Abstract
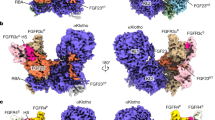
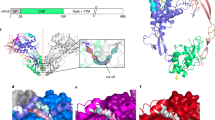
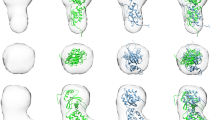
The fibroblast growth factors (FGFs) form a large family of structurally related, multifunctional proteins that regulate various biological responses1. They mediate cellular functions by binding to transmembrane FGF receptors2, which are protein tyrosine kinases. FGF receptors are activated by oligomerization3, and both this activation and FGF-stimulated biological responses require heparin-like molecules as well as FGF4. Heparins are linear anionic polysaccharide chains; they are typically heterogeneously sulphated on alternating L-iduronic and D-glucosamino sugars, and are nearly ubiquitous in animal tissues as heparan sulphate proteoglycans on cell surfaces and in the extracellular matrix. Although several crystal structures have been described for FGF molecules in complexes with heparin-like sugars5,6,7, the nature of a biologically active complex has been unknown until now. Here we describe the X-ray crystal structure, at 2.9 Å resolution, of a biologically active dimer of human acidic FGF in a complex with a fully sulphated, homogeneous heparin decassacharide. The dimerization of heparin-linked acidic FGF observed here is an elegant mechanism for the modulation of signalling through combinatorial homodimerization and heterodimerization of the 12 known members of the FGF family.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Basilico, C. & Moscatelli, D. The FGF family of growth factors and oncogenes. Adv. Cancer Res. 59, 115–165 (1992).
Jaye, M., Schlessinger, J. & Dionne, C. A. Fibroblast growth factor receptor tyrosine kinases: molecular analysis and signal transduction. Biochem. Biophys. Acta 1135, 185–199 (1992).
Schlessinger, J. Signal transduction by allosteric receptor oligomerization. Trends Biochem. Sci. 13, 443–447 (1988).
Spivak-Kroizman, T. et al. Heparin-induced oligomerization of FGF molecules is responsible for FGF receptor dimerization, activation, and cell proliferation. Cell 79, 1015–1024 (1994).
Zhu, X., Hsu, B. T. & Rees, D. C. Structural studies of the binding of the anti-ulcer drug sucrose octasulfate to acidic fibroblast growth factor. Structure 1, 27–34 (1993).
Ornitz, D. M. et al. FGF binding and FGF receptor activation by synthetic heparan-derived di- and trisaccharides. Science 268, 432–436 (1995).
Faham, S., Hileman, R. E., Fromm, J. R., Linhardt, R. J. & Rees, D. C. Heparin structure and interactions with basic fibroblast growth factor. Science 271, 1116–1120 (1996).
Mulloy, B., Forster, M. J., Jones, C. & Davies, D. B. NMR and molecular-modelling studies of the solution conformation of heparin. Biochem. J. 293, 849–858 (1993).
Burgess, W. H., Shaheen, A. M., Hampton, B., Donohue, P. J. & Winkles, J. A. Structure-function studies of heparin-binding (acidic fibroblast) growth factor-1 using site-directed mutagenesis. J. Cell Biol. 45, 131–138 (1991).
Thompson, L. D., Pantoliano, M. W. & Springer, B. A. Energetic characterization of the basic fibroblast growth factor-heparin interaction: identification of the heparin binding domain. Biochemistry 33, 3831–3840 (1994).
Ornitz, D. M., et al. Heparin is required for cell-free binding of basic fibroblast growth factor to a soluble receptor and for mitogenesis in whole cells. Mol. Cell. Biol. 12, 240–247 (1992).
Moy, F. J. et al. Properly oriented heparin-decasaccharide-induced dimers are the biologically active form of basic fibroblast growth factor. Biochemistry 36, 4782–4791 (1997).
Herr, A. B., Ornitz, D. M., Sasisekharan, R., Venkataraman, G. & Waksman, G. Heparin-induced self-association of fibroblast growth factor-2: evidence for two oligomerization processes. J. Biol. Chem. 272, 16382–16389 (1997).
Springer, B. A. et al. Identification and concerted function of two receptor binding surfaces on basic fibroblast growth factor required for mitogenesis. J. Biol. Chem. 269, 26879–26884 (1994).
Kan, M. et al. An essential heparin-binding domain in the fibroblast growth factor receptor kinase. Science 259, 1918–1921 (1993).
Weismann, C. et al. Crystal structure at 1.7 Å resolution of VEGF in complex with domain 2 of the Flt-1 receptor. Cell 91, 695–704 (1997).
van Holde, K. E. & Weischet, W. O. Boundary analysis of sedimentation-velocity experiments with monodisperse and paucidisperse solutes. Biopolymers 17, 1387–1403 (1978).
Huang, J., Mohammadi, M., Rodrigues, G. A. & Schlessingerr, J. Reduced activation of RAF-1 and MAP kinase by a fibroblast growth factor receptor mutant deficient in stimulation of phosphotidylinositol hydrolysis. J. Biol. Chem. 270, 5065–5072 (1995).
Jaye, M. et al. Human endothelial cell growth factor: cloning, nucleotide sequence, and chromosome localization. Science 233, 541–545 (1986).
Hendrickson, W. A., Horton, J. R. & LeMaster, D. M. Selenomethionyl proteins produced for analysis by multiwavelength anomalous diffraction (MAD): a vehicle for direct determination of three-dimensional structure. EMBO J. 9, 1665–1672 (1990).
Otwinowski, Z. DENZO 1-56-62(SERC Daresbury Laboratory, Warrington, UK, 1993).
Collaborative Computational Project no. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Hendrickson, W. A. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254, 51–58 (1991).
Brünger, A. T. X-PLOR Version 3.1, A System for X-ray Crystallography and NMR(Yale University Press, New Haven and London, 1992).
Jones, T. A., Zou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for binding protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Evans, S. V. SETOR: hardware lighted three-dimensional solid model representations of macromolecules. J. Mol. Graph. 11, 134–138 (1993).
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank C. Bingman, M. Cuff, L. Shapiro, H. Yamaguchi and C. Ogata for help in data collection; J. Arnez, K. Drickamer, B. Honig, S. Hubbard, J. Passner, M. Quesenberry, L. Shapiro, and H. Yamaguchi for discussions; P. Östergaard (Novo Nordisk) for heparin oligosaccharides; M. Gawinowicz (HHMI Columbia) for amino-acid analysis; D. King for electrospray mass spectrometry; and C. Turgeon and J. Hansen for analytical ultracentrifugation. This work was supported in part by a grant from the NIH. Beamline X4A at the National Synchrotron Light Source, a DOE faciity, is supported by the HHMI.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
DiGabriele, A., Lax, I., Chen, D. et al. Structure of a heparin-linked biologically active dimer of fibroblast growth factor. Nature 393, 812–817 (1998). https://doi.org/10.1038/31741
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/31741
This article is cited by
-
Binding affinity estimation from restrained umbrella sampling simulations
Nature Computational Science (2022)
-
Bioactive polysaccharides from natural resources including Chinese medicinal herbs on tissue repair
Chinese Medicine (2018)
-
Basic fibroblast growth factor released from fucoidan-modified chitosan/alginate scaffolds for promoting fibroblasts migration
Journal of Polymer Research (2018)
-
Heparin: role in protein purification and substitution with animal-component free material
Applied Microbiology and Biotechnology (2018)
-
An integrated approach using orthogonal analytical techniques to characterize heparan sulfate structure
Glycoconjugate Journal (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.