenylalanine transfer RNA. I. Crystallographic refinement. J. Mol. Biol. 123, 607–630 (1978).
Hauenstein, S., Zhang, C.M., Hou, Y.M. & Perona, J.J. Shape-selective RNA recognition by cysteinyl-tRNA synthetase. Nat. Struct. Mol. Biol. 11, 1134–1141 (2004).
Goldgur, Y. et al. The crystal structure of phenylalanyl-tRNA synthetase from Thermus thermophilus complexed with cognate tRNAPhe. Structure 5, 59–68 (1997).
Hohn, M.J., Park, H.S., O'Donoghue, P., Schnitzbauer, M. & Soll, D. Emergence of the universal genetic code imprinted in an RNA record. Proc. Natl. Acad. Sci. USA 103, 18095–18100 (2006).
Nissen, P., Thirup, S., Kjeldgaard, M. & Nyborg, J. The crystal structure of Cys-tRNACys-EF-Tu-GDPNP reveals general and specific features in the ternary complex and in tRNA. Structure 7, 143–156 (1999).
Zhang, C.M., Perona, J.J., Ryu, K., Francklyn, C. & Hou, Y.M. Distinct kinetic mechanisms of the two classes of Aminoacyl-tRNA synthetases. J. Mol. Biol. 361, 300–311 (2006).
Ribas de Pouplana, L. & Schimmel, P. Aminoacyl-tRNA synthetases: potential markers of genetic code development. Trends Biochem. Sci. 26, 591–596 (2001).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Weeks, C.M. & Miller, R. Optimizing Shake-and-Bake for proteins. Acta Crystallogr. D Biol. Crystallogr. 55, 492–500 (1999).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Fukunaga, R. & Yokoyama, S. Crystal structure of leucyl-tRNA synthetase from the archaeon Pyrococcus horikoshii reveals a novel editing domain orientation. J. Mol. Biol. 346, 57–71 (2005).
Acknowledgements
We thank S. Sekine, T. Ito, T. Yanagisawa, A. Shimada, T. Sengoku, R. Ishii, M. Kawamoto, N. Shimizu, and H. Sakai for their help in data collection at SPring-8 and R. Akasaka for help in the analytical ultracentrifugation analysis. This work was supported by Grants-in-Aid for Scientific Research in Priority Areas from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan and by the RIKEN Structural Genomics/Proteomics Initiative of the National Project on Protein Structural and Functional Analyses, MEXT. R.F. was supported by Research Fellowships from the Japan Society for the Promotion of Science for Young Scientists.
Author information
Authors and Affiliations
Contributions
R.F. designed, performed and interpreted experiments and wrote the manuscript. S.Y. interpreted experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Structure of M. jannaschii SepRS. (PDF 530 kb)
Supplementary Fig. 2
Structure of A. fulgidus SepRS. (PDF 609 kb)
Supplementary Fig. 3
tRNA anticodon loop conformations. (PDF 382 kb)
Supplementary Fig. 4
Structure of the suppressor complexes of A. fulgidus SepRS. (PDF 1200 kb)
Supplementary Fig. 5
Engineering of A. fulgidus SepRS. (PDF 41 kb)
Rights and permissions
About this article
Cite this article
Fukunaga, R., Yokoyama, S. Structural insights into the first step of RNA-dependent cysteine biosynthesis in archaea. Nat Struct Mol Biol 14, 272–279 (2007). https://doi.org/10.1038/nsmb1219
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1219
This article is cited by
-
Structural basis for tRNA-dependent cysteine biosynthesis
Nature Communications (2017)
-
Biosynthesis and genetic encoding of phosphothreonine through parallel selection and deep sequencing
Nature Methods (2017)
-
Efficient genetic encoding of phosphoserine and its nonhydrolyzable analog
Nature Chemical Biology (2015)
-
The selective tRNA aminoacylation mechanism based on a single G•U pair
Nature (2014)
-
The mechanistic and evolutionary aspects of the 2′- and 3′-OH paradigm in biosynthetic machinery
Biology Direct (2013)