Abstract
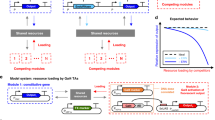
Engineering artificial gene networks from modular components is a major goal of synthetic biology. However, the construction of gene networks with predictable functions remains hampered by a lack of suitable components and the fact that assembled networks often require extensive, iterative retrofitting to work as intended. Here we present an approach that couples libraries of diversified components (synthesized with randomized nonessential sequence) with in silico modeling to guide predictable gene network construction without the need for post hoc tweaking. We demonstrate our approach in Saccharomyces cerevisiae by synthesizing regulatory promoter libraries and using them to construct feed-forward loop networks with different predicted input-output characteristics. We then expand our method to produce a synthetic gene network acting as a predictable timer, modifiable by component choice. We use this network to control the timing of yeast sedimentation, illustrating how the plug-and-play nature of our design can be readily applied to biotechnology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Andrianantoandro, E., Basu, S., Karig, D.K. & Weiss, R. Synthetic biology: new engineering rules for an emerging discipline. Mol. Syst. Biol. 2, 2006–0028 (2006).
Withers, S.T. & Keasling, J.D. Biosynthesis and engineering of isoprenoid small molecules. Appl. Microbiol. Biotechnol. 73, 980–990 (2007).
Weber, W. et al. A synthetic mammalian gene circuit reveals antituberculosis compounds. Proc. Natl. Acad. Sci. USA 105, 9994–9998 (2008).
Basu, S., Gerchman, Y., Collins, C.H., Arnold, F.H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
Lu, T.K. & Collins, J.J. Dispersing biofilms with engineered enzymatic bacteriophage. Proc. Natl. Acad. Sci. USA 104, 11197–11202 (2007).
Lu, T.K. & Collins, J.J. Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy. Proc. Natl. Acad. Sci. USA 106, 4629–4634 (2009).
Lee, S.K., Chou, H., Ham, T.S., Lee, T.S. & Keasling, J.D. Metabolic engineering of microorganisms for biofuels production: from bugs to synthetic biology to fuels. Curr. Opin. Biotechnol. 19, 556–563 (2008).
Guido, N.J. et al. A bottom-up approach to gene regulation. Nature 439, 856–860 (2006).
Hasty, J., McMillen, D. & Collins, J.J. Engineered gene circuits. Nature 420, 224–230 (2002).
Guet, C.C., Elowitz, M.B., Hsing, W. & Leibler, S. Combinatorial synthesis of genetic networks. Science 296, 1466–1470 (2002).
Levskaya, A. et al. Synthetic biology: engineering Escherichia coli to see light. Nature 438, 441–442 (2005).
Deans, T.L., Cantor, C.R. & Collins, J.J. A tunable genetic switch based on RNAi and repressor proteins for regulating gene expression in mammalian cells. Cell 130, 363–372 (2007).
Kramer, B.P. et al. An engineered epigenetic transgene switch in mammalian cells. Nat. Biotechnol. 22, 867–870 (2004).
Gardner, T.S., Cantor, C.R. & Collins, J.J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Weber, W. et al. A synthetic time-delay circuit in mammalian cells and mice. Proc. Natl. Acad. Sci. USA 104, 2643–2648 (2007).
Grilly, C., Stricker, J., Pang, W.L., Bennett, M.R. & Hasty, J. A synthetic gene network for tuning protein degradation in Saccharomyces cerevisiae. Mol. Syst. Biol. 3, 127 (2007).
Elowitz, M.B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008).
Tigges, M., Marquez-Lago, T.T., Stelling, J. & Fussenegger, M. A tunable synthetic mammalian oscillator. Nature 457, 309–312 (2009).
Marguet, P., Balagadde, F., Tan, C. & You, L. Biology by design: reduction and synthesis of cellular components and behaviour. J. R. Soc. Interface 4, 607–623 (2007).
Yokobayashi, Y., Weiss, R. & Arnold, F.H. Directed evolution of a genetic circuit. Proc. Natl. Acad. Sci. USA 99, 16587–16591 (2002).
Kobayashi, H. et al. Programmable cells: interfacing natural and engineered gene networks. Proc. Natl. Acad. Sci. USA 101, 8414–8419 (2004).
Acar, M., Becskei, A. & van Oudenaarden, A. Enhancement of cellular memory by reducing stochastic transitions. Nature 435, 228–232 (2005).
Blake, W.J. et al. Phenotypic consequences of promoter-mediated transcriptional noise. Mol. Cell 24, 853–865 (2006).
Pedraza, J.M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005).
Canton, B., Labno, A. & Endy, D. Refinement and standardization of synthetic biological parts and devices. Nat. Biotechnol. 26, 787–793 (2008).
Cox, R.S. III, Surette, M.G. & Elowitz, M.B. Programming gene expression with combinatorial promoters. Mol. Syst. Biol. 3, 145 (2007).
Gertz, J., Siggia, E.D. & Cohen, B.A. Analysis of combinatorial cis-regulation in synthetic and genomic promoters. Nature 457, 215–218 (2009).
Murphy, K.F., Balazsi, G. & Collins, J.J. Combinatorial promoter design for engineering noisy gene expression. Proc. Natl. Acad. Sci. USA 104, 12726–12731 (2007).
