Abstract
The opening and closing of two ring-shaped Mcm2–7 DNA helicases is necessary to license eukaryotic origins of replication, although the mechanisms controlling these events are unclear. The origin-recognition complex (ORC), Cdc6 and Cdt1 facilitate this process by establishing a topological link between each Mcm2–7 hexamer and origin DNA. Using colocalization single-molecule spectroscopy and single-molecule Förster resonance energy transfer (FRET), we monitored ring opening and closing of Saccharomyces cerevisiae Mcm2–7 during origin licensing. The two Mcm2–7 rings were open during initial DNA association and closed sequentially, concomitant with the release of their associated Cdt1. We observed that ATP hydrolysis by Mcm2–7 was coupled to ring closure and Cdt1 release, and failure to load the first Mcm2–7 prevented recruitment of the second Mcm2–7. Our findings identify key mechanisms controlling the Mcm2–7 DNA-entry gate during origin licensing, and reveal that the two Mcm2–7 complexes are loaded via a coordinated series of events with implications for bidirectional replication initiation and quality control.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Yardimci, H. & Walter, J.C. Prereplication-complex formation: a molecular double take? Nat. Struct. Mol. Biol. 21, 20–25 (2014).
Samel, S.A. et al. A unique DNA entry gate serves for regulated loading of the eukaryotic replicative helicase MCM2-7 onto DNA. Genes Dev. 28, 1653–1666 (2014).
Bochman, M.L. & Schwacha, A. The Mcm2-7 complex has in vitro helicase activity. Mol. Cell 31, 287–293 (2008).
Remus, D. et al. Concerted loading of Mcm2-7 double hexamers around DNA during DNA replication origin licensing. Cell 139, 719–730 (2009).
Aparicio, O.M., Weinstein, D.M. & Bell, S.P. Components and dynamics of DNA replication complexes in S. cerevisiae: redistribution of MCM proteins and Cdc45p during S phase. Cell 91, 59–69 (1997).
Fernández-Cid, A. et al. An ORC/Cdc6/MCM2-7 complex is formed in a multistep reaction to serve as a platform for MCM double-hexamer assembly. Mol. Cell 50, 577–588 (2013).
Sun, J. et al. Cryo-EM structure of a helicase loading intermediate containing ORC–Cdc6–Cdt1–MCM2–7 bound to DNA. Nat. Struct. Mol. Biol. 20, 944–951 (2013).
Ticau, S., Friedman, L.J., Ivica, N.A., Gelles, J. & Bell, S.P. Single-molecule studies of origin licensing reveal mechanisms ensuring bidirectional helicase loading. Cell 161, 513–525 (2015).
Sun, J. et al. Structural and mechanistic insights into Mcm2–7 double-hexamer assembly and function. Genes Dev. 28, 2291–2303 (2014).
Evrin, C. et al. A double-hexameric MCM2-7 complex is loaded onto origin DNA during licensing of eukaryotic DNA replication. Proc. Natl. Acad. Sci. USA 106, 20240–20245 (2009).
Bell, S.P. & Stillman, B. ATP-dependent recognition of eukaryotic origins of DNA replication by a multiprotein complex. Nature 357, 128–134 (1992).
Randell, J.C.W., Bowers, J.L., Rodríguez, H.K. & Bell, S.P. Sequential ATP hydrolysis by Cdc6 and ORC directs loading of the Mcm2-7 helicase. Mol. Cell 21, 29–39 (2006).
Kang, S., Warner, M.D. & Bell, S.P. Multiple functions for Mcm2-7 ATPase motifs during replication initiation. Mol. Cell 55, 655–665 (2014).
Coster, G., Frigola, J., Beuron, F., Morris, E.P. & Diffley, J.F.X. Origin licensing requires ATP binding and hydrolysis by the MCM replicative helicase. Mol. Cell 55, 666–677 (2014).
