Abstract
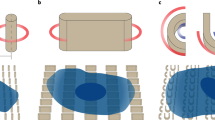
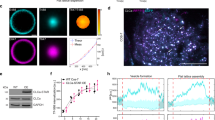
Clathrin-mediated endocytosis (CME) involves nanoscale bending and inward budding of the plasma membrane, by which cells regulate both the distribution of membrane proteins and the entry of extracellular species1,2. Extensive studies have shown that CME proteins actively modulate the plasma membrane curvature1,3,4. However, the reciprocal regulation of how the plasma membrane curvature affects the activities of endocytic proteins is much less explored, despite studies suggesting that membrane curvature itself can trigger biochemical reactions5,6,7,8. This gap in our understanding is largely due to technical challenges in precisely controlling the membrane curvature in live cells. In this work, we use patterned nanostructures to generate well-defined membrane curvatures ranging from +50 nm to −500 nm radius of curvature. We find that the positively curved membranes are CME hotspots, and that key CME proteins, clathrin and dynamin, show a strong preference towards positive membrane curvatures with a radius <200 nm. Of ten CME-related proteins we examined, all show preferences for positively curved membrane. In contrast, other membrane-associated proteins and non-CME endocytic protein caveolin1 show no such curvature preference. Therefore, nanostructured substrates constitute a novel tool for investigating curvature-dependent processes in live cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McMahon, H. T. & Boucrot, E. Molecular mechanism and physiological functions of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 12, 517–533 (2011).
Di Fiore, P. P. & von Zastrow, M. Endocytosis, signaling, and beyond. Cold Spring Harb. Perspect. Biol. 6, a016865 (2014).
Johannes, L., Wunder, C. & Bassereau, P. Bending ‘on the rocks’—a cocktail of biophysical modules to build endocytic pathways. Cold Spring Harb. Perspect. Biol. 6, a016741 (2014).
Kirchhausen, T., Owen, D. & Harrison, S. C. Molecular structure, function, and dynamics of clathrin-mediated membrane traffic. Cold Spring Harb. Perspect. Biol. 6, a016725 (2014).
McMahon, H. T. & Gallop, J. L. Membrane curvature and mechanisms of dynamic cell membrane remodelling. Nature 438, 590–596 (2005).
Liu, J., Sun, Y., Drubin, D. G. & Oster, G. F. The mechanochemistry of endocytosis. PLoS Biol. 7, e1000204 (2009).
Galic, M. et al. Dynamic recruitment of the curvature-sensitive protein ArhGAP44 to nanoscale membrane deformations limits exploratory filopodia initiation in neurons. eLife 3, e03116 (2014).
Larsen, J. B. et al. Membrane curvature enables N-Ras lipid anchor sorting to liquid-ordered membrane phases. Nat. Chem. Biol. 11, 192–194 (2015).
Epand, R. M., D'Souza, K., Berno, B. & Schlame, M. Membrane curvature modulation of protein activity determined by NMR. Biochim. Biophys. Acta. 1848, 220–228 (2015).
Iversen, L., Mathiasen, S., Larsen, J. B. & Stamou, D. Membrane curvature bends the laws of physics and chemistry. Nat. Cell Biol. 11, 822–825 (2015).
Wu, M. et al. Coupling between clathrin-dependent endocytic budding and F-BAR-dependent tubulation in a cell-free system. Nat. Cell Biol. 12, 902–908 (2010).
Lee, I.-H., Kai, H., Carlson, L.-A., Groves, J. T. & Hurley, J. H. Negative membrane curvature catalyzes nucleation of endosomal sorting complex required for transport (ESCRT)-III assembly. Proc. Natl Acad. Sci. USA 112, 15892–15897 (2015).
Galic, M. et al. External push and internal pull forces recruit curvature-sensing N-BAR domain proteins to the plasma membrane. Nat. Cell Biol. 14, 874–881 (2012).
Jeong, S., McDowell, M. T. & Cui, Y. Low-temperature self-catalytic growth of tin oxide nanocones over large areas. ACS Nano 5, 5800–5807 (2011).
Hanson, L., Lin, Z. C., Xie, C., Cui, Y. & Cui, B. Characterization of the cell-nanopillar interface by transmission electron microscopy. Nano Lett. 12, 5815–5820 (2012).
Mumm, F., Beckwith, K. M., Bonde, S., Martinez, K. L. & Sikorski, P. A transparent nanowire-based cell impalement device suitable for detailed cell-nanowire interaction studies. Small 9, 263–272 (2013).
Santoro, F. et al. Interfacing electrogenic cells with 3D nanoelectrodes: position, shape, and size matter. ACS Nano 8, 6713–6723 (2014).
Avinoam, O., Schorb, M., Beese, C. J., Briggs, J. A. & Kaksonen, M. Endocytic sites mature by continuous bending and remodeling of the clathrin coat. Science 348, 1369–1372 (2015).
Doyon, J. B. et al. Rapid and efficient clathrin-mediated endocytosis revealed in genome-edited mammalian cells. Nat. Cell Biol. 13, 331–337 (2011).
Taylor, M. J., Perrais, D. & Merrifield, C. J. A high precision survey of the molecular dynamics of mammalian clathrin-mediated endocytosis. PLoS Biol. 9, e1000604 (2011).
Ford, M. G. J. et al. Curvature of clathrin-coated pits driven by epsin. Nature 419, 361–366 (2002).
Peter, B. J. BAR domains as sensors of membrane curvature the amphiphysin BAR structure. Science 303, 495–499 (2004).
