Abstract
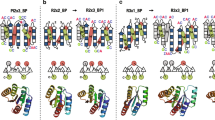
FOUR-RESIDUE β-turns and larger loop structures represent a significant fraction of globular protein surfaces and play an important role in determining the conformation and specificity of enzyme active sites and antibody-combining sites1,2. Turns are an attractive starting point to develop protein design methods, as they involve a small number of consecutive residues, adopt a limited number of defined conformations and are minimally constrained by packing interactions with the remainder of the protein. The ability to substitute oneβ-turn geometry for another will extend protein engineering beyond the redecoration of fixed backbone conformations to include local restructuring and the repositioning of surface side chains. To determine the feasibility and to examine the effect of such a structural modification on the fold and thermodynamic stability of a globular protein, we have substituted a five-residue turn sequence from concanavalin A for a type I' β-turn in staphylococcal nuclease. The resulting hybrid protein is folded and has full nuclease enzymatic activity but reduced thermodynamic stability. The crystal structure of the hybrid protein reveals that the guest turn sequence retains the conformation of the parent concanavalin A structure when substituted in the nuclease host.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Rose, G. D., Gierasch, L. M. & Smith, J. A. Adv. Prot. Chem. 37, 1–109 (1985).
Kuntz, I. D. J. Am. Chem. Soc. 94, 4,009–4,012 (1972).
Bajaj, M. & Blundell, T. A. A. Rev. Biophys. Bioengng 13, 453–492 (1984).
Smith, J. A. & Pease, L. G. Crit. Rev. Biochem. 8, 315–399 (1980).
Wilmot, C. N. & Thornton J. M. J. molec. Biol. 203, 221–232 (1988).
Kabsch, W. & Sander, C. Proc. natn. Acad. Sci. U.S.A. 81, 1,075–1,078 (1984).
Argos, P. J. molec. Biol. 197, 331–348 (1987).
Fine, R. M., Wang, H., Shenkin, P. S., Yarmush, D. L. & Levinthal, C. Proteins 1, 342–362 (1986).
Moult, J. & James, M. N. G. Proteins 4, 146–163 (1986).
Chothia, C. et al. Science 233, 755–758 (1986).
Reeke, G. N., Becker, J. W. & Edelman, G. M. J. biol. Chem. 250, 1525–1547 (1975).
Hardman, K. D. & Ainswirth, C. F. Biochemistry 11, 4910–45191 (1972).
Sibanda, B. L. & Thornton, J. M. Nature 316, 170–174 (1985).
Blundell, T. A., Sibanda, B. L., Sternberg, M. J. E. & Thornton, J. M. Nature 326, 347–352 (1987).
Cuatrecasus, P., Fuchs, S. & Anfinsen, C. B. J. biol. Chem. 242, 1541–1547 (1967).
Pace, C. N. Crit. Rev. Biochem. 3, 1–43 (1975).
Shortle, D. & Lin, B. Genetics 110, 539–555 (1985).
Shortle, D. & Meeker, A. K. Proteins 1, 81–89 (1986).
Burke, K. L., Dunn, G., Ferguson, M., Minor, P. D. & Almond, J. W. Nature 332, 81–82 (1988).
Murray, M. G. et al. Science 241, 213–215 (1988).
Small, D., Chou, P. Y. & Fasman, G. O. Biochem. biophys. Res. Commun. 79, 341–346 (1977).
Loll, P. J. & Lattman, E. E. Proteins (Submitted).
Arnone, A. et al. J. biol. Chem. 245, 2302–2316 (1971).
Cotton, F. A., Hazen, E. E. & Legg, M. J. Proc. natn. Acad. Sci. U.S.A. 76, 2,551–2,554 (1979).
Bernstein, F. C. et al. J. molec. Biol. 112, 535–542 (1977).
Lesk, A. M. & Hardman, K. D. Science 216, 539–540 (1982).
Evans, P. A., Kautz, R. A., Fox, R. O. & Dobson, C. M. Biochemistry 28, 362–370 (1989).
Arnone, A. et al. Proc. natn. Acad. Sci. U.S.A. 64, 420–427 (1969).
Xuong, N., Sullivan, D., Nielsen, C. & Hamlin, R. Acta Crystallogr. B41, 267–269 (1985).
Hendrickson, W. A. Meth. Enzym. 115, 252–271 (1985).
Jones, T. A. Meth. Enzym. 115, 157–171 (1985).
Mathews, B. W., & Czerwinski, E. W. Acta. Crystallogr. A31, 480–487 (1975).
Howard, A. J. et al. J. appl. Cryst. 20, 383–387 (1987).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hynes, T., Kautz, R., Goodman, M. et al. Transfer of a β-turn structure to a new protein context. Nature 339, 73–76 (1989). https://doi.org/10.1038/339073a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/339073a0
This article is cited by
-
Critical Role of a Loop at C-Terminal Domain on the Conformational Stability and Catalytic Efficiency of Chondroitinase ABC I
Molecular Biotechnology (2015)
-
Molecular Cloning and Evolutionary Analysis of Hemoglobin α-Chain Genes in Bats
Biochemical Genetics (2009)
-
In vitro evolution of thermodynamically stable turns
Nature Structural Biology (1996)
-
Mapping the structure of a non-native state of staphylococcal nuclease
Nature Structural Biology (1996)
-
The role of turns in the structure of an α-helical protein
Nature (1993)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.