Abstract
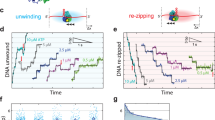
Helicases unwind dsDNA during replication, repair and recombination in an ATP-dependent reaction. The mechanism for helicase activity can be studied using oligonucleotide substrates to measure formation of single-stranded (ss) DNA from double-stranded (ds) DNA. This assay provides an 'all-or-nothing' readout because partially unwound intermediates are not detected. We have determined conditions under which an intermediate in the reaction cycle of Dda helicase can be detected by trapping a partially unwound substrate. The appearance of this intermediate supports a model in which each ssDNA product interacts with the helicase after unwinding has occurred. Kinetic analysis indicates that the intermediate appears during a slow step in the reaction cycle that is flanked by faster steps for unwinding. These observations demonstrate a complex mechanism containing nonuniform steps for a monomeric helicase. The potential biological significance of such a mechanism is discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Delagoutte, E. & von Hippel, P.H. Helicase mechanisms and the coupling of helicases within macromolecular machines. Part I: structures and properties of isolated helicases. Q. Rev. Biophys. 35, 431–478 (2002).
Delagoutte, E. & von Hippel, P.H. Helicase mechanisms and the coupling of helicases within macromolecular machines. Part II: integration of helicases into cellular processes. Q. Rev. Biophys. 36, 1–69 (2003).
von Hippel, P.H. Helicases become mechanistically simpler and functionally more complex. Nat. Struct. Mol. Biol. 11, 494–496 (2004).
Dillingham, M.S., Wigley, D.B. & Webb, M.R. Demonstration of unidirectional single-stranded DNA translocation by PcrA helicase: measurement of step size and translocation speed. Biochemistry 39, 205–212 (2000).
Velankar, S.S., Soultanas, P., Dillingham, M.S., Subramanya, H.S. & Wigley, D.B. Crystal structures of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism. Cell 97, 75–84 (1999).
Bianco, P.R. & Kowalczykowski, S.C. Translocation step size and mechanism of the RecBC DNA helicase. Nature 405, 368–372 (2000).
Hacker, K.J. & Alberts, B.M. Overexpression, purification, sequence analysis, and characterization of the T4 bacteriophage dda DNA helicase. J. Biol. Chem. 267, 20674–20681 (1992).
Jongeneel, C.V., Formosa, T. & Alberts, B.M. Purification and characterization of the bacteriophage T4 dda protein. A DNA helicase that associates with the viral helix-destabilizing protein. J. Biol. Chem. 259, 12925–12932 (1984).
Kadyrov, F.A. & Drake, J.W. UvsX recombinase and Dda helicase rescue stalled bacteriophage T4 DNA replication forks in vitro. J. Biol. Chem. 279, 35735–35740 (2004).
Ma, Y., Wang, T., Villemain, J.L., Giedroc, D.P. & Morrical, S.W. Dual functions of single-stranded DNA-binding protein in helicase loading at the bacteriophage T4 DNA replication fork. J. Biol. Chem. 279, 19035–19045 (2004).
Eoff, R.L., Spurling, T.L. & Raney, K.D. Chemically modified DNA substrates implicate the importance of electrostatic interactions for DNA unwinding by Dda helicase. Biochemistry 44, 666–674 (2005).
Raney, K.D. & Benkovic, S.J. Bacteriophage T4 Dda helicase translocates in a unidirectional fashion on single-stranded DNA. J. Biol. Chem. 270, 22236–22242 (1995).
Morris, P.D. et al. Evidence for a functional monomeric form of the bacteriophage T4 DdA helicase. Dda does not form stable oligomeric structures. J. Biol. Chem. 276, 19691–19698 (2001).
Nanduri, B., Byrd, A.K., Eoff, R.L., Tackett, A.J. & Raney, K.D. Pre-steady-state DNA unwinding by bacteriophage T4 Dda helicase reveals a monomeric molecular motor. Proc. Natl. Acad. Sci. USA 99, 14722–14727 (2002).
Byrd, A.K. & Raney, K.D. Increasing the length of the single-stranded overhang enhances unwinding of duplex DNA by bacteriophage T4 Dda helicase. Biochemistry 44, 12990–12997 (2005).
Levin, M.K., Wang, Y.H. & Patel, S.S. The functional interaction of the hepatitis C virus helicase molecules is responsible for unwinding processivity. J. Biol. Chem. 279, 26005–26012 (2004).
Tackett, A.J., Chen, Y., Cameron, C.E. & Raney, K.D. Multiple full-length NS3 molecules are required for optimal unwinding of oligonucleotide DNA in vitro. J. Biol. Chem. 280, 10797–10806 (2005).
Byrd, A.K. & Raney, K.D. Protein displacement by an assembly of helicase molecules aligned along single-stranded DNA. Nat. Struct. Mol. Biol. 11, 531–538 (2004).
Ali, J.A. & Lohman, T.M. Kinetic measurement of the step size of DNA unwinding by Escherichia coli UvrD helicase. Science 275, 377–380 (1997).
Lucius, A.L., Maluf, N.K., Fischer, C.J. & Lohman, T.M. General methods for analysis of sequential “n-step” kinetic mechanisms: application to single turnover kinetics of helicase-catalyzed DNA unwinding. Biophys. J. 85, 2224–2239 (2003).
Galletto, R., Jezewska, M.J. & Bujalowski, W. Unzipping mechanism of the double-stranded DNA unwinding by a hexameric helicase: quantitative analysis of the rate of the dsDNA unwinding, processivity and kinetic step-size of the Escherichia coli DnaB helicase using rapid quench-flow method. J. Mol. Biol. 343, 83–99 (2004).
