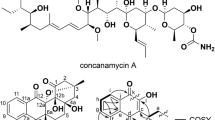
Abstract
Bacteria belonging to the order Actinomycetales produce most microbial metabolites thus far described, several of which have found applications in medicine and agriculture. However, most strains were discovered by their ability to produce a given molecule and are, therefore, poorly characterized physiologically and genetically. Thus, methodologies for genetic manipulation of actinomycetes are not available and efficient tools have been developed for just a few strains. This constitutes a serious limitation to applying molecular genetics approaches to strain development and structural manipulation of microbial metabolites. To overcome this hurdle, we have developed bacterial artificial chromosomes (BAC)1,2 that can be shuttled among Escherichia coli, where they replicate autonomously, and a suitable Streptomyces host, where they integrate site-specifically into the chromosome. The existence of gene clusters3 and of genetically amenable host strains, such as Streptomyces coelicolor or Streptomyces lividans4, makes this a sensible approach. We report here that 100 kb segments of actinomycete DNA can be cloned into these vectors and introduced into genetically accessible S. lividans, where they are stably maintained in integrated form in its chromosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Shizuya, H. et al. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc. Natl. Acad. Sci. USA 89, 8794–8797 (1992).
Ioannou, P.A. et al. A new bacteriophage P1-derived vector for the propagation of large human DNA fragments. Nat. Genet. 6, 84–89 (1994).
Martìn, J.F. & Liras, P. Organization and expression of genes involved in the biosynthesis of antibiotics and other secondary metabolites. Annu. Rev. Microbiol. 43, 173–206 (1989).
Hopwood, D.A. In: Proceedings from the Genetics and Molecular Biology and Industrial Microorganisms (GMBIM) conference (eds.Baltz, R.H., Hegeman, G.D. & Skatrud, P.L.) 1–6 (Society of Industrial Microbiology, Fairfax, VA; 1997).
Malpartida, F. & Hopwood, D.A. Molecular cloning of the whole biosynthetic pathway of a Streptomyces antibiotic and its expression in a heterologous host. Nature 309, 462–464 (1984).
Kao, C.M., Katz, L. & Khosla, C. Engineered biosynthesis of a complete macrolactone in a heterologous host. Science 265, 509–512 (1994).
Schwecke, T. et al. The biosynthetic gene cluster for the polyketide immunosuppressant rapamycin. Proc. Natl. Acad. Sci. USA 92, 7839–7843 (1995).
August P.R. et al. Biosynthesis of the ansamycin antibiotic rifamycin: deductions from the molecular analysis of the rif biosynthetic gene cluster of Amycolatopsis mediterranei S699. Chem. Biol. 5, 69–79 (1998).
Kuhstoss, S. & Rao, R.N. Analysis of the integration function of the streptomycete bacteriophage phi C31. J. Mol. Biol. 222, 897–908 (1991).
Hopwood, D.A. et al. Genetic manipulation of Streptomyces: a laboratory manual (The John Innes Foundation, Norwich, UK; 1985).
Ioannou, P.A. & de Jong, P.J. in Current protocols in human genetics (eds Dracopoli, N.C. et al.) 5.15.1–5.15.24 (Wiley, New York; 1996).
Kieser, H.M., Kieser, T. & Hopwood, D.A. A combined genetic and physical map of the chromosome of Streptomyces coelicolor A3(2). J. Bacteriol. 174, 5496–5507 (1992).
Redenbach, M. et al. A set of ordered cosmids and a detailed genetic and physical map of the 8Mb Streptomyces coelicolor A3(2) chromosome. Mol. Microbiol. 21, 77–96 (1996).
Leblond, P., Redenbach, M. & Cullum, J. Physical map of the Streptomyces lividans 66 genome and comparison with that of the related strain Streptomyces coelicolor A3(2). J. Bacteriol. 175, 3422–3429 (1993).
Xu, Y., Jiang, L., Murray, B.E. & Weinstock, G.M. Enterococcus faecalis antigens in human infections. Infect. Immun. 65, 4207–4215 (1997).
Brosch, R. et al. Use of a Mycobacterium tuberculosis H37Rv bacterial artificial chromosome library for genome mapping, sequencing, and comparative genomics. Infect. Immun. 66, 2221–2229 (1998).
Rondon, M.R., Raffel, S.J., Goodman, R.M. & Handelsman, J. Towards functional genomics in bacteria: analysis of expression in Escherichia coli from a bacterial artificial chromosome library of Bacillus cereus. Proc. Natl. Acad. Sci. USA 96, 6451–6455 (1999).
Hamilton, C.M., Frary, A., Lewis, C. & Tanskley, S.D. Stable transfer of intact high molecular weight DNA into plant chromosomes. Proc. Natl. Acad. Sci. USA 93, 9975–9979 (1996).
Liu, Y.-G. et al. Complementation of plant mutations with large genomic DNA fragments by a transformation-competent artificial chromosome vector accelerates positional cloning. Proc. Natl. Acad. Sci. USA 96, 6535–6540 (1999).
Birren, B. & Lai, E. Pulsed field gel electrophoresis: a practical guide (Academic Press, New York, NY; 1993).
Acknowledgements
We thank Luca Dolce and Rosa Alduina for performing some experiments on the ESAC library. We are grateful to Gianni Dehò and Charles Thompson for critical reading of the manuscript, and to David Hopwood and Pieter de Jong, and Sidney Kushner for plasmids and strains. This work was partially supported by the Italian CNR, PF Biotecnologie.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sosio, M., Giusino, F., Cappellano, C. et al. Artificial chromosomes for antibiotic-producing actinomycetes. Nat Biotechnol 18, 343–345 (2000). https://doi.org/10.1038/73810
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/73810
This article is cited by
-
Streptomyces: host for refactoring of diverse bioactive secondary metabolites
3 Biotech (2021)
-
Integrating vectors for genetic studies in the rare Actinomycete Amycolatopsis marina
BMC Biotechnology (2019)
-
Characterization and heterologous expression of the neoabyssomicin/abyssomicin biosynthetic gene cluster from Streptomyces koyangensis SCSIO 5802
Microbial Cell Factories (2018)
-
Characterization of the biosynthetic gene cluster (ata) for the A201A aminonucleoside antibiotic from Saccharothrix mutabilis subsp. capreolus
The Journal of Antibiotics (2017)