Abstract
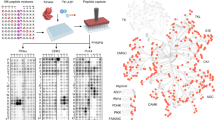
Protein phosphorylation is estimated to affect 30% of the proteome and is a major regulatory mechanism that controls many basic cellular processes1,2,3. Until recently, our biochemical understanding of protein phosphorylation on a global scale has been extremely limited; only one half of the yeast kinases have known in vivo substrates and the phosphorylating kinase is known for less than 160 phosphoproteins. Here we describe, with the use of proteome chip technology4, the in vitro substrates recognized by most yeast protein kinases5: we identified over 4,000 phosphorylation events involving 1,325 different proteins. These substrates represent a broad spectrum of different biochemical functions and cellular roles. Distinct sets of substrates were recognized by each protein kinase, including closely related kinases of the protein kinase A family and four cyclin-dependent kinases that vary only in their cyclin subunits. Although many substrates reside in the same cellular compartment or belong to the same functional category as their phosphorylating kinase, many others do not, indicating possible new roles for several kinases. Furthermore, integration of the phosphorylation results with protein–protein interaction6,7,8,9,10 and transcription factor binding data11,12 revealed novel regulatory modules. Our phosphorylation results have been assembled into a first-generation phosphorylation map for yeast. Because many yeast proteins and pathways are conserved, these results will provide insights into the mechanisms and roles of protein phosphorylation in many eukaryotes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Manning, G., Plowman, G. D., Hunter, T. & Sudarsanam, S. Evolution of protein kinase signalling from yeast to man. Trends Biochem. Sci. 27, 514–520 (2002)
Ficarro, S. B. et al. Phosphoproteome analysis by mass spectrometry and its application to Saccharomyces cerevisiae. Nature Biotechnol. 20, 301–305 (2002)
Cohen, P. The regulation of protein function by multisite phosphorylation—a 25 year update. Trends Biochem. Sci. 25, 596–601 (2000)
Zhu, H. et al. Global analysis of protein activities using proteome chips. Science 293, 2101–2105 (2001)
Zhu, H. et al. Analysis of yeast protein kinases using protein chips. Nature Genet. 26, 283–289 (2000)
Gavin, A. C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002)
Ito, T. et al. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc. Natl Acad. Sci. USA 98, 4569–4574 (2001)
Uetz, P. et al. A comprehensive analysis of protein–protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000)
Xenarios, I. et al. DIP: the database of interacting proteins. Nucleic Acids Res. 28, 289–291 (2000)
Bader, G. D. & Hogue, C. W. BIND—a data specification for storing and describing biomolecular interactions, molecular complexes and pathways. Bioinformatics 16, 465–477 (2000)
Horak, C. E. et al. Complex transcriptional circuitry at the G1/S transition in Saccharomyces cerevisiae. Genes Dev. 16, 3017–3033 (2002)
Lee, T. I. et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298, 799–804 (2002)
Toda, T., Cameron, S., Sass, P., Zoller, M. & Wigler, M. Three different genes in S. cerevisiae encode the catalytic subunits of the cAMP-dependent protein kinase. Cell 50, 277–287 (1987)
Pan, X. & Heitman, J. Cyclic AMP-dependent protein kinase regulates pseudohyphal differentiation in Saccharomyces cerevisiae. Mol. Cell. Biol. 19, 4874–4887 (1999)
Jonassen, I., Collins, J. F. & Higgins, D. G. Finding flexible patterns in unaligned protein sequences. Protein Sci. 4, 1587–1595 (1995)
Hanrahan, J. & Snyder, M. Cytoskeletal activation of a checkpoint kinase. Mol. Cell 12, 663–673 (2003)
Barral, Y., Parra, M., Bidlingmaier, S. & Snyder, M. Nim1-related kinases coordinate cell cycle progression with the organization of the peripheral cytoskeleton in yeast. Genes Dev. 13, 176–187 (1999)
Mewes, H. W., Albermann, K., Heumann, K., Liebl, S. & Pfeiffer, F. MIPS: a database for protein sequences, homology data and yeast genome information. Nucleic Acids Res. 25, 28–30 (1997)
Ubersax, J. A. et al. Targets of the cyclin-dependent kinase Cdk1. Nature 425, 859–864 (2003)
