Abstract
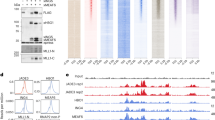
HIRA maps to the DiGeorge/velocardiofacial syndrome critical region (DGCR) at 22q11 (Refs 1,2) and encodes a WD40 repeat protein similar to yeast Hir1p and Hir2p. These transcriptional co-repressors regulate cell cycle-dependent histone gene transcription3, possibly by remodelling local chromatin structure. We report an interaction between HIRA and the transcription factor Pax3. Pax3 haploinsufficiency results in the mouse splotch and human Waardenburg syndrome (WSI and WSIII) phenotypes. Mice homozygous for Pax3 mutations die in utero with a phenocopy of DGS, or neonatally with neural tube defects. HIRA was also found to interact with core histones. Thus, altered stoichiometry of complexes containing HIRA may be important for the development of structures affected in WS and DGS.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Halford, S. et al. Isolation of a putative transcriptional regulator from the region of 22q11 deleted in DiGeorge Syndrome, Shprintzen syndrome and familial congenital heart disease. Hum. Mol. Genet. 2, 2099–2107 (1993).
Lamour, V. et al. A human homolog of the S. cerevisiae HIR1 and HIR2 transcriptional repressors cloned from the DiGeorge syndrome critical region. Hum. Mol. Genet. 4, 791–799 (1995).
Spector, M.S., Raff, A., DeSilva, H., Lee, K. & Osley, M.A. Hir1p and Hir2p function as transcriptional corepressors to regulate histone gene transcription in the Saccharomyces cerevisiae cell cycle. Mol. Cell. Biol. 17, 545 –552 (1997).
Dallapiccola, B., Pizzuti, A. & Novelli, G. How many breaks do we need to CATCH on 22q11? Am. J. Hum. Genet. 59, 7–11 (1996).
Kurahashi, H. et al. Deletion mapping of 22q11 in CATCH22 syndrome: identification of a second critical region. Am. J. Hum. Genet. 58, 1377–1881 (1996).
Rizzu, P. et al. Cloning and comparative mapping of a gene deleted in DiGeorge and velocardiofacial syndromes conserved in the C.elegans genome. Mamm. Genome 7, 639–643 (1996).
Carlson, C. et al. Molecular definition of 22q11 deletions in 151 VCFS patients: integration within a 10kb physical map. Am. J. Hum. Genet. 61, 620–629 (1997).
Budarf, M.L. & Emanuel, B.S. Progress in the autosomal segmental aneusomy syndromes (SASs): single or multi-locus disorders? Hum. Mol. Genet. 6, 1657–1665 (1997).
Kirby, M.L. & Waldo, K.L. Role of the neural crest in congenital heart disease. Circulation 82, 332– 340 (1990).
Manley, N. & Capecchi, M.R. The role of Hoxa-3 in mouse thymus and thyroid development. Development 121, 1989–2003 (1995).
Roberts, C., Daw, S., Halford, S. & Scambler, P.J. Cloning and Developmental Expression Analysis of Chick Hira, a candidate gene for DiGeorge syndrome. Hum. Mol. Genet. 6, 237– 246 (1997).
Wilming, L.G., Snoeren, C.A.S., van Rijswijk, A., Grosveld, F. & Meijers, C. The murine homologue of HIRA, a DiGeorge syndrome candidate gene, is expressed in embryonic structures affected in CATCH22 patients. Hum. Mol. Genet. 6, 247–258 (1997).
Lorain, S. et al. Core histones and HIRIP3, a novel histone-binding protein, directly interact with the WD repeat protein HIRA. Mol. Cell. Biol. in press (1998).
Goulding, M.D., Chalepakis, G., Deutsch, E., Erselius, J.R. & Gruss, P. Pax-3, a novel murine DNA binding protein expressed during early embryogenesis. EMBO J. 10, 1135–1147 (1991).
Zappavigna, V., Falciola, L., Citterich, M.H., Mavilio, F. & Bianchi, M.E. HMG1 interacts with Hox proteins and enhances their DNA binding and transcriptional activation. EMBO J. 15, 5981–4991 ( 1996).
Jostes, B., Walther, C. & Gruss, P. The murine paired box gene, Pax7, is expressed specifically during the development of the nervous and muscular system. Mech. Dev. 33, 27–38 ( 1991).