Bowers, J.L., Randell, J.C.W., Chen, S. & Bell, S.P. ATP hydrolysis by ORC catalyzes reiterative Mcm2-7 assembly at a defined origin of replication. Mol. Cell 16, 967–978 (2004).
Chang, F. et al. Cdc6 ATPase activity disengages Cdc6 from the pre-replicative complex to promote DNA replication. eLife 4, 1–42 (2015).
Frigola, J., Remus, D., Mehanna, A. & Diffley, J.F.X. ATPase-dependent quality control of DNA replication origin licensing. Nature 495, 339–343 (2013).
Deegan, T.D. & Diffley, J.F. MCM: one ring to rule them all. Curr. Opin. Struct. Biol. 37, 145–151 (2016).
Costa, A. et al. The structural basis for MCM2-7 helicase activation by GINS and Cdc45. Nat. Struct. Mol. Biol. 18, 471–477 (2011).
Lyubimov, A.Y., Costa, A., Bleichert, F., Botchan, M.R. & Berger, J.M. ATP-dependent conformational dynamics underlie the functional asymmetry of the replicative helicase from a minimalist eukaryote. Proc. Natl. Acad. Sci. USA 109, 11999–12004 (2012).
Li, N. et al. Structure of the eukaryotic MCM complex at 3.8 Å. Nature 524, 186–191 (2015).
Schepers, A. & Diffley, J.F. Mutational analysis of conserved sequence motifs in the budding yeast Cdc6 protein. J. Mol. Biol. 308, 597–608 (2001).
Donovan, S., Harwood, J., Drury, L.S. & Diffley, J.F. Cdc6p-dependent loading of Mcm proteins onto pre-replicative chromatin in budding yeast. Proc. Natl. Acad. Sci. USA 94, 5611–5616 (1997).
Zhai, Y. et al. Open-ringed structure of the Cdt1–MCM2–7 complex as a precursor of the MCM double hexamer. Nat. Struct. Mol. Biol. http://dx.doi.org/10.1038/nsmb.3374 (2016).
Samson, R.Y., Abeyrathne, P.D. & Bell, S.D. Mechanism of Archaeal MCM helicase recruitment to DNA replication origins. Mol. Cell 61, 287–296 (2016).
Kelch, B.A. The lord of the rings: structure and mechanism of the sliding clamp loader. Biopolymers 105, 532–546 (2016).
Yeeles, J.T.P., Deegan, T.D., Janska, A., Early, A. & Diffley, J.F.X. Regulated eukaryotic DNA replication origin firing with purified proteins. Nature 519, 431–435 (2015).
Chen, I., Dorr, B.M. & Liu, D.R. A general strategy for the evolution of bond-forming enzymes using yeast display. Proc. Natl. Acad. Sci. USA 108, 11399–11404 (2011).
Kindermann, M., Sielaff, I. & Johnsson, K. Synthesis and characterization of bifunctional probes for the specific labeling of fusion proteins. Bioorg. Med. Chem. Lett. 14, 2725–2728 (2004).
Smith, B.A. et al. Three-color single molecule imaging shows WASP detachment from Arp2/3 complex triggers actin filament branch formation. eLife 2, e01008 (2013).
Friedman, L.J. & Gelles, J. Mechanism of transcription initiation at an activator-dependent promoter defined by single-molecule observation. Cell 148, 679–689 (2012).
Friedman, L.J., Chung, J. & Gelles, J. Viewing dynamic assembly of molecular complexes by multi-wavelength single-molecule fluorescence. Biophys. J. 91, 1023–1031 (2006).
Crawford, D.J., Hoskins, A.A., Friedman, L.J., Gelles, J. & Moore, M.J. Single-molecule colocalization FRET evidence that spliceosome activation precedes stable approach of 5′ splice site and branch site. Proc. Natl. Acad. Sci. USA 110, 6783–6788 (2013).
Hoskins, A.A. et al. Ordered and dynamic assembly of single spliceosomes. Science 331, 1289–1295 (2011).
