Abstract
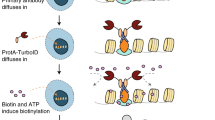
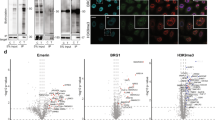
Although purification of biotinylated molecules is highly efficient, identifying specific sites of biotinylation remains challenging. We show that anti-biotin antibodies enable unprecedented enrichment of biotinylated peptides from complex peptide mixtures. Live-cell proximity labeling using APEX peroxidase followed by anti-biotin enrichment and mass spectrometry yielded over 1,600 biotinylation sites on hundreds of proteins, an increase of more than 30-fold in the number of biotinylation sites identified compared to streptavidin-based enrichment of proteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Green, N.M. Methods Enzymol. 184, 51–67 (1990).
Derakhshan, B., Wille, P.C. & Gross, S.S. Nat. Protoc. 2, 1685–1691 (2007).
Yang, J. et al. Nat. Protoc. 10, 1022–1037 (2015).
Hung, V. et al. Nat. Protoc. 11, 456–475 (2016).
Dieterich, D.C. et al. Nat. Protoc. 2, 532–540 (2007).
Lee, S.Y. et al. J. Am. Chem. Soc. 139, 3651–3662 (2017).
Szychowski, J. et al. J. Am. Chem. Soc. 132, 18351–18360 (2010).
Mertins, P. et al. Nat. Methods 10, 634–637 (2013).
Schiapparelli, L.M. et al. J. Proteome Res. 13, 3966–3978 (2014).
Hung, V. et al. Mol. Cell 55, 332–341 (2014).
Rhee, H.W. et al. Science 339, 1328–1331 (2013).
Loh, K.H. et al. Cell 166, 1295–1307.e21 (2016).
Mick, D.U. et al. Dev. Cell 35, 497–512 (2015).
Roux, K.J., Kim, D.I., Raida, M. & Burke, B. J. Cell Biol. 196, 801–810 (2012).
Schopfer, L.M., Champion, M.M., Tamblyn, N., Thompson, C.M. & Lockridge, O. Anal. Biochem. 345, 122–132 (2005).
Kamer, K.J. & Mootha, V.K. Nat. Rev. Mol. Cell Biol. 16, 545–553 (2015).
Baughman, J.M. et al. Nature 476, 341–345 (2011).
De Stefani, D., Raffaello, A., Teardo, E., Szabò, I. & Rizzuto, R. Nature 476, 336–340 (2011).
Martell, J.D. et al. Nat. Biotechnol. 30, 1143–1148 (2012).
UniProt Consortium. Nucleic Acids Res. 43, D204–D212 (2015).
Neupert, W. & Herrmann, J.M. Annu. Rev. Biochem. 76, 723–749 (2007).
Dimmer, K.S. et al. Hum. Mol. Genet. 17, 201–214 (2008).
Meier, S., Neupert, W. & Herrmann, J.M. J. Cell Biol. 170, 881–888 (2005).
Walker, J.E. & Dickson, V.K. Biochim. Biophys. Acta 1757, 286–296 (2006).
Rappsilber, J., Mann, M. & Ishihama, Y. Nat. Protoc. 2, 1896–1906 (2007).
Udeshi, N.D., Mertins, P., Svinkina, T. & Carr, S.A. Nat. Protoc. 8, 1950–1960 (2013).
Lam, S.S. et al. Nat. Methods 12, 51–54 (2015).
Calvo, S.E., Clauser, K.R. & Mootha, V.K. Nucleic Acids Res. 44, D1251–D1257 (2016).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E.L. J. Mol. Biol. 305, 567–580 (2001).
Acknowledgements
Portions of this work were supported by HHMI Collaborative Innovation Awards to S.A.C. and A.Y.T. (Norbert Perrimon PI). K.P. was supported by a National Science Foundation Graduate Research Fellowship and a Stanford Graduate Fellowship. T.A. is supported by the Helen Hay Whitney Foundation fellowship. V.K.M. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
N.D.U. and S.A.C. conceived of the study; N.D.U., K.P., T.S., and S.F. performed experiments; N.D.U., K.P., T.S., S.F., S.A.M., O.A., K.K., K.C., D.R., T.A., V.K.M., A.Y.T., and S.A.C. contributed to experimental design, data analysis, and data interpretation. N.D.U. and S.A.C. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Enrichment with anti-biotin antibody significantly increases detection of biotinylated peptides in complex samples.
(A) Structure of NHS-biotin (B) Results of spike-in studies. Bar plot shows the number of unique biotinylated peptides identified at each biotinylated:non-biotinylated peptide spike-in level. Error bars indicate standard deviation across three replicates of enrichment (C) Results of antibody titration experiments. Plot shows the number of distinct biotinylated peptides (red) and the number of distinct non-biotinylated peptides (black) from enrichment of 1 mg of peptides from 1:1000 spike-in samples. Error bars indicate range across two replicates of enrichment. (D) Results of testing different vendor antibodies.
