Abstract
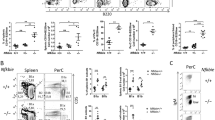
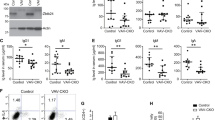
The gene FUS (also known as TLS (for translocated in liposarcoma) and hnRNP P2) is translocated with the gene encoding the transcription factor ERG-1 in human myeloid leukaemias1,2,3. Although the functions of wild-type FUS are unknown, the protein contains an RNA-recognition motif and is a component of nuclear riboprotein complexes4,5. FUS resembles a transcription factor in that it binds DNA, contributes a transcriptional activation domain to the FUS–ERG oncoprotein and interacts with several transcription factors in vitro6,7,8. To better understand FUS function in vivo, we examined the consequences of disrupting Fus in mice. Our results indicate that Fus is essential for viability of neonatal animals, influences lymphocyte development in a non-cell-intrinsic manner, has an intrinsic role in the proliferative responses of B cells to specific mitogenic stimuli and is required for the maintenance of genomic stability. The involvement of a nuclear riboprotein in these processes in vivo indicates that Fus is important in genome maintenance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pereira, D.S. et al. Retroviral transduction of TLS-ERG initiates a leukemogenic program in normal human hematopoietic cells. Proc. Natl Acad. Sci. USA 95, 8239–8244 ( 1998).
Ichikawa, H., Shimizu, K., Hayashi, Y. & Ohki, M. An RNA-binding protein gene, TLS /FUS, is fused to ERG in human myeloid leukemia with t(16;21) chromosomal translocation. Cancer Res. 54, 2865–2868 (1994).
Hiyoshi, M., Oh, K.R., Yamane, T. & Tatsumi, N. Acute non-lymphoblastic leukaemia with t(16;21): case report with a review of the literature. Clin. Lab Haematol. 17, 243–246 (1995).
Zinszner, H., Albalat, R. & Ron, D. A novel effector domain from the RNA-binding protein TLS or EWS is required for oncogenic transformation by CHOP. Genes Dev. 8, 2513–2526 ( 1994).
Calvio, C., Neubauer, G., Mann, M. & Lamond, A.I. Identification of hnRNP P2 as TLS/FUS using electrospray mass spectrometry. RNA 1, 724–733 ( 1995).
Prasad, D.D., Ouchida, M., Lee, L., Rao, V.N. & Reddy, E.S. TLS/FUS fusion domain of TLS/FUS-erg chimeric protein resulting from the t(16;21) chromosomal translocation in human myeloid leukemia functions as a transcriptional activation domain. Oncogene 9, 3717–3729 (1994).
Powers, C.A., Mathur, M., Raaka, B.M., Ron, D. & Samuels, H.H. TLS (translocated-in-liposarcoma) is a high-affinity interactor for steroid, thyroid hormone, and retinoid receptors. Mol. Endocrinol. 12, 4–18 (1998 ).
Hallier, M., Lerga, A., Barnache, S., Tavitian, A. & Moreau-Gachelin, F. The transcription factor Spi-1/PU.1 interacts with the potential splicing factor TLS. J. Biol. Chem. 273, 4838–4842 (1998).
Hicks, G.G. et al. Functional genomics in mice by tagged sequence mutagenesis . Nature Genet. 16, 338– 344 (1997).
Aman, P. et al. Expression patterns of the human sarcoma-associated genes FUS and EWS and the genomic structure of FUS. Genomics 37, 1–8 (1996).
Carsetti, R., Kohler, G. & Lamers, M.C. Transitional B cells are the target of negative selection in the B cell compartment. J. Exp. Med. 181, 2129–2140 (1995).
Horwitz, B.H., Scott, M.L., Cherry, S.R., Bronson, R.T. & Baltimore, D. Failure of lymphopoiesis after adoptive transfer of NF- κB-deficient fetal liver cells. Immunity 6, 765–772 (1997).
Hardin, J.D. et al. Bone marrow B lymphocyte development in c-abl-deficient mice . Cell. Immunol. 165, 44– 54 (1995).
Schwartzberg, P.L. et al. Mice homozygous for the ablm1 mutation show poor viability and depletion of selected B and T cell populations. Cell 65, 1165–1175 (1991).
Perrotti, D. et al. TLS/FUS, a pro-oncogene involved in multiple chromosomal translocations, is a novel regulator of BCR/ABL-mediated leukemogenesis. EMBO J. 17, 4442–4455 ( 1998).
