Abstract
Tropical mountains are hot spots of biodiversity and endemism1,2,3, but the evolutionary origins of their unique biotas are poorly understood4. In varying degrees, local and regional extinction, long-distance colonization, and local recruitment may all contribute to the exceptional character of these communities5. Also, it is debated whether mountain endemics mostly originate from local lowland taxa, or from lineages that reach the mountain by long-range dispersal from cool localities elsewhere6. Here we investigate the evolutionary routes to endemism by sampling an entire tropical mountain biota on the 4,095-metre-high Mount Kinabalu in Sabah, East Malaysia. We discover that most of its unique biodiversity is younger than the mountain itself (6 million years), and comprises a mix of immigrant pre-adapted lineages and descendants from local lowland ancestors, although substantial shifts from lower to higher vegetation zones in this latter group were rare. These insights could improve forecasts of the likelihood of extinction and ‘evolutionary rescue’7 in montane biodiversity hot spots under climate change scenarios.
Similar content being viewed by others
Main
In mountainous areas of the humid tropics, steep environmental gradients coincide with high primary productivity and relative climatic stability to sustain large numbers of species, often with striking degrees of endemism at higher elevations1,8. It has therefore been recognized that tropical mountains are biodiversity hot spots of great conservation value1, especially because endemics on mountain tops are vulnerable to becoming trapped and then annihilated as a result of global warming2,3.
The evolutionary origins of these unique biotas, however, are poorly understood4. Like other insular habitats9, the endemic biota of an isolated mountain results from complex dynamics among colonization, in situ speciation, and local extinction. Each of these factors is dependent on the age and size of the habitat, and on the environmental contrast between the insular habitat and its matrix10. In the case of a tropical mountain top, an added complication is the fact that climate fluctuations may have widened and restricted the geographic range over which the montane conditions have extended in the past, meaning that parts of the endemic biota may be relicts, and other components may be novel in character4,5.
Disentangling these possibilities for a single tropical montane biodiversity hot spot requires molecular phylogenetic study of a large number of fauna and flora elements. However, with only few exceptions11,12, evolutionary studies in such hot spots have been limited to single taxa13,14. This precludes broad understanding of the evolutionary and biogeographic origins of an endemic biota as a whole4.
We investigated the evolutionary routes to endemism by sampling an entire tropical mountain biota on the UNESCO World Heritage site of Gunung Kinabalu in Sabah, East Malaysia. We demonstrate that most of its unique biodiversity is younger than the mountain itself and comprises a mix of immigrant pre-adapted lineages as well as descendants from local lowland ancestors.
At 4,095 m, Kinabalu is the tallest mountain between the Himalayas and New Guinea. It is a solitary ‘sky island’, having emerged during the Pliocene and early Pleistocene as a granite pluton within the surrounding sandstone of the Crocker Range, the latter having formed much earlier, between the Eocene and the early Miocene15. Because of the area’s tectonic activity, as well as Pleistocene sea level changes, the exact historical progression of its elevation above sea level is not known, but it is likely that a major rise, even beyond today’s elevation, of Kinabalu, as well as the central spine of the Crocker Range, took place between 6 million years ago and today (for more geological background see Methods). Since the early days of its exploration16, Kinabalu has been famous for its extremely high biological diversity, especially its richness in endemic species, with endemism proportions reaching 25–30% for some taxa17.
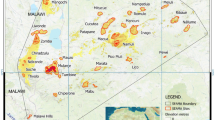
To unravel the origins of the exceptionally rich Kinabalu biota, we mounted a Malaysian–Dutch expedition in which 47 taxonomists worked at 37 localities, spanning the full range of elevations (Fig. 1). We used Sanger sequencing to sequence one or more fast-evolving loci for 1,852 individuals, belonging to 18 genera representing gastropods, annelids, insects, arachnids, vertebrates, pteridophytes, bryophytes, and angiosperms. We also obtained 3.7 million basidiomycete and glomeromycete ITS2 rDNA sequences from soil cores with ion semiconductor sequencing. In addition, we retrieved data from eight previously published single-taxon studies on vascular plants.
Inset left, location of the study area in the World and in Borneo. Inset middle, detail of the summit trail in Kinabalu Park. The eight expedition stations in Kinabalu Park and Crocker Range Park are indicated with red markers, ten additional sampling sites with blue markers. Not indicated separately are 15 sites along the summit trail, and four sites very close to Mahua, Gunung Alab, and Inobong.