Supplementary Figure 2 Labeling the mitochondrial matrix proteome in living cell.
(A) Anti V5- (top) and and anti-streptavidin (bottom) western blots of SILAC HEK 293T cells labeled with mitochondrial matrix-APEX and biotin phenol. A merged image is also shown. (B) Confocal fluorescence imaging of mitochondrial matrix-APEX2 labeling. Human embryonic kidney (HEK) 293T cells stably expressing matrix-APEX2 were incubated with biotin-phenol (BP) for 30 min and then treated live for 1 min with 1 mM H2O2 to initiate biotinylation (top row). A sample in which both BP and H2O2 were omitted was prepared in parallel as a negative control (bottom row). Cells were fixed and then stained with a NeutrAvidin-Alexa Fluor 647 (AF647) conjugate to visualize biotinylated proteins and an anti-V5 antibody to visualize matrix-APEX2 localization. DIC, differential interference contrast image. White horizontal scale bars = 25 μm.
Supplementary Figure 3 Analysis of SILAC data from the mitochondrial matrix streptavidin enriched proteomic experiments.
(A) Correlation between SILAC log2 H/L ratios for replicates 1, 2 and 3. A high correlation is seen, as expected. R^2 values are shown for all detected proteins. (B) Cutoff determination for the mitochondrial matrix streptavidin enriched proteomic experiment. Histograms (left panel) plot true positives (TP) in green, and false positives (FP) in red. Receiver-operating characteristic analysis (ROC, right panel) was performed for each replicate, setting the SILAC ratio cutoff value such that true positives were maximized and false positives were minimized.
Supplementary Figure 4 Fragment ions specific to biotin phenol can be used to enhance spectral interpretation.
(A) Structure of biotinylated tyrosine (B) Structure and mass of the immonium ion of biotinylated tyrosine (C) Structure and mass of the biotin phenol cleavage product produced upon HCD fragmentation.
Supplementary Figure 5 Majority of biotinylated peptides identified in antibody enrichment experiments are modified at tyrosine.
Bar plot shows the number of distinct biotinylation sites identified at tyrosine (Y), tryptophan (W), and cysteine (C) from SILAC labeled HEK 293T cells stably expressing matrix-APEX2. The percent of each site at a given amino acid is calculated relative to the total number of biotinylation sites.
Supplementary Figure 6 Conservative sorting model21 for LETM1.
(Left Panel) Many inner membrane proteins are imported via the Stop-Transfer pathway, where proteins are arrested at the TIM23 complex and laterally instead into the membrane. (Right Panel) Another, smaller class of inner membrane proteins are processed via a Conservative Sorting pathway. In this pathway, proteins initially translocate to the matrix prior to insertion in the inner membrane. We propose that biotinylation of Y141 may occur on a processing intermediate of LETM1 if the protein is processed via the Conservative Sorting pathway.
21Neupert, W. & Herrmann, J.M. Translocation of proteins into mitochondria. Annu Rev Biochem 76, 723-749 (2007).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 4 and 9 (PDF 1123 kb)
Supplementary Table 1
NHS-Biotin Peptide Spike-ins (XLSX 59612 kb)
Supplementary Table 2
Comparison of anti-biotin antibody and NeutrAvidin enrichment (XLSX 6946 kb)
Supplementary Table 3
Proteins: APEX labelling Streptavidin Enrichment (XLSX 189 kb)
Supplementary Table 5
Biotinylation Sites: APEX labelling Streptavidin Enrichment (XLSX 21 kb)
Supplementary Table 6
Biotinylation Sites: APEX labelling Anti-Biotin Antibody Enrichment; all (XLSX 625 kb)
Supplementary Table 7
Biotinylation Sites: APEX labelling Anti-Biotin Antibody Enrichment; >= 2 replicates (XLSX 373 kb)
Supplementary Table 8
Coverage of OXPHOS proteins (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Udeshi, N., Pedram, K., Svinkina, T. et al. Antibodies to biotin enable large-scale detection of biotinylation sites on proteins. Nat Methods 14, 1167–1170 (2017). https://doi.org/10.1038/nmeth.4465
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4465
This article is cited by
-
Mitochondrial matrix RTN4IP1/OPA10 is an oxidoreductase for coenzyme Q synthesis
Nature Chemical Biology (2024)
-
A modification-centric assessment tool for the performance of chemoproteomic probes
Nature Chemical Biology (2022)
-
Lima1 mediates the pluripotency control of membrane dynamics and cellular metabolism
Nature Communications (2022)
-
A proximity biotinylation-based approach to identify protein-E3 ligase interactions induced by PROTACs and molecular glues
Nature Communications (2022)
-
In vivo interactome profiling by enzyme‐catalyzed proximity labeling
Cell & Bioscience (2021)