Abstract
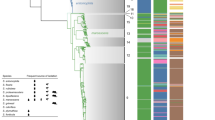
Genomic islands, such as pathogenicity islands, contribute to the evolution and diversification of microbial life1. Here we report on the Widespread Colonization Island, which encompasses the tad (tight adherence) locus for colonization of surfaces and biofilm formation by the human pathogen Actinobacillus actinomycetemcomitans. At least 12 of the 14 genes at the tad locus are required for tenacious biofilm formation and synthesis of bundled Flp pili (fibrils) that mediate adherence. The pilin subunit2, Flp1, remains inside the cell in tad-locus mutants, indicating that these genes encode a secretion system for export and assembly of fibrils. We found tad-related regions in a wide variety of Bacterial and Archaeal species3, and their sequence characteristics indicate possible horizontal transfer. To test the hypothesis of horizontal transfer, we compared the phylogeny of the tad locus to a robust organismal phylogeny using statistical tests of congruence and tree reconciliation techniques. Our analysis strongly supports a complex history of gene shuffling by recombination and multiple horizontal transfers, duplications and losses. We present evidence for a specific horizontal transfer event leading to the establishment of this region as a determinant of disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hacker, J. & Kaper, J.B. Pathogenicity islands and the evolution of microbes. Annu. Rev. Microbiol. 54, 641–679 (2000).
Inoue, T. et al. Molecular characterization of low-molecular-weight component protein, Flp, in Actinobacillus actinomycetemcomitans fimbriae. Microbiol. Immunol. 42, 253–258 (1998).
Kachlany, S.C., Planet, P.J., DeSalle, R., Fine, D.H. & Figurski, D.H. Genes for tight adherence of Actinobacillus actinomycetemcomitans: from plaque to plague to pond scum. Trends Microbiol. 9, 429–437 (2001).
Fine, D.H. et al. Colonization and persistence of rough and smooth colony variants of Actinobacillus actinomycetemcomitans in the mouths of rats. Arch. Oral Biol. 46, 1065–1078 (2001).
Kachlany, S.C. et al. flp-1, the first representative of a new pilin gene subfamily, is required for non-specific adherence of Actinobacillus actinomycetemcomitans. Mol. Microbiol. 40, 542–554 (2001).
Kachlany, S.C. et al. Nonspecific adherence by Actinobacillus actinomycetemcomitans requires genes widespread in bacteria and archaea. J. Bacteriol. 182, 6169–6176 (2000).
Fuller, T.E., Kennedy, M.J. & Lowery, D.E. Identification of Pasteurella multocida virulence genes in a septicemic mouse model using signature-tagged mutagenesis. Microb. Pathog. 29, 25–38 (2000).
Nika, J.R. et al. Haemophilus ducreyi requires the flp gene cluster for microcolony formation in vitro. Infect. Immun. 70, 2965–2975 (2002).
Skerker, J.M. & Shapiro, L. Identification and cell cycle control of a novel pilus system in Caulobacter crescentus. EMBO J. 19, 3223–3234 (2000).
Sommer, J.M. & Newton, A. Turning off flagellum rotation requires the pleiotropic gene pleD: pleA, pleC, and pleD define two morphogenic pathways in Caulobacter crescentus. J. Bacteriol. 171, 392–401 (1989).
Thomson, V.J., Bhattacharjee, M.K., Fine, D.H., Derbyshire, K.M. & Figurski, D.H. Direct selection if IS903 transposon insertions by use of a broad-host-range vector: isolation of catalase-deficient mutants of Actinobacillus actinomycetemcomitans. J. Bacteriol. 181, 7298–7307 (1999).
Haase, E.M., Zmuda, J.L. & Scannapieco, F.A. Identification and molecular analysis of rough-colony-specific outer membrane proteins of Actinobacillus actinomycetemcomitans. Infect. Immun. 67, 2901–2908 (1999).
Planet, P.J., Kachlany, S.C., DeSalle, R. & Figurski, D.H. Phylogeny of genes for secretion NTPases: identification of the widespread tadA subfamily and development of a diagnostic key for gene classification. Proc. Natl. Acad. Sci. USA 98, 2503–2508 (2001).
Thomas, N.A., Mueller, S., Klein, A. & Jarrell, K.F. Mutants in flaI and flaJ of the archaeon Methanococcus voltae are deficient in flagellum assembly. Mol. Microbiol. 46, 879–887 (2002).