We analysed all data within a phylogenetic framework to estimate the times of origin of endemic species, and to determine whether endemic species had descended from local or distant congeners (Methods, Fig. 2, Extended Data Table 1 and Extended Data Figs 1, 2, 3, 4, 5, 6, 7, 8, 9). Note that we define ‘endemic’ as restricted to the area in which our expedition took place. Although the present study offers the most comprehensive evolutionary analysis of any mountain biota to date, the taxa covered are, by necessity, an uneven and fragmentary sampling of the full diversity. Nonetheless, we expect that our results are representative for the Kinabalu biota as a whole, as our selected taxa encompass organisms with a wide variety of phylogenetic backgrounds, ecologies, and life history traits.
a, Centric endemism (Ptomaphaginus). b, Eccentric endemism (Impatiens). c, Centric endemism superimposed on eccentric endemism (detail of the Cortinarius phylogeny). Clades are coloured with the same colour scheme as in Fig. 1, by elevational zone at which they occur (see legend). High-elevation endemics in a and b are indicated in red. For support values and other details on these and other trees, see Extended Data Figs 1, 2, 3, 4.
Similar to Mesoamerican endemic cloud forest seed plants and vertebrates11, our molecular dating results show that the estimated mean stem-node ages of 33 endemic species span a wide range, from 0.12 million years to 14.6 million years (Fig. 3). However, 76% of these fall within the past 6 million years, the time span during which Kinabalu is likely to have reached its present elevation. Only two endemics, the frog Kalophrynus baluensis and the flowering plant Ilex kinabaluensis, are markedly older than the mountain itself. These may be explained as artefactual if we failed to identify the closest non-endemic sister lineage, thereby inflating their reconstructed age, or if these species are actually not endemics, but more widespread. Alternatively, they may truly be old endemics that evolved during cooler periods at lower elevations in Borneo before Kinabalu’s formation.
Elevations (mid-points, minima and maxima) and dates of origination derived from molecular dating (averages and 95% credible intervals) for endemic species; eccentric species (see main text) are indicated with a black circle. The vertical dashed line indicates the oldest possible date for Kinabalu to have reached its current elevation. For details, see Extended Data Figs 5 and 6.
Our phylogenetic and biogeographic analyses (Extended Data Table 1 and Extended Data Figs 1, 2, 3, 4, 7, 8, 9) suggest the existence of two categories of endemics18: ‘eccentric’ (12 taxa) and ‘centric’ (25 taxa). The eccentric type of endemic has sister taxa that occur either in temperate climates (seven cases) or in other tropical mountains outside of Borneo (five cases). To this group belong all bryophytes, pteridophytes, some of the fungal lineages and also the endemics in the flowering plant groups Hedyotis, Ilex, Impatiens, Ranunculus and Euphrasia, and the animals Coeliccia and Tritetrabdella. Eccentric endemics predominantly occur at high elevations (mean lower elevational boundary, 2,212 m; s.d., 837 m), they are strict Kinabalu endemics (they do not occur on nearby, lower mountains), and are further characterized by high dispersal capacities (one, two and seven clades with eccentric endemism have small, medium and large dispersal, respectively).
The centric type of endemic has sister taxa that occur locally, in the Borneo lowlands. Six out of the eight endemic animal species, some of the fungal lineages, the conifers, and 17 of the 20 flowering plants belong to this type. Centric endemics occur on average at lower elevations (mean lower elevational boundary, 1,724 m; s.d., 728 m) and tend to have lower dispersal capacities than the eccentric endemics (eight, seven and ten clades with centric endemism have small, medium and large dispersal, respectively). Some centric endemics are not strict Kinabalu endemics, as they are also found on other mountains in Kinabalu’s vicinity. Our ancestral state reconstructions (Supplementary Table 6) confirm a pattern of altitudinal bottom-up cladogenesis in the centric endemics, with 8 out of 11 species ranging partially or entirely outside of the 95% credibility interval for the reconstructed elevation of the most recent common ancestor, and the remaining 3 out of 11 falling entirely within this interval. When we performed the same analysis with Kitayama’s (ref. 19) seven vegetation zones, rather than elevation, we found that only three of these eight elevation shifts may represent a shift towards a higher vegetation zone. This suggests that, even in high-elevation centric endemics, substantial niche shifts are rare. Niche conservatism is probably even underestimated, since the Massenerhebung effect causes an upward shift of vegetation zones on Kinabalu compared with other Bornean mountains14,20.