Alper, H., Moxley, J., Nevoigt, E., Fink, G.R. & Stephanopoulos, G. Engineering yeast transcription machinery for improved ethanol tolerance and production. Science 314, 1565–1568 (2006).
Jensen, P.R. & Hammer, K. Artificial promoters for metabolic optimization. Biotechnol. Bioeng. 58, 191–195 (1998).
Hammer, K., Mijakovic, I. & Jensen, P.R. Synthetic promoter libraries–tuning of gene expression. Trends Biotechnol. 24, 53–55 (2006).
Alper, H., Fischer, C., Nevoigt, E. & Stephanopoulos, G. Tuning genetic control through promoter engineering. Proc. Natl. Acad. Sci. USA 102, 12678–12683 (2005).
Jensen, P.R. & Hammer, K. The sequence of spacers between the consensus sequences modulates the strength of prokaryotic promoters. Appl. Environ. Microbiol. 64, 82–87 (1998).
Hillen, W. & Berens, C. Mechanisms underlying expression of Tn10 encoded tetracycline resistance. Annu. Rev. Microbiol. 48, 345–369 (1994).
Johnston, M. & Davis, R.W. Sequences that regulate the divergent GAL1–GAL10 promoter in Saccharomyces cerevisiae. Mol. Cell. Biol. 4, 1440–1448 (1984).
Cormack, B.P. et al. Yeast-enhanced green fluorescent protein (yEGFP)a reporter of gene expression in Candida albicans. Microbiology 143, 303–311 (1997).
Schirmaier, F. & Philippsen, P. Identification of two genes coding for the translation elongation factor EF-1 alpha of S. cerevisiae. EMBO J. 3, 3311–3315 (1984).
Floer, M., Bryant, G.O. & Ptashne, M. HSP90/70 chaperones are required for rapid nucleosome removal upon induction of the GAL genes of yeast. Proc. Natl. Acad. Sci. USA 105, 2975–2980 (2008).
Mangan, S. & Alon, U. Structure and function of the feed-forward loop network motif. Proc. Natl. Acad. Sci. USA 100, 11980–11985 (2003).
Scrable, H. & Stambrook, P.J. Activation of the lac repressor in the transgenic mouse. Genetics 147, 297–304 (1997).
Kaplan, S., Bren, A., Dekel, E. & Alon, U. The incoherent feed-forward loop can generate non-monotonic input functions for genes. Mol. Syst. Biol. 4, 203 (2008).
Entus, R., Aufderheide, B., Herbert, M. & Sauro, M.H. Design and implementation of three incoherent feed-forward motif based biological concentration sensors. Syst. Synth. Biol. 1, 119–128 (2007).
Tsang, J., Zhu, J. & van Oudenaarden, A. MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Mol. Cell 26, 753–767 (2007).
Wang, X., Hao, N., Dohlman, H.G. & Elston, T.C. Bistability, stochasticity, and oscillations in the mitogen-activated protein kinase cascade. Biophys. J. 90, 1961–1978 (2006).
Strogatz, S.H . Nonlinear Dynamics and Chaos: With Applications to Physics, Biology, Chemistry, and Engineering (Addison-Wesley, Reading, Massachusetts, 1994).
Guo, B., Styles, C.A., Feng, Q. & Fink, G.R. A Saccharomyces gene family involved in invasive growth, cell-cell adhesion, and mating. Proc. Natl. Acad. Sci. USA 97, 12158–12163 (2000).
Verstrepen, K.J. & Klis, F.M. Flocculation, adhesion and biofilm formation in yeasts. Mol. Microbiol. 60, 5–15 (2006).
Smukalla, S. et al. FLO1 is a variable green beard gene that drives biofilm-like cooperation in budding yeast. Cell 135, 726–737 (2008).
Rosenfeld, N., Young, J.W., Alon, U., Swain, P.S. & Elowitz, M.B. Accurate prediction of gene feedback circuit behavior from component properties. Mol. Syst. Biol. 3, 143 (2007).
Acknowledgements
We thank P.R. Jensen for advice on promoter library synthesis methods, K. Verstrepen for guidance and materials relating to yeast flocculation and H.H. Lee for valuable ideas in genetic device construction. This work was supported by the National Institutes of Health (NIH) through the NIH Director's Pioneer Award Program, grant number DP1 OD003644, the National Science Foundation Frontiers in Integrative Biological Research program and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
T.E., X.W. and J.J.C. designed the study; T.E. performed the experiments; X.W. conducted the modeling work; T.E., X.W. and J.J.C. analyzed the data and wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Methods (PDF 579 kb)
Rights and permissions
About this article
Cite this article
Ellis, T., Wang, X. & Collins, J. Diversity-based, model-guided construction of synthetic gene networks with predicted functions. Nat Biotechnol 27, 465–471 (2009). https://doi.org/10.1038/nbt.1536
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1536
This article is cited by
-
Generative Models
Erkenntnis (2023)
-
Engineering eukaryote-like regulatory circuits to expand artificial control mechanisms for metabolic engineering in Saccharomyces cerevisiae
Communications Biology (2022)
-
2D printed multicellular devices performing digital and analogue computation
Nature Communications (2021)
-
Exploring the effect of network topology, mRNA and protein dynamics on gene regulatory network stability
Nature Communications (2021)
-
Engineering living therapeutics with synthetic biology
Nature Reviews Drug Discovery (2021)