Abstract
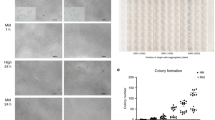
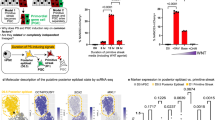
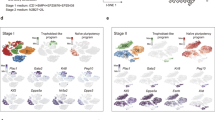
The generation of research-quality, clinically relevant cell types in vitro from human pluripotent stem cells requires a detailed understanding of the equivalent human cell types. Here we analysed 134 human embryonic and fetal samples from 6 to 20 developmental weeks and identified the stages at which cKIT+ primordial germ cells (PGCs), the precursors of gametes, undergo whole-genome epigenetic reprogramming with global depletion of 5mC, H3K27me3 and H2A.Z, and the time at which imprint erasure is initiated and 5hmC is present. Using five alternative in vitro differentiation strategies combined with single-cell microfluidic analysis and a bona fide human cKIT+ PGC signature, we show the stage of cKIT+ PGC formation in the first 16 days of differentiation. Taken together, our study creates a resource of human germ line ontogeny that is essential for future studies aimed at in vitro differentiation and unveiling the mechanisms necessary to pass human DNA from one generation to the next.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Ohinata, Y. et al. A signaling principle for the specification of the germ cell lineage in mice. Cell 137, 571–584 (2009).
Hayashi, K. et al. Offspring from oocytes derived from in vitro primordial germ cell-like cells in mice. Science 16, 971–975 (2012).
Reijo, R. et al. Mouse autosomal homolog of DAZ, a candidate male sterility gene in humans, is expressed in male germ cells before and after puberty. Genomics 35, 346–352 (1996).
Park, T. S. et al. Derivation of primordial germ cells from human embryonic and induced pluripotent stem cells is significantly improved by co-culture with human fetal gonadal cells. Stem Cells 27, 783–795 (2009).
Gaskell, T. L., Esnal, A., Robinson, L. L. L., Anderson, R. A. & Saunders, P. T. K. Immunohistochemical profiling of germ cells within the human fetal testis: identification of three subpopulations. Biol. Reprod. 71, 2012–2021 (2004).
Robinson, L., Gaskell, T., Saunders, P. & Anderson, R. Germ cell specific expression of c-kit in the human fetal gonad. Mol. Human Reprod. 7, 845 (2001).
Høyer, P. E., Byskov, A. G. & Møllgård, K. Stem cell factor and c-Kit in human primordial germ cells and fetal ovaries. Mol. Cell. Endocrinol. 234, 1–10 (2005).
Pauls, K. Spatial expression of germ cell markers during maturation of human fetal male gonads: an immunohistochemical study. Hum. Reprod. 21, 397–404 (2005).
Kerr, C., Hill, C., Blumenthal, P. & Gearhart, J. Expression of pluripotent stem cell markers in the human fetal ovary. Hum. Reprod. 23, 589–599 (2008).
Kerr, C. L., Hill, C. M., Blumenthal, P. D. & Gearhart, J. D. Expression of pluripotent stem cell markers in the human fetal testis. Stem Cells 26, 412–421 (2008).
Seki, Y. et al. Extensive and orderly reprogramming of genome-wide chromatin modifications associated with specification and early development of germ cells in mice. Dev. Biol. 278, 440–458 (2005).
Hajkova, P. et al. Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 452, 877–881 (2008).
Hajkova, P. et al. Genome-wide reprogramming in the mouse germ line entails the base excision repair pathway. Science 329, 78–82 (2010).
Popp, C. et al. Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463, 1101–1105 (2010).
Sinclair, A. H. et al. A gene from the human sex-determining region encodes a protein with homology to a conserved DNA-binding motif. Nature 346, 240–244 (1990).
Atlasi, Y., Mowla, S. J., Ziaee, S. A. M., Gokhale, P. J. & Andrews, P. W. OCT4 spliced variants are differentially expressed in human pluripotent and nonpluripotent cells. Stem Cells 26, 3068–3074 (2008).