Our multi-taxon study shows that Kinabalu’s biodiversity hot spot is of recent origin. This means the mountain is an evolutionary cradle, accumulating neo-endemics, as has been suggested for other young, high-elevation biodiversity hot spots at low latitudes21, such as the Tibetan plateau22, the Andean highlands8, and Afrotropical volcanoes23. However, probably as a consequence of the rapid emergence of the mountain15 and its unique alpine summit conditions19, many of these neo-endemics have not evolved by drastic niche shifts from local ancestors, but rather by immigration of pre-adapted propagules from elsewhere. This explains the multiple independent colonization events in some taxa (for example, Glomus, Rhododendron and Coeliccia). In addition, local lowland taxa have also generated montane species, but although some of these have reached vegetation zones above 2,000 m, most do not show substantial niche shifts away from their ancestral niche. The fact that the endemic biota of Kinabalu appears to be composed largely of pre-adapted (eccentric) species and locally derived (centric), ecologically conserved endemics is in line with niche conservatism24.
We suggest that our novel approach of molecular dating of multiple clades be applied to larger communities in this and other tropical montane biodiversity hot spots25. In combination, such information should allow a detailed dissection of the relative roles of ecological speciation, colonization and habitat filtering in the formation of endemic biotas in this and other tropical mountains. Moreover, such understanding could improve predictions of the likelihood of extinction and evolutionary rescue of endemic species experiencing changing climate conditions7.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Expedition
Material was sampled chiefly between September 10th and 25th, 2012, by the participants of the Crocker Range/Kinabalu Scientific Expedition, which was conceived, organized, funded and conducted jointly by Sabah Parks (Malaysia) and Naturalis Biodiversity Center (The Netherlands). The expedition was set up with the explicit aim to investigate evolutionary patterns of endemism (see main text), and participants were selected to maximize taxon coverage and ecological diversity. The 47 expedition members were asked to identify one or more congeneric species complexes (hereafter termed ‘genera’) within their taxon of expertise that included high elevation endemics and preferably also more widespread lowland species (although there is a possibility that this approach would leave out any monotypic endemics, in the groups studied by us, such species are not known). These, plus outgroup species, were then sampled (using taxon-specific methods; see Supplementary Table 2 for further details) at eight expedition stations located at a range of elevations (Fig. 1), and also in additional locations in both parks and their vicinity. Data on time, latitude and longitude, elevation, and habitat were collected with each sample. Material was subsampled for DNA analysis, mostly in the field, by preserving tissues or entire organisms, usually either in pure ethanol or in silica gel (see Supplementary Table 2 for taxon-specific subsampling and storage). In the case of fungi, mycelial DNA was obtained by bulk extraction from composite soil cores (see Supplementary Table 2 and below). In addition to the data collected during the expedition, the expedition members were asked to provide a complete assessment of elevational and geographical distribution for each of the species within their taxon of expertise, based on published sources and previous collections.
Molecular analysis
Upon arrival in the Naturalis DNA barcoding facility, subsamples were processed in a semi-automated way. DNA extractions on animal material were performed on either tissue samples or individual arthropod legs, or using whole arthropods in a non-destructive extraction method. DNA was extracted using the NucleoMag 96 Tissue kit (Macherey-Nagel Gmbh & Co., Düren, Germany) on the KingFisher Flex magnetic particle processor (Thermo Scientific). DNA extractions on plant tissue were performed on dried leaf samples, using the NucleoMag 96 Plant kit (Macherey-Nagel Gmbh & Co., Düren, Germany). The KingFisher Flex protocols were slightly adjusted from the manufacturer’s protocol (http://www.thermoscientific.com) to include longer mixing of beads with lysed sample and a drying step before elution. A volume of 150 µl was used for elution in both animal and plant samples. Soil samples containing fungal mycelia were processed by pooling 40 cores (for sampling details, see Supplementary Table 2), resulting in a composite soil sample for each site. Genomic DNA was extracted from 1 g of this sample using the NucleoSpin Soil kit (Macherey-Nagel Gmbh & Co., Düren, Germany), according to the manufacturer’s protocol. For each sample DNA extraction was carried out twice and replicates were combined.
For each specimen, the accepted barcode marker for its taxon was sequenced, in many cases augmented with one or more other phylogenetically informative loci. Supplementary Tables 1 and 2 provide lists of the loci and primers used for each genus included in the study. Where relevant for outgroup comparison or further geographic resolution, we then obtained additional sequences from Borneo or elsewhere from GenBank or from our own, unrelated studies. GenBank accession codes for these are given in Supplementary Table 2. After sequencing, (remains of) voucher specimens were mostly deposited in the collections of Sabah Parks and/or Naturalis Biodiversity Center. Voucher collection codes are listed in Supplementary Table 2.
