Abstract
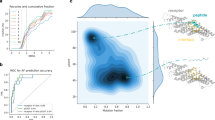
Peptide recognition modules (PRMs) are used throughout biology to mediate protein-protein interactions, and many PRMs are members of large protein domain families. Members of these families are often quite similar to each other, but each domain recognizes a distinct set of peptides, raising the question of how peptide recognition specificity is achieved using similar protein domains. The analysis of individual protein complex structures often gives answers that are not easily applicable to other members of the same PRM family. Bioinformatics-based approaches, one the other hand, may be difficult to interpret physically. Here we integrate structural information with a large, quantitative data set of SH2-peptide interactions to study the physical origin of domain-peptide specificity. We develop an energy model, inspired by protein folding, based on interactions between the amino acid positions in the domain and peptide. We use this model to successfully predict which SH2 domains and peptides interact and uncover the positions in each that are important for specificity. The energy model is general enough that it can be applied to other members of the SH2 family or to new peptides, and the cross-validation results suggest that these energy calculations will be useful for predicting binding interactions. It can also be adapted to study other PRM families, predict optimal peptides for a given SH2 domain, or study other biological interactions, e.g. protein-DNA interactions.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wunderlich, Z., Mirny, L. An optimized energy potential can predict SH2 domain-peptide interactions. Nat Prec (2008). https://doi.org/10.1038/npre.2008.1881.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2008.1881.1