Abstract
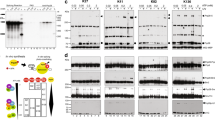
SRp38 is an atypical SR protein that functions as a general splicing repressor when dephosphorylated. We now show that phosphorylated SRp38 functions as a sequence-specific splicing activator. Unlike characterized splicing activators, SRp38 functions in the absence of other SR proteins but requires a cofactor for activity. SRp38 was able to induce formation of splicing complex A in the absence of the cofactor, but this factor was necessary for progression to complexes B and C. Mechanistically, SRp38 strengthens the ability of the U1 and U2 small nuclear ribonucleoproteins to stably recognize the pre-mRNA. Extending these findings, analysis of alternative splicing of pre-mRNA encoding the glutamate receptor B revealed that SRp38 alters its splicing pattern in a sequence-specific manner. Together, our data demonstrate that SRp38, in addition to its role as a splicing repressor, can function as an unusual sequence-specific splicing activator.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Black, D.L. Protein diversity from alternative splicing: a challenge for bioinformatics and post-genome biology. Cell 103, 367–370 (2000).
Johnson, J.M. et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science 302, 2141–2144 (2003).
Jurica, M.S. & Moore, M.J. Pre-mRNA splicing: awash in a sea of proteins. Mol. Cell 12, 5–14 (2003).
Smith, C.W. & Valcarcel, J. Alternative pre-mRNA splicing: the logic of combinatorial control. Trends Biochem. Sci. 25, 381–388 (2000).
Stamm, S. Regulation of alternative splicing by reversible protein phosphorylation. J. Biol. Chem. 283, 1223–1227 (2008).
David, C.J. & Manley, J.L. The search for alternative splicing regulators: new approaches offer a path to a splicing code. Genes Dev. 22, 279–285 (2008).
Wang, J., Smith, P.J., Krainer, A.R. & Zhang, M.Q. Distribution of SR protein exonic splicing enhancer motifs in human protein-coding genes. Nucleic Acids Res. 33, 5053–5062 (2005).
Tanaka, K., Watakabe, A. & Shimura, Y. Polypurine sequences within a downstream exon function as a splicing enhancer. Mol. Cell. Biol. 14, 1347–1354 (1994).
Tacke, R., Tohyama, M., Ogawa, S. & Manley, J.L. Human TRA2 proteins are sequence-specific activators of pre-mRNA splicing. Cell 93, 139–148 (1998).
Goren, A. et al. Comparative analysis identifies exonic splicing regulatory sequences—the complex definition of enhancers and silencers. Mol. Cell 22, 769–781 (2006).
Zheng, Z.M. Regulation of alternative RNA splicing by exon definition and exon sequences in viral and mammalian gene expression. J. Biomed. Sci. 11, 278–294 (2004).
Manley, J.L. & Tacke, R. SR proteins and splicing control. Genes Dev. 10, 1569–1579 (1996).
Graveley, B.R. Sorting out the complexity of SR protein functions. RNA 6, 1197–1211 (2000).
Hastings, M.L. & Krainer, A.R. Pre-mRNA splicing in the new millennium. Curr. Opin. Cell Biol. 13, 302–309 (2001).
Wu, J.Y. & Maniatis, T. Specific interactions between proteins implicated in splice site selection and regulated alternative splicing. Cell 75, 1061–1070 (1993).
Kohtz, J.D. et al. Protein-protein interactions and 5′-splice-site recognition in mammalian mRNA precursors. Nature 368, 119–124 (1994).
Shen, H. & Green, M.R. RS domains contact splicing signals and promote splicing by a common mechanism in yeast through humans. Genes Dev. 20, 1755–1765 (2006).
Hertel, K.J. & Graveley, B.R. RS domains contact the pre-mRNA throughout spliceosome assembly. Trends Biochem. Sci. 30, 115–118 (2005).
Matlin, A.J., Clark, F. & Smith, C.W. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol. 6, 386–398 (2005).