PCR reactions generally contained 18.75 µl ultrapure H2O, 2.5 µl 10× PCR buffer, 1.0 µl 10 mM of each primer, 0.5 µl 2.5 mM dNTPs and 0.25 µl 5U µl−1 Qiagen Taq. Most PCRs were performed using an initial denaturation step of 180 s at 94 °C, followed by 40 cycles of 15 s at 94 °C, 30 s at 50 °C and 40 s at 72 °C, and ended with a final extension of 300 s at 72 °C and pause at 12 °C. Bi-directional Sanger sequencing was performed at either Macrogen (http://www.macrogen.com) or BaseClear (http://www.baseclear.com). Sequences were edited manually (primers trimmed off and obvious reading errors corrected) using Sequencher 4.10.1 (Gene Codes Corporation). For the fungal soil samples, the amplicon library was sequenced using an Ion 318TM Chip in an Ion Torrent Personal Genome Machine (PGM; Life Technologies, Guilford, CT, USA). PCR, sequencing, raw data filtering and all bioinformatic analyses were performed as described in detail previously26. From these fungal soil sequences, we selected non-redundant sequences of the ectomycorrhizal genus Cortinarius and the arbuscular mycorrhizal genus Glomus because they were distributed along the entire altitudinal range of our sampling.
Phylogenetic analysis
We performed initial exploratory alignment and created neighbour-joining trees. This allowed an assessment of individuals that (on the basis of cursory morphological examination) had been included as potential (undescribed) members of the species complexes of choice. In some cases, the sequence analysis showed that these individuals were in fact unrelated and such sequences were then removed before further analysis. Then, using Geneious (version 6.0.5; created by Biomatters; available from http://www.geneious.com/) as a workbench, sequence data were aligned with MUSCLE25. We ran ModelTest27 to determine, using Akaike Information Criterion (AIC), the best model of nucleotide substitution per alignment, for each genetic marker separately. We then performed a Bayesian analysis using MrBayes 3.2.2 (ref. 28) with settings for multiple markers unlinked, running two replicates of a minimum of 107 generations each. Convergence was checked in Tracer (v1.5; ref. 29) and, after discarding a 25% burn-in, the resulting majority-rule consensus trees were preserved. For the fungal data sets, multiple sequence alignments were built using MUSCLE and, due to the large size of these data sets, phylogenetic analyses of maximum likelihood (ML) were carried out in Garli 2.0 (ref. 30) using the GTR+I+G model with estimated values of the alpha parameter of the gamma distribution of variable sites (G) and proportion of invariable sites (I). Clade support was assessed by non-parametric bootstrap analyses with 100 pseudo-replicates.
Molecular dating
The times of divergence of Mount Kinabalu endemics from their most closely related extant relatives were estimated with a molecular clock approach. We included not only our own data, but also incorporated published studies on genera other than the ones we sampled. For each genus, taxon sampling and molecular data used are specified in Supplementary Table 4. All data sets were assembled in Geneious (ver. 5.5.6; created by Biomatters; available from http://www.geneious.com/). Data sets were pruned to contain a single taxon per species by selecting a random sequence from each species. Sequence alignments were obtained with the MAFFT (ver. 6.814b) alignment tool implemented in Geneious. Divergence time estimates were obtained using a relaxed molecular clock approach with BEAST (ver. 2.1.3; ref. 31). For each DNA region used, an unlinked substitution model was implemented as selected with jModeltest (ver. 2.1.5; ref. 27) under the AIC. For the tree prior, a Yule birth rate prior was selected with a flat prior on the Yule birth rate. A single clock and tree model was used for each analysis. All analyses were run with an uncorrelated log-normal clock model, to which an exponential prior was assigned for both the mean (ucldMEAN; mean set at 10.0) and the standard deviation (ucldStdev; mean set at 0.33). Time calibration was done using: (1) fossil constraints; (2) secondary calibration points; or (3) substitution rates, depending on the information available for each particular genus. In cases where suitable fossil or secondary calibration information was not available, we used a priori information about the substitution rate of one of the gene partitions—as opposed to the ‘fix the mean substitution rate’ option across all partitions—to optimize the analyses. The full details on calibration methods used for each taxon are given in Supplementary Table 5. For each run, posterior distributions of parameters were approximated using a Markov chain Monte Carlo analysis of 5 × 107 generations, sampling every 5,000 generations, discarding a burn-in of 5 × 106 generations (10%). Convergence of the chains was checked by evaluating the ESS values of each parameter with Tracer v1.6.
Biogeographical, elevational and vegetation zone analyses
For each genus, an independent estimation of the dispersal neighbourhood size was obtained from the spatial ecology literature on the genus concerned or related genera, and/or expert knowledge. We used this information to classify dispersal neighbourhood (two standard deviations for the distribution of distances between birth and reproduction for individuals of a certain species) in the genus in question as ‘small’, <1 km (mostly flightless animals), ‘medium’, 1–100 km (flying or ballooning animals, and plants dispersed by flying animals), or ‘large’, >100 km (plants and fungi with very small wind-dispersed seeds or spores). An overview of dispersal assessments is in Supplementary Table 3.