Loog, M. & Morgan, D. O. Cyclin specificity in the phosphorylation of cyclin-dependent kinase substrates. Nature 434, 104–108 (2005)
Gruhler, A. et al. Quantitative phosphoproteomics applied to the yeast pheromone signalling pathway. Mol. Cell Proteomics 4, 310–327 (2005)
Huse, M. & Kuriyan, J. The conformational plasticity of protein kinases. Cell 109, 275–282 (2002)
Moffat, J. & Andrews, B. Late-G1 cyclin-CDK activity is essential for control of cell morphogenesis in budding yeast. Nature Cell Biol. 6, 59–66 (2004)
Breitkreutz, B. J., Stark, C. & Tyers, M. Osprey: a network visualization system. Genome Biol. 4, R22.1–4 (2003)
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003)
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003)
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999)
Acknowledgements
We thank J. Tang for help with the initial Cdc28 preparations, and D. Gelperin, J. Mok and K. Wise for comments on the manuscript. M.S. was funded by grants from the NIH; J.P., G.D. and J.F. were funded by NIH predoctoral fellowships, and B.A. and M.T. were funded by grants from the Canadian Institutes of Health Research. M.J.R.S. was funded by a project grant from the Wellcome Trust, UK.Author Contributions Assay development was performed by H.Z., J.P., G.D., G.M. and M.S. Proteome chips were prepared by G.M., B.S. and P.F.P. at Invitrogen. G.J. contributed transcription factors for the arrays. Kinase assays were performed by J.P., G.D., H.Z. and M.S. Most kinases were prepared by J.P. and G.D. Additional kinases were provided by A.B., R.S., R.R.M., M.C.S., N.R., S.J.L., A.S.M., M.J.R.S., D.F.S, C.D.V., M.T. and B.A. Data analysis was performed by X.Z., J.P. and G.D. Consensus mapping was by H.G. In vitro solution validations were performed by G.M. and L.M. In vivo substrate validations were performed by J.P., G.D. and J.F. Most assays and analyses were performed in the laboratory of M.S. with contributions from M.G.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
G.M., L.M., B.S. and P.F.P. are employed by Invitrogen. M.S. has financial interests in Invitrogen.
Supplementary information
Supplementary Tables
Supplementary Table 1 shows kinases and their targets that share same MIPS functionary categories. Supplementary Table 2 lists all the known kinase-substrate phosphorylation events found in our study. (PDF 138 kb)
Supplementary Figures
Supplementary Figure 1 shows the distribution of phosphorylation events. Number of kinases for each substrate (top). Number of substrates for each kinase (bottom). Supplementary Figure 2 details solution validation of yeast proteome identified protien kinase substrates. (PDF 194 kb)
Supplementary Data 1
List of kinases analysed by our protein chip assay. (PDF 3 kb)
Supplementary Data 2
This file summarizes the substrate report for all the kinases tested. * This file was incorrectly uploaded as Supplementary Data 3 at time of publication. This was corrected on 03 February 2006. (XLS 950 kb)
Supplementary Data 3
All 8 regulatory modules including phosphorylation links, protein-protein interaction and transcriptional regulation. * This file was incorrectly uploaded as Supplementary Data 2 at time of publication. This was corrected on 03 February 2006. (XLS 1292 kb)
Rights and permissions
About this article
Cite this article
Ptacek, J., Devgan, G., Michaud, G. et al. Global analysis of protein phosphorylation in yeast. Nature 438, 679–684 (2005). https://doi.org/10.1038/nature04187
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04187
This article is cited by
-
RGS4 impacts carbohydrate and siderophore metabolism in Trichoderma reesei
BMC Genomics (2023)
-
In situ digestion-assisted multi-template imprinted nanoparticles for efficient analysis of protein phosphorylation
Microchimica Acta (2023)
-
Analog-sensitive Cdk1 as a tool to study mitotic exit: protein phosphatase 1 is required downstream from Cdk1 inactivation in budding yeast
Chromosome Research (2023)
-
Prediction of serine phosphorylation sites mapping on Schizosaccharomyces Pombe by fusing three encoding schemes with the random forest classifier
Scientific Reports (2022)
-
Drug-dependent growth curve reshaping reveals mechanisms of antifungal resistance in Saccharomyces cerevisiae
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.