Friedman, L.J. & Gelles, J. Multi-wavelength single-molecule fluorescence analysis of transcription mechanisms. Methods 86, 27–36 (2015).
Acknowledgements
We are grateful to the members of the Bell laboratory for useful discussions, B. Sauer for comments on the manuscript, and L.D. Lavis (Janelia Research Campus, Ashburn, Virginia, USA) for providing Janelia fluorophores. This work was supported by the NIH (grants R01 GM52339 (to S.P.B.) and R01 GM81648 (to J.G.); pre-doctoral training grant GM007287 to S.T.) and the G. Harold and Leila Y. Matthews Foundation (grant to J.G.). S.P.B. is an investigator with the Howard Hughes Medical Institute. This work was supported in part by the Koch Institute (support grant P30-CA14051 from the NCI to T. Jacks, Koch Institute for Integrative Cancer Research, MIT, Cambridge, Massachusetts, USA). We thank the Koch Institute Swanson Biotechnology Center for technical support, specifically the Biopolymers and Genomics cores.
Author information
Authors and Affiliations
Contributions
S.T. performed all experiments with feedback from J.G., L.J.F. and S.P.B., except for the ensemble FRET studies, which were performed by K.C. I.R.C. prepared essential reagents. S.T., L.J.F. and J.G. analyzed the data. S.P.B. wrote the paper with input from all other authors. S.P.B. and J.G. directed the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Additional characterization of Mcm2–725FRET and Mcm2–725quench.
(A) Illustration of the approximate locations of SNAP and CLIP protein (1EH8) insertions in Mcm2–7 hexamer (3JA8; ref 21) used for labeling Mcm2–725FRET. SNAP and CLIP were inserted after amino acid 591 in Mcm5 and 721 in Mcm2, respectively. The distance between the two insertion sites is approximately 64 Å.
(B) Purified helicase-loading proteins. The indicated purified protein preparations were separated by SDS-PAGE and stained with Coomassie. Note that the insertion of SNAP or CLIP into Mcm5 and Mcm2 respectively caused them to comigrate with other Mcm subunits. The gray square covers other preparations not relevant to this study.
(C) Helicase-loading activity of Mcm2–725FRET and Mcm2–725quench. Bulk helicase-loading assays were performed using unlabeled, fluorescently-labeled Cdt1–Mcm2–725FRET, or fluorophore- and quencher-labeled Cdt1–Mcm2–725quench. All assays were performed with purified Cdc6, ORC, and origin DNA. All samples were subjected to a high-salt wash and DNA associated proteins were separated by SDS-PAGE and detected with Krypton protein stain (Thermo Fisher).
(D) Relative helicase-loading activity of Mcm2–725FRET and Mcm2–725quench relative to unlabeled Mcm2–7. Duplicate helicase-loading assays were performed as described in (C) with the indicated Mcm2–7 complexes. DNA-associated Mcm2–7 was quantified using AlphaView software (ProteinSimple).
Supplementary Figure 2 FRET is observed only when donor and acceptor fluorophores are on the same Mcm2–7.
Records of fluorescence co-localized with individual origin DNA molecules for Mcm2-725FRET complexes in which only Mcm2 or Mcm5 in a single hexamer was labeled (due to incomplete labeling). Acceptor-excited (i, A ex, A em) and donor-excited (ii, D ex, D em and D ex, A em) emission records are shown together with calculated donor-excited total emission (iii) and EFRET (iv). EFRET is only shown when D and A fluorophores are present (left example); in this case the first Mcm2–7 is labeled with D fluorophore only and the second Mcm2–7 is labeled with A fluorophore only and no EFRET increase after binding is observed.
(A) The first Mcm2–7 is only labeled with the acceptor fluorophore and the second Mcm2–7 is only labeled with the donor fluorophore.
(B) The DNA-associated Mcm2–7 is labeled with the acceptor fluorophore only.
(C) The DNA-associated Mcm2–7 is labeled with the donor fluorophore only.
Supplementary Figure 3 Mcm2–7 hexamers associate with DNA in an open-gate conformation and subsequently close.