Ancestral ranges were estimated for each taxon group under maximum likelihood with BioGeoBears32 using the three-parameter DEC+J model. The ultrametric tree from the Bayesian relaxed molecular clock analysis was used as input tree. This tree was pruned to include the clade with the endemic species and at least one deeper node (for most groups two deeper nodes were included). Each species was scored to be absent or present in Mount Kinabalu (K), Borneo (B), and the rest of the world (W). For each group the analysis was run with equal probability for exchange between the three areas over the entire diversification time of the tree. The maximum number of areas was assigned as the maximum number of areas observed in the extant taxa (either 2 or 3). The optimizations, with the ancestral range with the highest likelihood for each node, are shown in Extended Data Figs 7, 8, 9). We classify a ‘W’ to ‘K’ or ‘KW’ shift in the lineage leading to a species endemic to Kinabalu as ‘eccentric endemism’. All other ancestral ranges leading to Kinabalu endemics are scored as ‘centric’.
For each genus sampled during the expedition that showed centric endemism, we added the elevation at which it had been collected as a continuous character. In BayesTraits 2.0 (ref. 33) we reconstructed ancestral states for elevation at the node leading to the endemic species and its sister clade using the continuous random walk model which implements a generalized least squares model for continuously varying traits. The elevation of the most-recent common ancestor of the node of interest was inferred using a MCMC approach on a set of 2,700 trees randomly sampled after the phylogenetic inference with MrBayes (excluding burn-in). The MCMC chain of BayesTraits was run for 106 generations, discarding the results sampled during the first 105 generations as burn-in. We then inspected whether the current range of elevation of the endemic species stretches beyond the 95% credible interval for the most-recent common ancestor. If it stretches above this interval, we considered this indicative of a ‘bottom-up’ scenario, whereas the reverse would suggest a ‘top-down’ scenario. The results are summarized in Supplementary Table 6.
In parallel with the elevational analysis, we conducted an analysis in which we coded the vegetation belt(s) as taxon traits. We used the zones defined for Mt. Kinabalu in ref. 19, namely: (i) lowland vegetation (below 600 m); (ii) hill vegetation (600–1,200 m); (iii) lower montane vegetation (1,200–2,000 m); (iv) upper montane vegetation (2,000–2,800 m); (v) subalpine vegetation (2,800–3,400 m); (vi) upper subalpine vegetation (3,400–3,700 m); (vii) alpine vegetation (above 3,700 m). The probabilities at each vegetation belt for the most-recent common ancestor of the node leading to the endemic clade and its sister group were inferred with the continuous-time Markov model for discrete data implemented in BayesTraits 2.0. The same input trees were used as for the elevation reconstruction, and the MCMC was run for 106 generations, discarding the results sampled during the first 105 generations as burn-in. The resulting probabilities are shown in Supplementary Table 7.
Geological summary
The following is based on refs 15,34 and R. Hall, personal communication, 2014.
Orogeny in the region began at about 23 million years ago (Mya) as a result of the northward movement of the Australian plate leading to the collision of Australia and Southeast Asia in what today is eastern Indonesia. Orogeny in the Kinabalu area began at about 20 Mya after collision of north Borneo with the South China continental crust. This led to the emergence of most of Sabah. Soon after this emergence much of Sabah became submerged, although a north–south-oriented mountain belt formed of tightly folded mudstones and sandstones (the future Crocker Range) remained. At the same time, the climate changed from seasonal to ever-wet. Mapping of former shelf edges offshore shows that later the Crocker Range widened with time, implying that it became higher in its central parts. Between 8 and 7 Mya a granitic body intruded the crust and cooled rapidly from ∼7 to 5 Mya. Melting could have resulted from a build-up of heat from radioactive decay in the deep crust, or from decompression melting following extension. After 6 Mya (the precise timing and duration is not known) the area (including the surrounding Crocker Range) rose further, elevating to the point that it was glaciated recently, possibly after a lithospheric root detached and sank into the deeper mantle, leading new mantle material to flow into the vacated space below the Crocker Range. Extensive glacial weathering of the summit suggests the mountain had reached its present elevation by at least 1 Mya. The Pinosuk gravels on Kinabalu’s south-eastern slope are the accumulated erosional debris of young glaciation. Today, the mountain is still rising by 0.5 mm per year.