Wang, Z., Hoffmann, H.M. & Grabowski, P.J. Intrinsic U2AF binding is modulated by exon enhancer signals in parallel with changes in splicing activity. RNA 1, 21–35 (1995).
Zuo, P. & Maniatis, T. The splicing factor U2AF35 mediates critical protein-protein interactions in constitutive and enhancer-dependent splicing. Genes Dev. 10, 1356–1368 (1996).
Eldridge, A.G., Li, Y., Sharp, P.A. & Blencowe, B.J. The SRm160/300 splicing coactivator is required for exon-enhancer function. Proc. Natl. Acad. Sci. USA 96, 6125–6130 (1999).
Li, Y. & Blencowe, B.J. Distinct factor requirements for exonic splicing enhancer function and binding of U2AF to the polypyrimidine tract. J. Biol. Chem. 274, 35074–35079 (1999).
Ryner, L.C. et al. Control of male sexual behavior and sexual orientation in Drosophila by the fruitless gene. Cell 87, 1079–1089 (1996).
Shin, C., Feng, Y. & Manley, J.L. Dephosphorylated SRp38 acts as a splicing repressor in response to heat shock. Nature 427, 553–558 (2004).
Shin, C. & Manley, J.L. The SR protein SRp38 represses splicing in M phase cells. Cell 111, 407–417 (2002).
Shi, Y. & Manley, J.L. A complex signaling pathway regulates SRp38 phosphorylation and pre-mRNA splicing in response to heat shock. Mol. Cell 28, 79–90 (2007).
Shin, C., Kleiman, F.E. & Manley, J.L. Multiple properties of the splicing repressor SRp38 distinguish it from typical SR proteins. Mol. Cell. Biol. 25, 8334–8343 (2005).
Prasad, J., Colwill, K., Pawson, T. & Manley, J.L. The protein kinase Clk/Sty directly modulates SR protein activity: both hyper- and hypophosphorylation inhibit splicing. Mol. Cell. Biol. 19, 6991–7000 (1999).
Ruskin, B., Zamore, P.D. & Green, M.R. A factor, U2AF, is required for U2 snRNP binding and splicing complex assembly. Cell 52, 207–219 (1988).
Komatsu, M., Kominami, E., Arahata, K. & Tsukahara, T. Cloning and characterization of two neural-salient serine/arginine-rich (NSSR) proteins involved in the regulation of alternative splicing in neurones. Genes Cells 4, 593–606 (1999).
Sommer, B. et al. Flip and flop: a cell-specific functional switch in glutamate-operated channels of the CNS. Science 249, 1580–1585 (1990).
Chen, X. et al. Tra2βl regulates P19 neuronal differentiation and the splicing of FGF-2R and GluR-B minigenes. Cell Biol. Int. 28, 791–799 (2004).
Tacke, R. & Manley, J.L. The human splicing factors ASF/SF2 and SC35 possess distinct, functionally significant RNA binding specificities. EMBO J. 14, 3540–3551 (1995).
Tacke, R., Chen, Y. & Manley, J.L. Sequence-specific RNA binding by an SR protein requires RS domain phosphorylation: creation of an SRp40-specific splicing enhancer. Proc. Natl. Acad. Sci. USA 94, 1148–1153 (1997).
Graveley, B.R. & Maniatis, T. Arginine/serine-rich domains of SR proteins can function as activators of pre-mRNA splicing. Mol. Cell 1, 765–771 (1998).
Shen, H., Kan, J.L. & Green, M.R. Arginine-serine-rich domains bound at splicing enhancers contact the branchpoint to promote prespliceosome assembly. Mol. Cell 13, 367–376 (2004).
Xiao, S.H. & Manley, J.L. Phosphorylation of the ASF/SF2 RS domain affects both protein-protein and protein-RNA interactions and is necessary for splicing. Genes Dev. 11, 334–344 (1997).