Warthemann, R., Eildermann, K., Debowski, K. & Behr, R. False-positive antibody signals for the pluripotency factor OCT4A (POU5F1) in testis-derived cells may lead to erroneous data and misinterpretations. Mol. Hum. Reprod. 18, 605–612 (2012).
Ruzov, A. et al. Lineage-specific distribution of high levels of genomic 5-hydroxymethylcytosine in mammalian development. Cell Res. 21, 1332–1342 (2011).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Ko, M. et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468, 839–843 (2010).
Koh, K. P. et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Stem Cell 8, 200–213 (2011).
Thomson, J. A. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Diaz Perez, S. V. et al. Derivation of new human embryonic stem cell lines reveals rapid epigenetic progression in vitro that can be prevented by chemical modification of chromatin. Hum. Mol. Genet. 21, 751–764 (2012).
Clark, A. T. Spontaneous differentiation of germ cells from human embryonic stem cells in vitro. Hum. Mol. Genet. 13, 727–739 (2004).
Zwaka, T. P. A germ cell origin of embryonic stem cells? Development 132, 227–233 (2005).
Bao, S. et al. The germ cell determinant blimp1 is not required for derivation of pluripotent stem cells. Cell Stem Cell 11, 110–117 (2012).
Kamei, K. et al. Microfluidic image cytometry for quantitative single-cell profiling of human pluripotent stem cells in chemically defined conditions. Lab. Chip 10, 1113–1119 (2010).
Briddell, R. et al. Further phenotypic characterization and isolation of human hematopoietic progenitor cells using a monoclonal-antibody to the c-kit receptor. Blood 79, 3159–3167 (1992).
Anderson, R., Fulton, N., Cowan, G., Coutts, S. & Saunders, P. Conserved and divergent patterns of expression of DAZL, VASA and OCT 4 in the germ cells of the human fetal ovary and testis. BMC Dev. Biol. 7, 136 (2007).
Bhutani, N. et al. Reprogramming towards pluripotency requires AID-dependent DNA demethylation. Nature 463, 1042–1047 (2010).
Maiti, A. & Drohat, A. C. Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J. Biol. Chem. 286, 35334–35338 (2011).
Hajkova, P. et al. Epigenetic reprogramming in mouse primordial germ cells. Mech. Dev. 117, 15–23 (2002).
Ohinata, Y., Sano, M., Shigeta, M., Yamanaka, K. & Saitou, M. A comprehensive, non-invasive visualization of primordial germ cell development in mice by the Prdm1-mVenus and Dppa3-ECFP double transgenic reporter. Reproduction 136, 503–514 (2008).
Kurimoto, K. et al. Complex genome-wide transcription dynamics orchestrated by Blimp1 for the specification of the germ cell lineage in mice. Genes Dev. 22, 1617–1635 (2008).
Kee, K., Gonsalves, J. M., Clark, A. T. & Pera, R. A. R. Bone morphogenetic proteins induce germ cell differentiation from human embryonic stem cells. Stem. Cells Dev. 15, 831–837 (2006).
Kee, K., Angeles, V. T., Flores, M., Nguyen, H. N. & Pera, R. A. R. Human DAZL, DAZ and BOULE genes modulate primordial germ-cell and haploid gamete formation. Nature 462, 222–225 (2009).
Aflatoonian, B. et al. In vitro post-meiotic germ cell development from human embryonic stem cells. Hum. Reprod. 24, 3150–3159 (2009).
Tilgner, K. et al. Expression of GFP under the control of the RNA helicase VASA permits fluorescence-activated cell sorting isolation of human primordial germ cells. Stem Cells 28, 84–92 (2010).
Panula, S. et al. Human germ cell differentiation from fetal-and adult-derived induced pluripotent stem cells. Hum. Mol. Genet. 20, 752–762 (2011).
Medrano, J. V., Ramathal, C., Nguyen, H. N., Simon, C. & Reijo Pera, R. A. Divergent RNA-binding proteins, DAZL and VASA, induce meiotic progression in human germ cells derived in vitro. Stem Cells 30, 441–451 (2012).