Shafman, T. et al. Interaction between ATM protein and c-Abl in response to DNA damage. Nature 387, 520– 523 (1997).
Baskaran, R. et al. Ataxia telangiectasia mutant protein activates c-Abl tyrosine kinase in response to ionizing radiation. Nature 387 , 516–519 (1997).
Barlow, C. et al. Atm-deficient mice: a paradigm of ataxia telangiectasia. Cell 86, 159–171 ( 1996).
Xu, Y. et al. Targeted disruption of ATM leads to growth retardation, chromosomal fragmentation during meiosis, immune defects, and thymic lymphoma. Genes Dev. 10, 2411–2422 (1996).
Xu, Y. & Baltimore, D. Dual roles of ATM in the cellular response to radiation and in cell growth control. Genes Dev. 10, 2401–2410 (1996).
Bertrand, P., Akhmedov, A.T., Delacote, F., Durrbach, A. & Lopez, B.S. Human POMp75 is identified as the pro-oncoprotein TLS/FUS: both POMp75 and POMp100 DNA homologous pairing activities are associated to cell proliferation. Oncogene 18, 4515–4521 (1999).
Derr, L.K., Strathern, J.N. & Garfinkel, D.J. RNA-mediated recombination in S. cerevisiae. Cell 67, 355–364 ( 1991).
Maldonado, E. et al. A human RNA polymerase II complex associated with SRB and DNA-repair proteins. Nature 381, 86– 89 (1996).
Moore, J.K. & Haber, J.E. Capture of retrotransposon DNA at the sites of chromosomal double-strand breaks. Nature 383, 644–646 (1996).
Teng, S.C., Kim, B. & Gabriel, A. Retrotransposon reverse-transcriptase-mediated repair of chromosomal breaks. Nature 383, 641– 644 (1996).
Schreiber, E., Matthias, P., Muller, M.M. & Schaffner, W. Rapid detection of octamer binding proteins with mini-extracts, prepared from a small number of cells. Nucleic Acids Res. 17, 6419–6420 (1989).
Zinszner, H., Immanuel, D., Yin, Y., Liang, F.X. & Ron, D. A topogenic role for the oncogenic N-terminus of TLS: nucleolar localization when transcription is inhibited. Oncogene 14, 451–461 ( 1997).
Singh, N. et al. Cutting edge: activation of NK T cells by CD1d and α-galactosylceramide directs conventional T cells to the acquisition of a Th2 phenotype. J. Immunol. 163, 2373–2377 (1999).
Fukasawa, K., Wiener, F., Vande-Woude, G.F. & Mai, S. Genomic instability and apoptosis are frequent in p53 deficient young mice . Oncogene 15, 1295–1302 (1997).
Gaubatz, J.W. Extrachromosomal circular DNAs and genomic sequence plasticity in eukaryotic cells. Mutat. Res. 237, 271– 292 (1990).
Acknowledgements
We thank B. Collins for histology expertise; D. Houston for haematology expertise; D. McFarland for help with flow-cytometric analyses; K. Cann for assistance preparing the manuscript; and D. Ron for providing Fus antibody reagents and exchanging data before publication. This work was supported by Public Health Service Grants (R01HG00684 and R01RR13166 to H.E.R.), a grant from the Kleberg Foundation and Cancer Center (Core; P30CA42014) and a grant from Cancer Care Manitoba (CL2737 to G.G.H.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hicks, G., Singh, N., Nashabi, A. et al. Fus deficiency in mice results in defective B-lymphocyte development and activation, high levels of chromosomal instability and perinatal death . Nat Genet 24, 175–179 (2000). https://doi.org/10.1038/72842
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/72842
This article is cited by
-
Multi-omic and functional analysis for classification and treatment of sarcomas with FUS-TFCP2 or EWSR1-TFCP2 fusions
Nature Communications (2024)
-
The RNA-binding protein FUS/TLS interacts with SPO11 and PRDM9 and localize at meiotic recombination hotspots
Cellular and Molecular Life Sciences (2023)
-
Amyotrophic Lateral Sclerosis, FUS and Protein Synthesis Defects
Stem Cell Reviews and Reports (2023)
-
Modelling amyotrophic lateral sclerosis in rodents
Nature Reviews Neuroscience (2022)
-
Antisense oligonucleotide silencing of FUS expression as a therapeutic approach in amyotrophic lateral sclerosis
Nature Medicine (2022)