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
GenBank accession numbers for newly determined sequences are: KF369344, KF492848, KF839944–KF839949, KM378672–KM378719, KM489525–KM489544, KM982697–KM982715, KP152323–KP152394, KP978313–KP979097, KR002164–KR002181, KR030167–KR030238 and KR232819–KR232836. A large portion of the data is also publicly available on BOLD (http://www.boldsystems.org), as dataset DS–KINA (http://dx.doi.org/10.5883/DS-KINA). Voucher specimens have been archived in public natural history collections.
References
Körner, C. & Spehn, E. M. Mountain Biodiversity: a Global Assessment (Parthenon, Boca Raton, 2002)
Chen, I. C. et al. Elevation increases in moth assemblages over 42 years on a tropical mountain. Proc. Natl Acad. Sci. USA 106, 1479–1483 (2009)
La Sorte, F. A. & Jetz, W. Projected range contractions of montane biodiversity under global warming. Proc. R. Soc. Lond. B 277, 3401–3410 (2010)
Graham, C. H. et al. The origin and maintenance of montane diversity: integrating evolutionary and ecological processes. Ecography 37, 711–719 (2014)
Cadena, C. D. et al. Latitude, elevational climatic zonation and speciation in New World vertebrates. Proc. R. Soc. Lond. B 279, 194–201 (2012)
Rodríguez-Castañeda, G. et al. Tropical forests are not flat: how mountains affect herbivore diversity. Ecol. Lett. 13, 1348–1357 (2010)
Schiffers, K., Borne, E. C., Lavergne, S., Thuiller, W. & Travis, J. M. J. Limited evolutionary rescue of locally adapted populations facing climate change. Phil. Trans. R. Soc. Lond. B 368, 20120083 (2013)
Hoorn, C., Mosbrugger, V., Mulch, A. & Antonelli, A. Biodiversity from mountain building. Nat. Geosci. 6, 154 (2013)
Warren, B. H. et al. Islands as model systems in ecology and evolution: prospects fifty years after MacArthur–Wilson. Ecol. Lett. 18, 200–217 (2015)
Watson, D. M. A conceptual framework for studying species composition in fragments, islands and other patchy ecosystems. J. Biogeogr. 29, 823–834 (2002)
Ornelas, J. F. et al. Comparative phylogeographic analyses illustrate the complex evolutionary history of threatened cloud forests of northern Mesoamerica. PLoS ONE 8, e56283 (2013)
Madriñán, S., Cortés, A. J. & Richardson, J. E. Páramo is the world’s fastest evolving and coolest biodiversity hotspot. Front. Genet. 4, 192 (2013)
Barkman, T. J. & Simpson, B. B. Origin of high-elevation Dendrochilum species (Orchidaceae) endemic to Mount Kinabalu, Sabah, Malaysia. Syst. Bot. 26, 658–669 (2001)
Gawin, D. F. et al. Patterns of avian diversification in Borneo: the case of the endemic Mountain Black-eye (Chlorocharis emiliae). Auk 131, 86–99 (2014)
Cottam, M. A. et al. Neogene rock uplift and erosion in northern Borneo: evidence from the Kinabalu granite, Mount Kinabalu. J. Geol. Soc. Lond. 170, 805–816 (2013)
Whitehead, J. Exploration of Mount Kina Balu, North Borneo (Gurney & Jackson, London, 1893)
Wong, K. M. & Phillips, A. Kinabalu, Summit of Borneo (Sabah Society & Sabah Parks, Kota Kinabalu, 1996)
van Steenis, C. G. G. J. Plant geography of the mountain flora of Mt. Kinabalu. Proc. R. Soc. Lond. B 161, 7–38 (1964)
Kitayama, K. An altitudinal transect study of the vegetation on Mount Kinabalu, Borneo. Vegetatio 102, 149–171 (1992)
Grubb, P. J. Interpretation of Massenerhebung effect on tropical mountains. Nature 229, 44–45 (1971)
Favre, A. et al. The role of the uplift of the Qinghai–Tibetan Plateau for the evolution of Tibetan biotas. Biol. Rev. Camb. Philos. Soc. 90, 236–253 (2015)
Schwery, O. et al. As old as the mountains: the radiations of the Ericaceae. New Phytol. 207, 355–367 (2015)
Price, T. D. et al. Niche filling slows the diversification of Himalayan songbirds. Nature 509, 222–225 (2014)
Crisp, M. D. et al. Phylogenetic biome conservatism on a global scale. Nature 458, 754–756 (2009)
Culmsee, H. & Leuschner, C. Consistent patterns of elevational change in tree taxonomic and phylogenetic diversity across Malesian mountain forests. J. Biogeogr. 40, 1997–2010 (2013)
Geml, J. et al. Large-scale fungal diversity assessment in the Andean Yungas forests reveals strong community turnover among forest types along an altitudinal gradient. Mol. Ecol. 23, 2452–2472 (2014)
