Abstract
During meiotic prophase, the meiosis-specific telomere-binding protein TERB1 regulates chromosome movement required for homologous pairing and recombination by interacting with the telomeric shelterin subunit TRF1. Here, we report the crystal structure of the TRF1-binding motif of human TERB1 in complex with the TRFH domain of TRF1. Notably, specific disruption of the TERB1–TRF1 interaction by a point mutation in the mouse Terb1 gene results in infertility only in males. We find that this mutation causes an arrest in the zygotene–early pachytene stage and mild telomere abnormalities of autosomes but unpaired X and Y chromosomes in pachytene, leading to massive spermatocyte apoptosis. We propose that the loss of telomere structure mediated by the TERB1–TRF1 interaction significantly affects homologous pairing of the telomere-adjacent pseudoautosomal region (PAR) of the X and Y chromosomes in mouse spermatocytes. Our findings uncover a specific mechanism of telomeres that surmounts the unique challenges of mammalian X–Y pairing in meiosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Referenced accessions
NCBI Reference Sequence
Protein Data Bank
References
Handel, M.A. & Schimenti, J.C. Genetics of mammalian meiosis: regulation, dynamics and impact on fertility. Nat. Rev. Genet. 11, 124–136 (2010).
Petronczki, M., Siomos, M.F. & Nasmyth, K. Un ménage à quatre: the molecular biology of chromosome segregation in meiosis. Cell 112, 423–440 (2003).
Kleckner, N. Meiosis: how could it work? Proc. Natl. Acad. Sci. USA 93, 8167–8174 (1996).
Keeney, S., Giroux, C.N. & Kleckner, N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88, 375–384 (1997).
San Filippo, J., Sung, P. & Klein, H. Mechanism of eukaryotic homologous recombination. Annu. Rev. Biochem. 77, 229–257 (2008).
Neale, M.J. & Keeney, S. Clarifying the mechanics of DNA strand exchange in meiotic recombination. Nature 442, 153–158 (2006).
de Massy, B. Initiation of meiotic recombination: how and where? Conservation and specificities among eukaryotes. Annu. Rev. Genet. 47, 563–599 (2013).
Baudat, F., Imai, Y. & de Massy, B. Meiotic recombination in mammals: localization and regulation. Nat. Rev. Genet. 14, 794–806 (2013).
Syrjänen, J.L., Pellegrini, L. & Davies, O.R. A molecular model for the role of SYCP3 in meiotic chromosome organisation. eLife 3, e02963 (2014).
Bolcun-Filas, E. & Schimenti, J.C. Genetics of meiosis and recombination in mice. Int. Rev. Cell Mol. Biol. 298, 179–227 (2012).
Hirose, Y. et al. Chiasmata promote monopolar attachment of sister chromatids and their co-segregation toward the proper pole during meiosis I. PLoS Genet. 7, e1001329 (2011).
Bascom-Slack, C.A., Ross, L.O. & Dawson, D.S. Chiasmata, crossovers, and meiotic chromosome segregation. Adv. Genet. 35, 253–284 (1997).
Boateng, K.A., Bellani, M.A., Gregoretti, I.V., Pratto, F. & Camerini-Otero, R.D. Homologous pairing preceding SPO11-mediated double-strand breaks in mice. Dev. Cell 24, 196–205 (2013).
Brown, P.W. et al. Meiotic synapsis proceeds from a limited number of subtelomeric sites in the human male. Am. J. Hum. Genet. 77, 556–566 (2005).
Koszul, R., Kim, K.P., Prentiss, M., Kleckner, N. & Kameoka, S. Meiotic chromosomes move by linkage to dynamic actin cables with transduction of force through the nuclear envelope. Cell 133, 1188–1201 (2008).
Koszul, R. & Kleckner, N. Dynamic chromosome movements during meiosis: a way to eliminate unwanted connections? Trends Cell Biol. 19, 716–724 (2009).
Lee, C.Y. et al. Mechanism and regulation of rapid telomere prophase movements in mouse meiotic chromosomes. Cell Rep. 11, 551–563 (2015).
Hiraoka, Y. & Dernburg, A.F. The SUN rises on meiotic chromosome dynamics. Dev. Cell 17, 598–605 (2009).
Ding, X. et al. SUN1 is required for telomere attachment to nuclear envelope and gametogenesis in mice. Dev. Cell 12, 863–872 (2007).