Kelly, B. T. et al. AP2 controls clathrin polymerization with a membrane-activated switch. Science 345, 459–463 (2014).
Henne, W. M. et al. FCHo proteins are nucleators of clathrin-mediated endocytosis. Science 328, 1281–1284 (2010).
Dautry-Varsat, A., Ciechanover, A. & Lodish, H. F. pH and the recycling of transferrin during receptor-mediated endocytosis. Proc. Natl Acad. Sci. USA 80, 2258–2262 (1983).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657 (2006).
Posor, Y. et al. Spatiotemporal control of endocytosis by phosphatidylinositol-3,4-bisphosphate. Nature 499, 233–237 (2013).
Chaudhary, N. et al. Endocytic crosstalk: cavins, caveolins, and caveolae regulate clathrin-independent endocytosis. PLoS Biol. 12, e1001832 (2014).
Grassart, A. et al. Actin and dynamin2 dynamics and interplay during clathrin-mediated endocytosis. J. Cell Biol. 205, 721–735 (2014).
Dominguez, R. & Holmes, K. C. Actin structure and function. Annu. Rev. Biophys. 40, 169–186 (2011).
Hanson, L. et al. Vertical nanopillars for in situ probing of nuclear mechanics in adherent cells. Nat. Nanotech. 10, 554–562 (2015).
Santoro, F. et al. Revealing the cell-material interface by FIB-SEM. BioRxiv http://dx.doi.org/10.1101/123794 (2017).
Gorfe, A. A. & Hocker, H. J. Membrane Targeting: Methods (John Wiley & Sons, 2001).
Lin, W. C., Yu, C. H. & Triffo, S. Supported membrane formation, characterization, functionalization, and patterning for application in biological science and technology. Curr. Protoc. Chem. Biol. 2, 235–269 (2010).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Aguet, F., Antonescu, C. N., Mettlen, M., Schmid, S. L. & Danuser, G. Advances in analysis of low signal-to-noise images link dynamin and AP2 to the functions of an endocytic checkpoint. Dev. Cell 26, 279–291 (2013).
Acknowledgements
We thank Y. Miao and S.H. Hong of the D.G.D. group in UC Berkeley for valuable discussion as well as helpful comments on genome-edited cell lines and endocytic lifetime analysis; K. Shen in Stanford for generous support on spinning disk confocal microscopy, M. Galic of the T. Meyer group in Stanford for suggestions and amphiphysin1-YFP plasmid; S. Guo of the B.C. group for constructing mCherry-CAAX plasmid, as well as A. McGuire, C. Xie and Z. Lin of the B.C. group in Stanford for advice and help on the nanostructure fabrication. We also thank Q. Ong and L. Kaplan of the B.C. group for comments on the manuscript. Fabrication and characterization of nanostructures were conducted in Stanford Nanofabrication Facility (SNF) and Stanford Nano Shared Facilities (SNSF). Spinning disk confocal with perfect focus for lifetime measurement was conducted in Cell Science Imaging Facility (CSIF) at Stanford University. This work was supported by the National Science Foundation (CAREER award no. 1055112), the National Institutes of Health (NIH; grant no. NS057906), a Searle Scholar award, a Packard Science and Engineering Fellowship (to B.C.), NIH fellowship 1F32 GM113379-01A1 (to J.R.M.), Studying Abroad Scholarship (to H.-Y. L.), Arnold O. Beckman Postdoctoral Fellowship (to M.A.), Heart Rhythm Research Fellowship (to F.S.) and the NIH grant R35GM118149 (to D.G.D.).
Author information
Authors and Affiliations
Contributions
W.Z., B.C., Y.C. and D.G.D conceived the study and designed the experiment. W.Z. fabricated the nanostructure substrates, and performed most of experiments. L.H. performed TEM measurements. F.S. conducted the FIB-SEM characterization. H.-Y.L. performed most of the endocytic protein test on nanobar arrays and the quantification and statistical analysis. W.Z., P.D.C. and B.C. developed the Matlab code for the dynamic analysis. W.Z. analysed most of the data. M.A. analysed the AP2/Dynamin2 movies. A.G. and J.R.M. provided and characterized the genome-edited cell line. W.Z. and B.C. wrote the manuscript. All the authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 1444 kb)
Supplementary information
Supplementary Movie 1 (MOV 2581 kb)
Supplementary information
Supplementary Movie 2 (MOV 9838 kb)
Supplementary information
Supplementary Movie 3 (AVI 11798 kb)
Supplementary information
Supplementary Movie 4 (MOV 18936 kb)
Rights and permissions
About this article
Cite this article
Zhao, W., Hanson, L., Lou, HY. et al. Nanoscale manipulation of membrane curvature for probing endocytosis in live cells. Nature Nanotech 12, 750–756 (2017). https://doi.org/10.1038/nnano.2017.98
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2017.98
This article is cited by
-
Molecular mechanism of GPCR spatial organization at the plasma membrane
Nature Chemical Biology (2024)
-
Identifying molecular tags selectively retained on the surface of brain endothelial cells to generate artificial targets for therapy delivery
Fluids and Barriers of the CNS (2023)
-
Sensing their plasma membrane curvature allows migrating cells to circumvent obstacles
Nature Communications (2023)
-
A mechanosensitive caveolae–invadosome interplay drives matrix remodelling for cancer cell invasion
Nature Cell Biology (2023)
-
Curved adhesions mediate cell attachment to soft matrix fibres in three dimensions
Nature Cell Biology (2023)