(A) Additional representative records of fluorescence co-localized with individual origin DNA molecules for experiments using Mcm2–725FRET in which both D and A were present on both hexamers. Panels i-iv, black arrows and black bars are as described in Fig. 1A. Blue, green and red arrows indicate short-lived second Mcm2–7 associations; blue, Mcm2–7 labeled with D and A fluorophores; green, Mcm2–7 labeled with D fluorophore only (due to incomplete labeling); red, Mcm2–7 labeled with A fluorophore only.
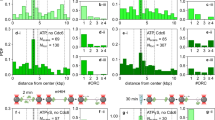
(B) Left: Record of mean EFRET values after initial Mcm2–725FRET DNA association (N= 57 molecules, ± s.e.m.). Only first Mcm2–7 associations where both D and A fluorophores were present for >100s and for which no second Mcm2–7 association was detected within 100s were selected for this analysis. Right: Evolution of the EFRET distribution for the same 57 complexes. The plot is a two-dimensional Gaussian kernel histogram with bandwidths 5.4 s and 0.05 on the time and EFRET axes, respectively (https://github.com/gelles-brandeis/jganalyze). A total of 6197 time points (≤ 500s) were plotted. For both plots, arrival of the first Mcm2–725FRET has been set to time zero.
(C) Histograms of EFRET values recorded in the indicated time intervals after association of the second Mcm2–725FRET with origin DNA already bound to a first Mcm2–725FRET. Time windows are as in Fig. 1C. Fits to the sum (cyan) of three Gaussians (red) yielded fit parameters shown in Supplementary Table 2. For accurate EFRET determination, only the 21 second Mcm2–7 associations lasting >150 s where both D and A fluorophores were present for both the first and the second Mcm2–7 molecules were analyzed. Associations where an additional Mcm2–7 molecule associated within 150s of the second Mcm2–7 association were eliminated from this analysis. The same 21 molecules were used for each histogram with data from 132 (0-15s), 484 (15-75s) and 574 (75-150s) total time points plotted.
(D) Left: Record of mean EFRET values after initial Mcm2–72C5FRET DNA association (N = 71 molecules, ± s.e.m.). Mcm2–72C5FRET has the same site of Mcm5 fluorophore attachment but the site of Mcm2 fluorophore attachments is now at the C-terminus. Molecules were selected for analysis by the same criteria as in (B). Right: Evolution of the EFRET distribution for the same set of molecules. The plot is a two-dimensional Gaussian kernel histogram (https://github.com/gelles-brandeis/jganalyze) with bandwidths 2.7 s and 0.05 on the time and EFRET axes, respectively. For both plots, arrival of the first Mcm2–725FRET has been set to time zero.
Supplementary Figure 4 Kinetic analysis of helicase loading with Mcm2-Mcm5 gate–modified Mcm2–7 complexes.
(A) Comparison of Cdc6 release times after loading of the first Mcm2–725FRET* or Mcm2–74SNAP. Fraction of fluorophore-labeled Cdc6 (Cdc6SORT549) released after the first Mcm2–725FRET* (red, N = 56) or first Mcm2–74SNAP (the singly-tagged Mcm2–7 used in ref. 8, gray, N = 82) are plotted. Red lines represent the 95% confidence interval (CI) for the release of Cdc6 associated with Mcm2–725FRET*.
(B) Comparison of Cdc6SORT549 release times after loading of the second Mcm2–725FRET* or Mcm2–74SNAP. Fraction of Cdc6 released after the arrival of the second Mcm2–725FRET* (blue, N = 27 molecules) or the second Mcm2–74SNAP (from ref. 8, gray, N = 70 molecules) are plotted. Blue lines represent the 95% CI for the release of Cdc6 associated with the second Mcm2–725FRET*.