Mansouri, A., Stoykova, A., Torres, M. & Gruss, P. Dysgenesis of cephalic neural crest derivatives in Pax7-/- mutant mice. Development 122, 831–838 (1996).
Conway, S.J., Henderson, D.J. & Copp, A.J. Pax3 is required for cardiac neural crest migration in the mouse: evidence from the splotch (Sp2H) mutant. Development 124, 505–514 ( 1997).
Read, A.P. & Newton, V.E. Waardenburg syndrome. J. Med. Genet. 34, 656–665 (1997).
Franz, T. Persistent truncus arteriosus in the Splotch mutant mouse. Anat. Embryol. 180, 457–464 ( 1989).
Zlotogora, J., Lerer, I., Bar-David, S., Ergaz, Z. & Abeliovich, D. Homozygosity for Waardenburg syndrome. Am. J. Hum. Genet. 56, 1173–1178 (1995).
Steel, K.P. & Smith, R.J. Normal hearing in Splotch (Sp/+), the mouse homologue of Waardenburg syndrome type I. Nature Genet. 2, 75–79 (1992 ).
Daw, S.C.M. et al. A common region of 10p deleted in DiGeorge and velo-cardio-facial syndrome. Nature Genet. 13, 458– 460 (1996).
Edmundson, D.G., Smith, M.M. & Roth, S.Y. Repression domain of the yeast global repressor Tup1 interacts directly with histones H3 and H4. Genes Dev. 10, 1247–1259 (1996).
Gould, A. Functions of mammalian Polycomb group and trithorax group related genes. Curr. Opin. Genet. Dev. 7, 488– 494 (1997).
Hollenberg, S.M., Sternglanz, R., Cheng, P.F. & Weintraub, H. Identification of a new family of tissue-specific basic helix-loop-helix proteins with a two-hybrid system. Mol. Cell. Biol. 15, 3813–3822 (1995).
Bours, V. et al. The oncoprotein BCL-3 directly transactivates through kappaB motifs via association with DNA-binding p50b homodimers. Cell 72, 729–739 (1993).
Harlow, E. & Lane, D. Antibodies: A Laboratory Manual. (Cold Spring Harbor Laboratory, New York, 1988).
Moase, C.E. & Trasler, D.G. Delayed neural crest emigration from Sp and Spd mouse neural tube explants. Teratology 42, 171–182 ( 1990).
Chalepakis, G., Jones, F.S., Edelman, G.M. & Gruss, P. Pax3 contains domains for transcription activation and transcription inhibition. Proc. Natl Acad. Sci. USA 91, 12745– 12749 (1994).
Acknowledgements
We would like to thank A. Copp and D. Henderson for Pax3 antibodies, the Pax3-pClneo construct and their generous help. S. Hollenberg supplied two-hybrid reagents, protocols and advice. C. Meijers kindly provided anti-Hira antibody. This work was supported by the British Heart Foundation, the Birth Defects Foundation and MRC (UK), the Ligue Nationale contre le Cancer, AFM, ARC (France) and NATO and the Human Frontier Science Program (UK and France).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Magnaghi, P., Roberts, C., Lorain, S. et al. HIRA, a mammalian homologue of Saccharomyces cerevisiae transcriptional co-repressors, interacts with Pax3. Nat Genet 20, 74–77 (1998). https://doi.org/10.1038/1739
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/1739
This article is cited by
-
Description of novel variants in consanguineous Pakistani families affected with intellectual disability
Genes & Genomics (2023)
-
In the line-up: deleted genes associated with DiGeorge/22q11.2 deletion syndrome: are they all suspects?
Journal of Neurodevelopmental Disorders (2019)
-
Synergy of Hir1, Ssn6, and Snf2 global regulators is the functional determinant of a Mac1 transcriptional switch in S. cerevisiae copper homeostasis
Current Genetics (2019)
-
UBN1/2 of HIRA complex is responsible for recognition and deposition of H3.3 at cis-regulatory elements of genes in mouse ES cells
BMC Biology (2018)
-
Genetic and physical interaction of Meis2, Pax3 and Pax7 during dorsal midbrain development
BMC Developmental Biology (2012)