Daniel, K. et al. Mouse CCDC79 (TERB1) is a meiosis-specific telomere associated protein. BMC Cell Biol. 15, 17 (2014).
Shibuya, H., Ishiguro, K. & Watanabe, Y. The TRF1-binding protein TERB1 promotes chromosome movement and telomere rigidity in meiosis. Nat. Cell Biol. 16, 145–156 (2014).
Shibuya, H. et al. MAJIN links telomeric DNA to the nuclear membrane by exchanging telomere cap. Cell 163, 1252–1266 (2015).
Viera, A. et al. CDK2 regulates nuclear envelope protein dynamics and telomere attachment in mouse meiotic prophase. J. Cell Sci. 128, 88–99 (2015).
Tu, Z. et al. Speedy A-Cdk2 binding mediates initial telomere-nuclear envelope attachment during meiotic prophase I independent of Cdk2 activation. Proc. Natl. Acad. Sci. USA 114, 592–597 (2017).
Perry, J., Palmer, S., Gabriel, A. & Ashworth, A. A short pseudoautosomal region in laboratory mice. Genome Res. 11, 1826–1832 (2001).
Kauppi, L., Jasin, M. & Keeney, S. The tricky path to recombining X and Y chromosomes in meiosis. Ann. NY Acad. Sci. 1267, 18–23 (2012).
Kauppi, L. et al. Distinct properties of the XY pseudoautosomal region crucial for male meiosis. Science 331, 916–920 (2011).
Chen, Y. et al. A shared docking motif in TRF1 and TRF2 used for differential recruitment of telomeric proteins. Science 319, 1092–1096 (2008).
Wan, B. et al. SLX4 assembles a telomere maintenance toolkit by bridging multiple endonucleases with telomeres. Cell Rep. 4, 861–869 (2013).
Rai, R. et al. NBS1 phosphorylation status dictates repair choice of dysfunctional telomeres. Mol. Cell 65, 801–817 e4 (2017).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
de la Fuente, R. et al. Meiotic pairing and segregation of achiasmate sex chromosomes in eutherian mammals: the role of SYCP3 protein. PLoS Genet. 3, e198 (2007).
de Vries, F.A.T. et al. Mouse Sycp1 functions in synaptonemal complex assembly, meiotic recombination, and XY body formation. Genes Dev. 19, 1376–1389 (2005).
Klutstein, M. & Cooper, J.P. The chromosomal courtship dance-homolog pairing in early meiosis. Curr. Opin. Cell Biol. 26, 123–131 (2014).
Scherthan, H. Analysis of telomere dynamics in mouse spermatogenesis. Methods Mol. Biol. 558, 383–399 (2009).
Scherthan, H. et al. Centromere and telomere movements during early meiotic prophase of mouse and man are associated with the onset of chromosome pairing. J. Cell Biol. 134, 1109–1125 (1996).
Hunter, N., Börner, G.V., Lichten, M. & Kleckner, N. Gamma-H2AX illuminates meiosis. Nat. Genet. 27, 236–238 (2001).
Celli, G.B. & de Lange, T. DNA processing is not required for ATM-mediated telomere damage response after TRF2 deletion. Nat. Cell Biol. 7, 712–718 (2005).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Kim, S.H., Kaminker, P. & Campisi, J. TIN2, a new regulator of telomere length in human cells. Nat. Genet. 23, 405–412 (1999).
Ye, J.Z. et al. POT1-interacting protein PIP1: a telomere length regulator that recruits POT1 to the TIN2/TRF1 complex. Genes Dev. 18, 1649–1654 (2004).
Frescas, D. & de Lange, T. TRF2-tethered TIN2 can mediate telomere protection by TPP1/POT1. Mol. Cell. Biol. 34, 1349–1362 (2014).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr. D Biol. Crystallogr. 62, 859–866 (2006).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Schrodinger, L. The PyMOL Molecular Graphics System, Version 1.8. (2015).
Moretti, P., Freeman, K., Coodly, L. & Shore, D. Evidence that a complex of SIR proteins interacts with the silencer and telomere-binding protein RAP1. Genes Dev. 8, 2257–2269 (1994).
Bastos, H. et al. Flow cytometric characterization of viable meiotic and postmeiotic cells by Hoechst 33342 in mouse spermatogenesis. Cytometry A 65, 40–49 (2005).
Getun, I.V., Torres, B. & Bois, P.R. Flow cytometry purification of mouse meiotic cells. J. Vis. Exp. 50, 2602 (2011).