Posada, D. & Crandall, K. A. Modeltest: testing the model of DNA substitution. Bioinformatics 14, 817–818 (1998)
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003)
Rambaut, A. & Drummond, A. J., Tracer v1.5. http://beast.bio.ed.ac.uk/Tracer (2009)
Zwickl, D. J. Genetic Algorithm Approaches for the Phylogenetic Analysis of Large Biological Sequence Datasets under the Maximum Likelihood Criterion (The University of Texas, Austin, 2006)
Bouckaert, R. et al. BEAST 2: a software platform for Bayesian evolutionary analysis. PLOS Comput. Biol. 10, e1003537 (2014)
Matzke, N. J. Probabilistic historical biogeography: new models for founder-event speciation, imperfect detection, and fossils allow improved accuracy and model-testing. Front. Biogeogr. 5, 242–248 (2013)
Meade, A. & Pagel M. BayesTraits, version 2.0. http://www.evolution.rdg.ac.uk (2015)
Hall, R. et al. The Geology of Mount Kinabalu (Sabah Parks, 2008)
Acknowledgements
We acknowledge the mountain guides of Sabah Parks for their help with material collection, ongoing assistance to continuing research in the area, and recent efforts following the 5 June 2015 earthquake. R. Butôt, M. de Jong, and B. Driessen carried out DNA extractions and PCRs. P. Wilkie provided a tissue sample for Payena. For the fungi, T. Semenova extracted DNA from the soil samples and E. Duijm generated the reads on the Ion Torrent sequencer. B. Blankevoort and E. J. Bosch prepared Fig. 1. J. Holloway (Natural History Museum, London, UK), I. Das (Universiti Malaysia Sarawak), and T. S. Liew (Naturalis, UMS) provided background on Kinabalu endemism, and R. Hall (University of London) on its geological history. R. Etienne (University of Groningen), R. Hall (University of London) and F. Bossuyt (Free University Brussels) read a draft of the paper. Collection and export permits were supplied by the Sabah Biodiversity Centre. Besides institutional support via Netherlands FES-funding, this project received funding via grants 821.01.002 and 825.12.007 of the Netherlands Organization for Scientific Research, the Alberta Mennega Foundation, the Ecology Fund of the Royal Netherlands Academy of Sciences, the Uyttenboogaart-Eliasen Foundation, and Pro Acarologia Basiliensis. G. van Uffelen, R. van Vugt, and P. Kessler (Botanic Gardens, Leiden) and T. de Winter, G. Thijsse, B. Hoeksema, H. Dautzenberg, J. Mols, and E. van Nieukerken (Naturalis) gave advice on transportation of specimens. Artist I. Kopelman and Naturalis general director E. van Huis (Naturalis) accompanied the expedition participants in the field. J. Gunsalam and P. Jilan (Sabah Parks Board of Trustees) officiated the opening and closing ceremony, respectively. P. Basintal, E. van Huis and E. Smets supervised the expedition as directors for the organizing institutions. Media coverage and communications were facilitated by J. van Alphen, A. Mohd. R. P. Dahlan, K. de Greef, A. Kromhout, J. Schilthuizen, E. E. Soudi, L. Stek, A. Taminiau, and M. Z. Abdul Wahab. Sabah Parks staff members L. Apin, B. Busin, B. Butit, J. Gampoyo, G. Gangko, B. Gangku, S. Gangku, P. Gimpiton, M. Gunggutou, T. Gunggutou, G. Gunsalam, J. Guntabid, I. Isnain, K. Ag. Jalil, Y. Johalin, R. M. Karim, Y. Kumin, D. Laban, A. Lahi, J. Lapidin, M. Latim, N. Maidin, N. Majuakim, R. Miadin, Y. Miki, M. Moguring, H. Mujih, F. Simin, K. C. Sion, D. Sumbin, D. Tingoh, E. Yakin, were invaluable for all manner of logistics and other support before, during, and after the expedition, in the field and at the stations and administrative offices. For the Forest Research Centre researchers, J. B. Dawat, M. Gumbilil, A. J. Jaimin, Jei. Jumian, Jem. Jumian, J. L. Yukang, P. J. Miun, M. A. T. Mustapha, and B. Saludin assisted in the field. T. Siew of Borneo Indah Sdn. Bhd. was the local supplier for consumables and field equipment. We thank the ESEB Outreach Fund and the Treub Foundation for funding outreach activities in Kinabalu Park and the Crocker Range Park. N. Raes helped with the niche modelling.
