Abstract
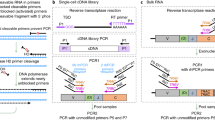
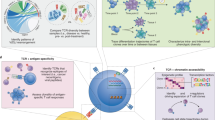
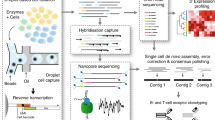
High-throughput sequencing of the variable domains of immune receptors (antibodies and T cell receptors (TCRs)) is of key importance in the understanding of adaptive immune responses in health and disease. However, the sequencing of both immune receptor chains (VH+VL or TCRβ/δ+TCRα/γ) at the single-cell level for typical samples containing >104 lymphocytes is problematic, because immune receptors comprise two polypeptide chains that are encoded by separate mRNAs. Here we present a technology that allows rapid and low-cost determination of a paired immune receptor repertoire from millions of cells with high precision (>97%). Flow focusing is used to encapsulate single cells in emulsions containing magnetic beads for mRNA capture. The mRNA transcripts are then reverse-transcribed, physically linked to their partners by overlap extension PCR, and interrogated by high-throughput paired-end Illumina sequencing. This protocol describes the construction and operation of the flow-focusing device in detail, as well as the bioinformatics pipeline used to interpret the data. The entire procedure can be performed by a single researcher in under 12 h of effort per sample.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Finco, O. & Rappuoli, R. Designing vaccines for the twenty-first century society. Front. Immunol. 5, 12 (2014).
Georgiou, G. et al. The promise and challenge of high-throughput sequencing of the antibody repertoire. Nat. Biotechnol. 32, 158–168 (2014).
Logan, A.C. et al. High-throughput VDJ sequencing for quantification of minimal residual disease in chronic lymphocytic leukemia and immune reconstitution assessment. Proc. Natl. Acad. Sci. USA 108, 21194–21199 (2011).
Mascola, J.R. & Haynes, B.F. Hiv-1 neutralizing antibodies: understanding nature′s pathways. Immunol. Rev. 254, 225–244 (2013).
Warren, E.H., Matsen, F.A.t. & Chou, J. High-throughput sequencing of B- and T-lymphocyte antigen receptors in hematology. Blood 122, 19–22 (2013).
Brezinschek, H.P., Foster, S.J., Dorner, T., Brezinschek, R.I. & Lipsky, P.E. Pairing of variable heavy and variable kappa chains in individual naive and memory B cells. J. Immunol. 160, 4762–4767 (1998).
Meijer, P.J . et al. Isolation of human antibody repertoires with preservation of the natural heavy and light chain pairing. J. Mol. Biol. 358, 764–772 (2006).
Poulsen, T.R., Meijer, P.J., Jensen, A., Nielsen, L.S. & Andersen, P.S. Kinetic, affinity, and diversity limits of human polyclonal antibody responses against tetanus toxoid. J. Immunol. 179, 3841–3850 (2007).
Smith, K. et al. Rapid generation of fully human monoclonal antibodies specific to a vaccinating antigen. Nat. Protoc. 4, 372–384 (2009).
Howie, B. et al. High-throughput pairing of T cell receptor alpha and beta sequences. Sci. Transl. Med. 7, 301ra131 (2015).
DeKosky, B.J. et al. High-throughput sequencing of the paired human immunoglobulin heavy and light chain repertoire. Nat. Biotechnol. 31, 166–169 (2013).
Turchaninova, M.A. et al. Pairing of T-cell receptor chains via emulsion PCR. Eur. J. Immunol. 43, 2507–2515 (2013).
Busse, C.E., Czogiel, I., Braun, P., Arndt, P.F. & Wardemann, H. Single-cell based high-throughput sequencing of full-length immunoglobulin heavy and light chain genes. Eur. J. Immunol. 44, 597–603 (2014).
DeKosky, B.J. et al. In-depth determination and analysis of the human paired heavy- and light-chain antibody repertoire. Nat. Med. 21, 86–91 (2015).
Doria-Rose, N.A. et al. Developmental pathway for potent V1V2-directed HIV-neutralizing antibodies. Nature 509, 55–62 (2014).
Berkland, C., Kim, K. & Pack, D.W. Fabrication of PLG microspheres with precisely controlled and monodisperse size distributions. J. Control. Release 73, 59–74 (2001).
Berkland, C., Pollauf, E., Pack, D.W. & Kim, K. Uniform double-walled polymer microspheres of controllable shell thickness. J. Control. Release 96, 101–111 (2004).
Lavinder, J.J. et al. Identification and characterization of the constituent human serum antibodies elicited by vaccination. Proc. Natl. Acad. Sci. USA 111, 2259–2264 (2014).
Beltramello, M. et al. The human immune response to dengue virus is dominated by highly cross-reactive antibodies endowed with neutralizing and enhancing activity. Cell Host Microbe 8, 271–283 (2010).
Walker, L.M. et al. Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 326, 285–289 (2009).
Liao, H.X. et al. High-throughput isolation of immunoglobulin genes from single human B cells and expression as monoclonal antibodies. J. Virol. Methods 158, 171–179 (2009).