Peters, A.H., Plug, A.W., van Vugt, M.J. & de Boer, P. A drying-down technique for the spreading of mammalian meiocytes from the male and female germline. Chromosome Res. 5, 66–68 (1997).
Scherthan, H. et al. Mammalian meiotic telomeres: protein composition and redistribution in relation to nuclear pores. Mol. Biol. Cell 11, 4189–4203 (2000).
Acknowledgements
We thank H. Shi, X. Huang, Q. Shi, M. Tong, and J. Li for discussion. We thank M. Han (Fudan University) for the SUN1 antibody. We thank L. Wu, D. Yao, and R. Zhang from BL18U1 and BL19U1 beamlines at NCPSS and Shanghai Synchrotron Radiation Facility (SSRF) for help with crystal data collection and processing. We thank Y. Yu, S. He, Y. Wang and S. Li from NCPSS for help with confocal microscopy and FACS. This work was supported by grants from the Ministry of Science and Technology of China (2013CB910402 to M.L.), the National Natural Science Foundation of China (31330040 and 31525007 to M.L., 31500625 to J.W.), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB08010201 to M.L.), the Outstanding Academic Leader Program of Science and Technology Commission of Shanghai Municipality (16XD1405000 to M.L. and C.H.), and the Youth Innovation Promotion Association of the Chinese Academy of Sciences (to J.W.). Y.L. acknowledges support from the Intramural Research Program of the National Institutes of Health, National Institute on Aging.
Author information
Authors and Affiliations
Contributions
J.L., C.H., J.W., and M.L. designed the study. J.L. crystallized the TERB1TBM–TRF1TRFH complex. J.W. collected X-ray data and determined the crystal structure. J.L., C.H., Y.C., and S.S. carried out biochemical assays. J.L., Y.L., C.L., Y.Z., and L.W. participated in the generation of the knock-in mice. J.L., C.H., and Y.C. performed the experiments to characterize the meiotic phenotype. C.H., J.W., and L.M. drafted the manuscript. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of the interaction between TRF1TRFH and TERB1TBM.
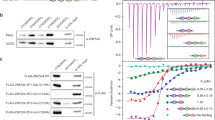
Yeast two-hybrid analysis of the interaction between TRF1TRFH with different fragments of TERB1. Interaction between LexA-TRF1TRFH and different GAD-TERB1 fragments was analyzed by measuring the β-galactosidase activity produced by the reporter gene. Data are averages of three independent β-galactosidase measurements normalized to the interaction of TRF1TRFH with TERB1522-622, arbitrarily set to 100. Error bars: standard deviation (s.d.); n=3.
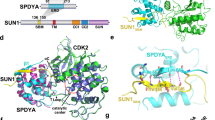
Supplementary Figure 2 Crystal structure of the TRF1TRFH and TERB1TBM complex.
(a) Stereo view of the TRF1TRFH-TERB1TBM complex. The electron density (2Fo-Fc) map of TERB1TBM, contoured at 1.0 σ, is shown in the complex. (b) The electrostatic surface potential of TRF1TRFH responsible for TERB1TBM binding (positive potential, blue; negative potential, red). TERB1TBM is in stick representation and colored in yellow.
Supplementary Figure 3 TERB1TBM and TIN2TBM occupy the same binding pocket on TRF1TRFH.
Electrostatic surface potential of superimposed complex structures, TRF1TRFH-TERB1TBM and TRF1TRFH-TIN2TBM are shown in (a), (b), and (c), respectively. TERB1TBM and TIN2TBM are in stick representation and colored in yellow and cyan.
Supplementary Figure 4 In vitro ITC measurement between TRF1TRFH and mutant TERB1TBM.
All the mutant TERB1TBM disrupt the interaction with TRF1TRFH, consistent with the Co-IP and co-localization data shown in Figure 2.
Supplementary Figure 5 The TERB1 647AEA649 mutant mice were generated using the CRISPR/Cas9 method.
(a) The Cas9/sgRNA-targeting sites in mouse Terb1 gene. The sgRNA-targeting sequence and the protospacer-adjacent motif (PAM) are underlined. The desired mutations on genomic DNA are indicated by lower-case letters. (b) Transgenic mouse offspring were genotyped by tail biopsy with PCR and DNA sequencing of the mutation loci. (c) Relative mRNA expression was determined in testes of 3-week old littermates by RT-qPCR using GAPDH as the loading control. The averaged value for WT was normalized to 1. Results were from three independent experiments. Error bars: standard deviation; n=3. (d) Western blot of TERB1 in testes of two-week-old littermates. β-actin was used as the loading control. The indicated numbers below the blot images represent the relative signals of anti-TERB1 normalized by the signals of anti-Actin. Band intensity of blots was quantified by ImageJ. The number for WT sample was set as 1. Uncropped blot images are shown in Supplementary Data Set 1.