Chuang, C. Y. et al. Meiotic Competent human germ cell-like cells derived from human embryonic stem cells induced by BMP4/WNT3A signaling and OCT4/EpCAM (epithelial cell adhesion molecule) selection. J. Biol. Chem. 287, 14389–14401 (2012).
Plath, K. Role of histone H3 Lysine 27 methylation in X inactivation. Science 300, 131–135 (2003).
Ficz, G. et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 473, 398–402 (2011).
Vincent, J. J. et al. Single cell analysis facilitates staging of blimp1-dependent primordial germ cells derived from mouse embryonic stem cells. PLoS ONE 6, e28960 (2011).
Boissonnas, C. et al. Specific epigenetic alterations of IGF2-H19 locus in spermatozoa from infertile men. Eue. J. Human Gene. 18, 73–80 (2009).
Geuns, E., Hilven, P., Van Steirteghem, A., Liebaers, I. & De Rycke, M. Methylation analysis of KvDMR1 in human oocytes. J. Med. Genet. 44, 144–147 (2006).
Kagami, M. et al. The IG-DMR and the MEG3-DMR at human chromosome 14q32.2: hierarchical interaction and distinct functional properties as imprinting control centers. PLoS Genet. 6, e1000992 (2010).
Zechner, U. et al. Quantitative methylation analysis of developmentally important genes in human pregnancy losses after ART and spontaneous conception. Mol. Hum. Reprod. 16, 704–713 (2010).
Acknowledgements
The authors would like to thank the UCLA Translational Pathology Core Laboratory and the UCLA Gene and Cellular Core Laboratory for some of the gonadal samples used in this study. We also thank J. Hargan-Calvopina, M. Oliveros-Etter and S. Diaz-Perez for critical reading of the manuscript, F. Codrea and J. Scholes for FACS and S. Peckman from the Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research for critical assistance with human subject and embryonic stem cell review. This work was supported primarily by fund number 1R01HD058047 from the Eunice Kennedy Shriver National Institute of Child Health & Human Development (NICHD; ATC), as well as the Iris Cantor-UCLA Women’s Health Pilot Project (ATC) and 1P01GM081621 from NIGMS. The Laboratory of Developmental Biology, University of Washington, Seattle is supported by NIH Award Number 5R24HD000836 from the NICHD. Human fetal tissue requests can be made to: bdrl@u.washington.edu.
Author information
Authors and Affiliations
Contributions
S.G. designed and performed the experiments, analysed data and wrote the manuscript; Z.L. performed flow analyses, RNA and DNA extraction for some gonadal samples, J.J.V. performed the single-cell analysis of female ovary at 14 weeks; K.X.Z. and M.P. performed the RNA-Seq data analysis; A.C. provided gonadal samples used in this study; A.T.C. designed the experiments, analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 937 kb)
Rights and permissions
About this article
Cite this article
Gkountela, S., Li, Z., Vincent, J. et al. The ontogeny of cKIT+ human primordial germ cells proves to be a resource for human germ line reprogramming, imprint erasure and in vitro differentiation. Nat Cell Biol 15, 113–122 (2013). https://doi.org/10.1038/ncb2638
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2638
This article is cited by
-
Histogenesis of intracranial germ cell tumors: primordial germ cell vs. embryonic stem cell
Child's Nervous System (2023)
-
In vitro differentiation of primed human induced pluripotent stem cells into primordial germ cell-like cells
Molecular Biology Reports (2023)
-
Porcine Primordial Germ Cell-Like Cells Generated from Induced Pluripotent Stem Cells Under Different Culture Conditions
Stem Cell Reviews and Reports (2022)
-
Proteome landscape and spatial map of mouse primordial germ cells
Science China Life Sciences (2021)
-
Female human primordial germ cells display X-chromosome dosage compensation despite the absence of X-inactivation
Nature Cell Biology (2020)