Wrammert, J. et al. Rapid cloning of high-affinity human monoclonal antibodies against influenza virus. Nature 453, 667–671 (2008).
Wardemann, H. et al. Predominant autoantibody production by early human B cell precursors. Science 301, 1374–1377 (2003).
Carlson, C.S. et al. Using synthetic templates to design an unbiased multiplex PCR assay. Nat. Commun. 4, 2680 (2013).
Ippolito, G.C. et al. Antibody repertoires in humanized NOD-scid-il2rγnull mice and human B cells reveals human-like diversification and tolerance checkpoints in the mouse. PLoS One 7, e35497 (2012).
Lim, T.S. et al. V-gene amplification revisited: an optimised procedure for amplification of rearranged human antibody genes of different isotypes. N. Biotechnol. 27, 108–117 (2010).
Brochet, X., Lefranc, M.P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–W508 (2008).
Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Greiff, V., Miho, E., Menzel, U. & Reddy, S.T. Bioinformatic and statistical analysis of adaptive immune repertoires. Trends Immunol 36, 738–749 (2014).
Saha, S. & Raghava, G.P. Algpred: prediction of allergenic proteins and mapping of IGE epitopes. Nucleic Acids Res. 34, W202–W209 (2006).
Magoc, T. & Salzberg, S.L. Flash: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. Pear: a fast and accurate Illumina paired-end read merger. Bioinformatics 30, 614–620 (2014).
Acknowledgements
We thank A. Kennedy for his assistance in making the borosilicate glass nozzles. Funding for this work was provided by Defense Threat Reduction Agency HDTRA1-12-C-0105 (G.G.), National Institutes of Health (NIH) Grant R01AI096228 (G.G.), and fellowships to B.J.D. from the Hertz Foundation, the University of Texas Donald D. Harrington Foundation and the National Science Foundation. H.T. was supported by Japan Society for the Promotion of Science (JSPS) Postdoctoral Fellowships for Research Abroad.
Author information
Authors and Affiliations
Contributions
J.R.M., B.J.D. and G.G. wrote the manuscript. J.R.M., B.J.D., H.T., A.D.E and G.G. designed the experiments. J.R.M., B.J.D. and H.T. performed the experiments and conducted bioinformatic analyses. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
G.G., B.J.D. and A.D.E. declare competing financial interests in the form of a patent filed by the University of Texas at Austin.
Integrated supplementary information
Supplementary Figure 1 Alignment of custom-designed glass nozzle.
A) PBS/PBS aqueous phases in oil; nozzle aperture is properly aligned. B) PBS/Lysis Buffer aqueous phases in oil; nozzle aperture is properly aligned. The presence of surfactants greatly reduces the width of the aqueous stream. C) PBS/Lysis Buffer aqueous phases in oil; aperture is off-center. D) PBS/Lysis Buffer aqueous phases in oil; aperture is so far off-center that the aqueous phase is coming into contact with the curvature of the glass nozzle. In all pictures, the stream is flowing right to left.
Supplementary Figure 2 Emulsions collected from flow-focusing device before and after centrifugation.
The top layer of mineral oil containing microfines (small emulsions) should be removed and discarded. The bottom layer of larger emulsions contains cells and beads, which should be retained. The color will change depending on the concentration of the poly(dT) beads.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Table 1 (PDF 476 kb)
Aqueous/oil interface for misaligned glass nozzle.
If the concentric metal tubes are not centered on the aperture of the nozzle, the aqueous stream will flicker or appear asymmetric. If the needle-to-aperture distance is too great (>2-3 mm), the aqueous stream will flicker and may form a second node. (MOV 2845 kb)
Aqueous/oil interface during the transition from the PBS/PBS aqueous phase to the PBS/Lysis Buffer aqueous phase.
The introduction of surfactant into the aqueous phase greatly reduces the diameter of the stream. The transition is also typically accompanied by air bubbles that are introduced while switching syringes. Sample collection should begin once the bubbles cease and the stream becomes stable. (MOV 1053 kb)
Supplementary Methods
Bioinformatics script written in Python to parse and pair the VH and VL sequences following IMGT gene annotation. (TXT 17 kb)
Rights and permissions
About this article
Cite this article
McDaniel, J., DeKosky, B., Tanno, H. et al. Ultra-high-throughput sequencing of the immune receptor repertoire from millions of lymphocytes. Nat Protoc 11, 429–442 (2016). https://doi.org/10.1038/nprot.2016.024
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.024
This article is cited by
-
High-throughput single-cell activity-based screening and sequencing of antibodies using droplet microfluidics
Nature Biotechnology (2020)
-
Massively parallel single-cell B-cell receptor sequencing enables rapid discovery of diverse antigen-reactive antibodies
Communications Biology (2019)
-
Unbiased quantification of immunoglobulin diversity at the DNA level with VDJ-seq
Nature Protocols (2018)
-
Recombinant human B cell repertoires enable screening for rare, specific, and natively paired antibodies
Communications Biology (2018)
-
Identification of tumor-reactive B cells and systemic IgG in breast cancer based on clonal frequency in the sentinel lymph node
Cancer Immunology, Immunotherapy (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.