Supplementary Figure 6 Hoechst 33342 fluorescence flow analysis of testicular cells from 6-week-old WT and Terb1AEA/AEA littermates.
(a) Left panel: Schematic diagram summarizes the Hoechst 33342 fluorescent profile of male germinal cells in adult spermatogenesis. Right panel: Hoechst 33342 fluorescence flow analysis of testicular cells from 6-week-old WT and Terb1AEA/AEA littermates. L-Z: subpopulation that mainly comprises leptotene and zygotene spermatocytes. P-D: subpopulation that mainly comprises pachytene and diplotene spermatocytes. Subpopulation was sorted and analyzed by immunofluorescence staining of SYCP3 and γ-H2AX or propidium iodide staining to determine the cell types (data not shown). (b) Quantification of spermatid population in all testicular cells. Statistical significances (*** p<0.01) were assessed by two-tailed t-tests. Error bars: standard deviation; n=4. (c) Quantification of L-Z and P-D subpopulation in all spermatocyte I (4N DNA content). Statistical significances (** p<0.05) were assessed by two-tailed t-tests. Error bars: standard deviation; n=4.
Supplementary Figure 7 Programmed DSB repair and chiasma formation autosomes are largely unaffected in Terb1AEA/AEA spermatocytes.
(a) WT and Terb1AEA/AEA pachytene spermatocytes stained for MLH1 (red), SYCP3 (green) and DAPI (blue). Scale bar: 10 μm. (b) Quantification of the number of MLH1 foci per cell from (a). The mean values were indicated. (c) WT and Terb1AEA/AEA spermatocytes stained for γ-H2AX (red), SYCP3 (green) and DAPI (blue). Scale bar: 10 μm.
Supplementary Figure 8 The 647AEA649 mutation of TERB1 results in telomere localization defect of SUN1.
(a) Cell lysates were prepared with testes from 3-week-old WT, heterozygous and Terb1AEA/AEA mice and immunoprecipitated with TERB1 antibody. Lysate samples and immunoprecipitates were blotted using antibodies as indicated. The indicated numbers below the blot images represent the relative signals of Co-IPed TRF1 normalized by the signals of IPed TERB1. Band intensity of blots was quantified by ImageJ. The number for WT sample was set as 1. Uncropped blot images are shown in Supplementary Data Set 1. (b) Localization of SUN1 (red) and SYCP3 (green) on pachytene spermatocyte spreads from WT and Terb1AEA/AEA mice. Scale bar: 10 μm. (c) Quantification of the relative intensity of SUN1 foci at the termini of synaptic chromosomes in WT and Terb1AEA/AEA pachytene spermatocytes. Three independent experiments with 6-week-old littermates showed similar results. More than 10 spermatocyte spreads were used for quantification for each group. (a.u., arbitrary units)
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1810 kb)
Life Sciences Reporting Summary
Life Sciences Reporting Summary (PDF 176 kb)
Supplementary Data Set 1
Uncropped gels of supplementary figures. (PDF 1457 kb)
Rights and permissions
About this article
Cite this article
Long, J., Huang, C., Chen, Y. et al. Telomeric TERB1–TRF1 interaction is crucial for male meiosis. Nat Struct Mol Biol 24, 1073–1080 (2017). https://doi.org/10.1038/nsmb.3496
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3496
This article is cited by
-
Loss of SUN1 function in spermatocytes disrupts the attachment of telomeres to the nuclear envelope and contributes to non-obstructive azoospermia in humans
Human Genetics (2023)
-
The double-stranded DNA-binding proteins TEBP-1 and TEBP-2 form a telomeric complex with POT-1
Nature Communications (2021)
-
Stage-resolved Hi-C analyses reveal meiotic chromosome organizational features influencing homolog alignment
Nature Communications (2021)
-
The SUN1-SPDYA interaction plays an essential role in meiosis prophase I
Nature Communications (2021)
-
The TERB1-TERB2-MAJIN complex of mouse meiotic telomeres dates back to the common ancestor of metazoans
BMC Evolutionary Biology (2020)