(C) Comparison of ORC release times after loading of the second Mcm2–725FRET* or second Mcm2–74SNAP. Fraction of fluorophore-labeled ORC (ORC1SORT549) released after arrival of the second Mcm2–725FRET* (red, N = 25 molecules) or the second Mcm2–74SNAP (from ref. 8, black, N = 98) are plotted. Red lines represent the 95% CI for the ORC release from Mcm2–725FRET*.
(D) Additional analysis of Mcm2–725quench. Record of population average Equench values after initial Mcm2–725quench DNA association (N = 61 molecules, ± s.e.m.). Only first Mcm2–7 associations where the D fluorophore was present for >100s and for which no second Mcm2–7 association was detected within 100s were analyzed. Equench was calculated as a fraction of the fluorescence intensity relative to the first time point of Mcm2–725quench association.
(E) Relative timing of second Mcm2–7 gate closing and initial double-hexamer interactions. At the indicated times the fraction of DNA molecules associated with two Mcm2–7 complexes that show double-hexamer interactions (green; N = 106 molecules, data taken from ref. 8, measured by EFRET between the first and second Mcm2–7) or the fraction of second Mcm2–7 with a closed gate (blue; N = 47 molecules) are plotted. Note that the vertical axis is inverted and that zero on the horizontal axis corresponds to the arrival of the second Mcm2–7. Inset shows mean times after second Mcm2–7 arrival (± 95% CI). Records for gate closure were selected as described for Fig. 2A.
Supplementary Figure 5 Further analysis of Mcm2–725FRET-5RA.
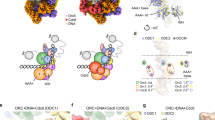
(A) Diagram illustrating the ATPase sites formed at the interface between each pair of Mcm2–7 subunits and the location of the Mcm5RA mutation relative to the Mcm2-Mcm5 gate (view is from the C-terminal end of Mcm2–7). The extension from each Mcm subunit represents the arginine finger required for ATP hydrolysis. The cut out of each subunit represents the primary ATP binding site including the Walker A and B motifs.
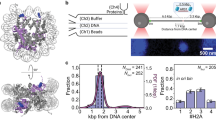
(B) Evolution of the EFRET distribution for Mcm2–725FRET-5RA associations where both D and A fluorophores were present (N = 33 molecules). The plot is a two-dimensional Gaussian kernel histogram with bandwidths 5.4 s and 0.05 on the time and EFRET axes, respectively. The initial binding of Mcm2–725FRET-5RA is set to time zero.
(C) Two example recordings of labeled Mcm2–74SNAP-mcm5RA/Cdt1SORT549 complexes associating with an individual origin DNA molecule. Both labeled proteins associate simultaneously with the DNA; in one example (left) they persist until the end of the recording (N = 57/109 molecules); in the other they dissociate simultaneously within experimental uncertainty (N = 40/109 molecules).
Supplementary Figure 6 Structures of SNAP-BHQ2 and SNAP-549-biotin.
Molecular structures of (A) SNAP-BHQ2 and (B) SNAP-549-Biotin whose synthesis is described in the Online Methods.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–3 (PDF 1538 kb)
Rights and permissions
About this article
Cite this article
Ticau, S., Friedman, L., Champasa, K. et al. Mechanism and timing of Mcm2–7 ring closure during DNA replication origin licensing. Nat Struct Mol Biol 24, 309–315 (2017). https://doi.org/10.1038/nsmb.3375
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3375
This article is cited by
-
A mechanism of origin licensing control through autoinhibition of S. cerevisiae ORC·DNA·Cdc6
Nature Communications (2022)
-
The structure of ORC–Cdc6 on an origin DNA reveals the mechanism of ORC activation by the replication initiator Cdc6
Nature Communications (2021)
-
DNA replication origins retain mobile licensing proteins
Nature Communications (2021)
-
Mechanism of head-to-head MCM double-hexamer formation revealed by cryo-EM
Nature (2019)
-
Abp1 promotes Arp2/3 complex-dependent actin nucleation and stabilizes branch junctions by antagonizing GMF
Nature Communications (2018)