Xiao, S.H. & Manley, J.L. Phosphorylation-dephosphorylation differentially affects activities of splicing factor ASF/SF2. EMBO J. 17, 6359–6367 (1998).
Dye, B.T., Buvoli, M., Mayer, S.A., Lin, C.H. & Patton, J.G. Enhancer elements activate the weak 3′ splice site of α-tropomyosin exon 2. RNA 4, 1523–1536 (1998).
Wang, J. & Bell, L.R. The Sex-lethal amino terminus mediates cooperative interactions in RNA binding and is essential for splicing regulation. Genes Dev. 8, 2072–2085 (1994).
Tacke, R. & Manley, J.L. Functions of SR and Tra2 proteins in pre-mRNA splicing regulation. Proc. Soc. Exp. Biol. Med. 220, 59–63 (1999).
Li, X., Shambaugh, M.E., Rottman, F.M. & Bokar, J.A. SR proteins Asf/SF2 and 9G8 interact to activate enhancer-dependent intron D splicing of bovine growth hormone pre-mRNA in vitro. RNA 6, 1847–1858 (2000).
Roscigno, R.F. & Garcia-Blanco, M.A. SR proteins escort the U4/U6.U5 tri-snRNP to the spliceosome. RNA 1, 692–706 (1995).
Tarn, W.Y. & Steitz, J.A. Modulation of 5′ splice site choice in pre-messenger RNA by two distinct steps. Proc. Natl. Acad. Sci. USA 92, 2504–2508 (1995).
Crovato, T.E. & Egebjerg, J. ASF/SF2 and SC35 regulate the glutamate receptor subunit 2 alternative flip/flop splicing. FEBS Lett. 579, 4138–4144 (2005).
Fisher, C.L. & Pei, G.K. Modification of a PCR-based site-directed mutagenesis method. Biotechniques 23, 570–571 574 (1997).
Arakawa, H., Lodygin, D. & Buerstedde, J.M. Mutant loxP vectors for selectable marker recycle and conditional knock-outs. BMC Biotechnol. 1, 7 (2001).
Das, R. & Reed, R. Resolution of the mammalian E complex and the ATP-dependent spliceosomal complexes on native agarose mini-gels. RNA 5, 1504–1508 (1999).
Wang, J. & Manley, J.L. Overexpression of the SR proteins ASF/SF2 and SC35 influences alternative splicing in vivo in diverse ways. RNA 1, 335–346 (1995).
Acknowledgements
We thank R. Lührmann (Max Planck Institute for Biophysical Chemistry, Germany) for providing purified U1 and U2 snRNP, T. Kashima (Columbia University, New York) for SR proteins and C. Shin (Columbia University, New York) for His-dSRp38. We thank A. Yang for help with the manuscript. We also thank members of the Manley laboratory for helpful discussions and comments. This work was supported by the US National Institutes of Health grant NIH GM48259.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 (PDF 186 kb)
Rights and permissions
About this article
Cite this article
Feng, Y., Chen, M. & Manley, J. Phosphorylation switches the general splicing repressor SRp38 to a sequence-specific activator. Nat Struct Mol Biol 15, 1040–1048 (2008). https://doi.org/10.1038/nsmb.1485
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1485
This article is cited by
-
Improved modeling of RNA-binding protein motifs in an interpretable neural model of RNA splicing
Genome Biology (2024)
-
lncR-GAS5 upregulates the splicing factor SRSF10 to impair endothelial autophagy, leading to atherogenesis
Frontiers of Medicine (2023)
-
Regulatory roles and mechanisms of alternative RNA splicing in adipogenesis and human metabolic health
Cell & Bioscience (2021)
-
Diversification of the muscle proteome through alternative splicing
Skeletal Muscle (2018)
-
SRSF10-mediated IL1RAP alternative splicing regulates cervical cancer oncogenesis via mIL1RAP-NF-κB-CD47 axis
Oncogene (2018)