Author information
Authors and Affiliations
Contributions
M. Sch., V.S.F.T.M., M.Sp., M.M.B., C.B.M., L.E.B., K.T.C.A.P., J.N., R.R., and M.L. conceived and guided the study. M.Sch. and V.S.F.T.M. wrote the paper. K.H., F.S. and K.B. administrated all samples and sequences. K.H. and L.E.B. were responsible for DNA sampling in the field. V.S.F.T.M., A.A., N.A., A.B., P.-P.C., A.Y.C.C., R.D., H.F., C.F.-v.S., J.G., R.G., P.H., P.I., I.I., M.J., H.K., E.K., P.K., F.L., R.J.M., L.N.M., N.N., J.T.P., H.R., S.S., A.S., R.M.S., P.-S.S., H.S., N.S., J.B.S., M.Sul., S.Sum., J.v.T., F.Y.Y.T., B.E.Y., C.B.M., R.R. and M.Sch. collected material in the field and identified it following current taxonomy. K.H., V.S.F.T.M., H.d.B., F.A.A.F., J.G., B.G., P.H., S.B.J., S.N., M.St., D.C.T. and M.Sch. conducted the analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Phylogenetic reconstructions for Anura, Araneae, Athyriaceae, Balsaminaceae and Begoniaceae.
Details of materials and methods used are given in Supplementary Table 2.
Extended Data Figure 2 Phylogenetic reconstructions for Coleoptera, Dicranaceae, Diptera and Cortinariaceae.
Details of materials and methods used are given in Supplementary Table 2.
Extended Data Figure 3 Phylogenetic reconstructions for Ericaceae, Gastropoda, Glomeraceae, Hemiptera and Hirudinea.
Details of materials and methods used are given in Supplementary Table 2.
Extended Data Figure 4 Phylogenetic reconstructions for Nepenthaceae, Odonata, Polygalaceae, Rubiaceae and Trombidiformes.
Details of materials and methods used are given in Supplementary Table 2.
Extended Data Figure 5 Chronograms for Athyrium, Begonia, Coeliccia, Dacrycarpus, Epirixanthes, Dendrochilum, Dicranoloma, Impatiens, Hygrobates, Hedyotis, Ilex and Myrmarachne.
See Supplementary Table 5 for more details. Node colours represent posterior probabilities. Endemic species printed in red.
Extended Data Figure 6 Chronograms for Euphrasia, Kalophrynus, Nepenthes, Paphiopedilum, Philalanka, Ptomaphaginus, Payena, Trichosanthes, Rhododendron, Ranunculus, Tritetrabdella and Weinmannia.
See Supplementary Table 5 for more details. Node colours represent posterior probabilities. Endemic species printed in red.
Extended Data Figure 7 Ancestral range estimations for Athyrium, Begonia, Coeliccia, Dacrycarpus, Dendrochilum, Dicranoloma, Epirixanthes, Euphrasia and Hedyotis.
See Methods for further details. The trees are alphabetically ordered by genus name.
Extended Data Figure 8 Ancestral range estimations for Hygrobates, Ilex, Impatiens, Kalophrynus, Myrmarachne, Nepenthes, Paphiopedilum, Payena and Philalanka.
See Methods for further details. The trees are alphabetically ordered by genus name.
Extended Data Figure 9 Ancestral range estimations for Ptomaphaginus, Ranunculus, Rhododendron, Teleopsis, Trichosanthes, Tritetrabdella and Weinmannia.
See Methods for further details. The trees are alphabetically ordered by genus name.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1, 3, 4, 6 and 7 (see separate files for Supplementary Tables 2 and 5). (PDF 3236 kb)
Supplementary Tables
This file contains Supplementary Table 2. (PDF 591 kb)
Supplementary Tables
This file contains supplementary Table 5. (XLSX 57 kb)
Source data
Rights and permissions
About this article
Cite this article
Merckx, V., Hendriks, K., Beentjes, K. et al. Evolution of endemism on a young tropical mountain. Nature 524, 347–350 (2015). https://doi.org/10.1038/nature14949
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14949
This article is cited by
-
The Sino-Himalayan flora evolved from lowland biomes dominated by tropical floristic elements
BMC Biology (2023)
-
A rare non-canonical splice site in Trema orientalis SYMRK does not affect its dual symbiotic functioning in endomycorrhiza and rhizobium nodulation
BMC Plant Biology (2023)
-
Evolutionary history of the Arctic flora
Nature Communications (2023)
-
The formation of the Indo-Pacific montane avifauna
Nature Communications (2023)
-
Implementing spatial analyses to measure angiosperm biodiversity from the high-altitude grasslands of the Atlantic forest
Alpine Botany (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.