Jeong, Y.J., Levin, M.K. & Patel, S.S. The DNA-unwinding mechanism of the ring helicase of bacteriophage T7. Proc. Natl. Acad. Sci. USA 101, 7264–7269 (2004).
Lucius, A.L. et al. DNA unwinding step-size of E. coli RecBCD helicase determined from single turnover chemical quenched-flow kinetic studies. J. Mol. Biol. 324, 409–428 (2002).
Okonogi, T.M., Alley, S.C., Harwood, E.A., Hopkins, P.B. & Robinson, B.H. Phosphate backbone neutralization increases duplex DNA flexibility: a model for protein binding. Proc. Natl. Acad. Sci. USA 99, 4156–4160 (2002).
Lucius, A.L. & Lohman, T.M. Effects of temperature and ATP on the kinetic mechanism and kinetic step-size for E. coli RecBCD helicase-catalyzed DNA unwinding. J. Mol. Biol. 339, 751–771 (2004).
Lucius, A.L., Jason, W.C. & Lohman, T.M. Fluorescence stopped-flow studies of single turnover kinetics of E. coli RecBCD helicase-catalyzed DNA unwinding. J. Mol. Biol. 339, 731–750 (2004).
Perkins, T.T., Li, H.W., Dalal, R.V., Gelles, J. & Block, S.M. Forward and reverse motion of single RecBCD molecules on DNA. Biophys. J. 86, 1640–1648 (2004).
Levin, M.K., Gurjar, M. & Patel, S.S. A Brownian motor mechanism of translocation and strand separation by hepatitis C virus helicase. Nat. Struct. Mol. Biol. 12, 429–435 (2005).
Tackett, A.J., Morris, P.D., Dennis, R., Goodwin, T.E. & Raney, K.D. Unwinding of unnatural substrates by a DNA helicase. Biochemistry 40, 543–548 (2001).
Galletto, R., Jezewska, M.J. & Bujalowski, W. Unzipping mechanism of the double-stranded DNA unwinding by a hexameric helicase: the effect of the 3′ arm and the stability of the dsDNA on the unwinding activity of the Escherichia coli DnaB helicase. J. Mol. Biol. 343, 101–114 (2004).
Arana, M.E., Haq, B., Tanguy Le, G.N. & Boehmer, P.E. Modulation of the herpes simplex virus type-1 UL9 DNA helicase by its cognate single-strand DNA-binding protein, ICP8. J. Biol. Chem. 276, 6840–6845 (2001).
He, X. & Lehman, I.R. Unwinding of a herpes simplex virus type 1 origin of replication (Ori(S)) by a complex of the viral origin binding protein and the single-stranded DNA binding protein. J. Virol. 74, 5726–5728 (2000).
Lee, S.S. & Lehman, I.R. Unwinding of the box I element of a herpes simplex virus type 1 origin by a complex of the viral origin binding protein, single-strand DNA binding protein, and single-stranded DNA. Proc. Natl. Acad. Sci. USA 94, 2838–2842 (1997).
Serebrov, V. & Pyle, A.M. Periodic cycles of RNA unwinding and pausing by hepatitis C virus NS3 helicase. Nature 430, 476–480 (2004).
Kawaoka, J., Jankowsky, E. & Pyle, A.M. Backbone tracking by the SF2 helicase NPH-II. Nat. Struct. Mol. Biol. 11, 526–530 (2004).
Acknowledgements
We would like to thank W. Bujalowski and A.L. Lucius for helpful discussions regarding general aspects of data analysis for DNA-unwinding reactions, with special thanks to A.L. Lucius for providing the initial Scientist script describing the inverse Laplace transform of Scheme 1. We would also like to thank M.K. Levin for helpful discussions. This work was supported by US National Institutes of Health grant R01 GM59400 (to K.D.R.) and the University of Arkansas for Medical Sciences Committee for Allocation of Graduate Student Research Funds (R.L.E.). Core facility support was provided by US National Institutes of Health grant P20 RR15569 (to F. Millet, P.I.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Position of the reannealing trap does not affect the appearance of the peak in ssDNA product. (PDF 72 kb)
Supplementary Fig. 2
Lowering the concentration of ATP slows the observed rate of helicase-catalyzed unwinding and delays the appearance of the intermediate. (PDF 54 kb)
Supplementary Fig. 3
Decreasing the concentration of ATP decreases the observed unwinding rate but does not affect the burst amplitude of ssDNA product. (PDF 53 kb)
Supplementary Fig. 4
Comparison of simulated eight-step time course with experimental unwinding data at low [ATP]. (PDF 60 kb)
Supplementary Fig. 5
Dda-catalyzed unwinding of a 120:60–mer results in the appearance of the trappable intermediate. (PDF 70 kb)
Rights and permissions
About this article
Cite this article
Eoff, R., Raney, K. Intermediates revealed in the kinetic mechanism for DNA unwinding by a monomeric helicase. Nat Struct Mol Biol 13, 242–249 (2006). https://doi.org/10.1038/nsmb1055
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1055
This article is cited by
-
Kinetic and structural mechanism for DNA unwinding by a non-hexameric helicase
Nature Communications (2021)
-
Coupling of DNA unwinding to nucleotide hydrolysis in a ring-shaped helicase
The EMBO Journal (2008)
-
Non-hexameric DNA helicases and translocases: mechanisms and regulation
Nature Reviews Molecular Cell Biology (2008)