Bhattacharjee, M.K., Kachlany, S.C., Fine, D.H. & Figurski, D.H. Nonspecific adherence and fibril biogenesis by Actinobacillus actinomycetemcomitans: TadA protein is an ATPase. J. Bacteriol. 183, 5927–5936 (2001).
Wang, Y., Goodman, S.D., Redfield, R.J. & Chen, C. Natural transformation and DNA uptake signal sequences in Actinobacillus actinomycetemcomitans. J. Bacteriol. 184, 3442–3449 (2002).
Doolittle, W.F. Phylogenetic classification and the universal tree. Science 284, 2124–2129 (1999).
Brochier, C., Bapteste, E., Moreira, D. & Philippe, H. Eubacterial phylogeny based on translational apparatus proteins. Trends Genet. 18, 1–5 (2002).
Jain, R., Rivera, M.C. & Lake, J.A. Horizontal gene transfer among genomes: the complexity hypothesis. Proc. Natl. Acad. Sci. USA 96, 3801–3806 (1999).
Farris, J.S., Kallersjo, M., Kluge, A.G. & Bult, C. Constructing a significance test for incongruence. Syst. Biol. 44, 570–572 (1995).
Brown, E.W., Kotewicz, M.L. & Cebula, T.A. Detection of recombination among Salmonella enterica strains using the incongruence length difference test. Mol. Phylogenet. Evol. 24, 102–120 (2002).
Dolphin, K., Belshaw, R., Orme, C.D. & Quicke, D.L. Noise and incongruence: interpreting results of the incongruence length difference test. Mol. Phylogenet. Evol. 17, 401–406 (2000).
Page, R.D. & Charleston, M.A. From gene to organismal phylogeny: reconciled trees and the gene tree/species tree problem. Mol. Phylogenet. Evol. 7, 231–240 (1997).
Charleston, M.A. Jungles: a new solution to the host/parasite phylogeny reconciliation problem. Math. Biosci. 149, 191–223 (1998).
Thornton, J.W. & DeSalle, R. A new method to localize and test the significance of incongruence: detecting domain shuffling in the nuclear receptor superfamily. Syst. Biol. 49, 183–201 (2000).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Nixon, K.C. The parsimony ratchet, a new method for rapid parsimony analysis. Cladistics 15, 407–414 (1999).
Schmidt, H.A., Strimmer, K., Vingron, M. & von Haeseler, A. TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics 18, 502–504 (2002).
Huelsenbeck, J.P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Bergey, D.H., Holt, J.G. & Krieg, N.R. Bergey's manual of systematic bacteriology vol. 4 (Williams & Wilkins, Baltimore, 1984).
Acknowledgements
We thank M. Charleston for early release of TreeMap2b and helpful discussion, I.N. Sarkar for computer assistance, the members of D.H. Figurski's, R.D.'s and D.H. Fine's laboratories for helpful comments and the multiple genome sequencing projects from which data were collected for this analysis. This work was funded by grants from the US National Institutes of Health (to D.H. Figurski and R.D.). R.D. is also supported by the Lewis B. and Dorothy Cullman Program for Molecular Systematics at the American Museum of Natural History. S.C.K. was partially supported by a training grant from the US National Institutes of Health to Columbia University. P.J.P. is supported by the Medical Scientist Training Program of Columbia University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Planet, P., Kachlany, S., Fine, D. et al. The Widespread Colonization Island of Actinobacillus actinomycetemcomitans. Nat Genet 34, 193–198 (2003). https://doi.org/10.1038/ng1154
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1154
This article is cited by
-
Distinctive gene and protein characteristics of extremely piezophilic Colwellia
BMC Genomics (2020)
-
Gene tree species tree reconciliation with gene conversion
Journal of Mathematical Biology (2019)
-
A Tad pilus promotes the establishment and resistance of Vibrio vulnificus biofilms to mechanical clearance
npj Biofilms and Microbiomes (2018)
-
Comparative genomics of Pseudomonas fluorescens subclade III strains from human lungs
BMC Genomics (2015)
-
Identification of putative adhesins of Actinobacillus suis and their homologues in other members of the family Pasteurellaceae
BMC Research Notes (2015)