Abstract
Vascular plants evolved in the Middle to Late Silurian period, about 420 million years ago1. The fossil record indicates that these primitive plants had branched stems with sporangia but no leaves. Leaf-like lateral outgrowths subsequently evolved on at least two independent occasions2,3,4. In extant plants, these events are represented by microphyllous leaves in lycophytes (clubmosses, spikemosses and quillworts) and megaphyllous leaves in euphyllophytes (ferns, gymnosperms and angiosperms). Our current understanding of how leaves develop is restricted to processes that operate during megaphyll formation. Because microphylls and megaphylls evolved independently, different mechanisms might be required for leaf formation. Here we show that this is not so. Gene expression data from a microphyllous lycophyte, phylogenetic analyses, and a cross-species complementation experiment all show that a common developmental mechanism can underpin both microphyll and megaphyll formation. We propose that this mechanism might have operated originally in the context of primitive plant apices to facilitate bifurcation. Recruitment of this pathway to form leaves occurred independently and in parallel in different plant lineages.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Edwards, D., Feehan, J. & Smith, D. G. A late Wenlock flora from County Tipperary, Ireland. Bot. J. Linn. Soc. 86, 19–36 (1983)
Gifford, E. M. & Foster, A. S. Morphology and Evolution of Vascular Plants (Freeman, New York, 1989)
Boyce, C. K. & Knoll, A. H. Evolution of developmental potential and the multiple independent origin of leaves in Paleozoic vascular plants. Paleobiology 28, 70–100 (2002)
Kenrick, P. & Crane, P. R. The Origin and Early Diversification of Land Plants: A Cladistic Study (Smithsonian Institution Press, London, 1997)
Jackson, D., Veit, B. & Hake, S. Expression of maize Knotted1 related homeobox genes in the shoot apical meristem predicts patterns of morphogenesis in the vegetative shoot. Development 120, 404–413 (1994)
Lincoln, C., Long, J., Yamaguchi, J., Serikawa, K. & Hake, S. A. Knotted1-like homeobox gene in Arabidopsis is expressed in the vegetative meristem and dramatically alters leaf morphology when overexpressed in transgenic plants. Plant Cell 6, 1859–1876 (1994)
Nishimura, A., Tamaoki, M., Sato, Y. & Matsuoka, M. The expression of tobacco Knotted1-type class1 homeobox genes correspond to regions predicted by the cytohistological zonation model. Plant J. 18, 337–347 (1999)
Long, J. A., Moan, E. I., Medford, J. I. & Barton, M. K. A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis . Nature 379, 66–69 (1996)
Sentoku, N. et al. Regional expression of the rice KN1-type homeobox gene family during embryo, shoot and flower development. Plant Cell 11, 1651–1663 (1999)
Hofer, J. M., Gourlay, C. W., Michael, A. & Ellis, T. H. Expression of a class 1 Knotted1-like homeobox gene is down-regulated in pea compound leaf primordia. Plant Mol. Biol. 45, 387–398 (2001)
Hareven, D., Gutfinger, T., Parnis, A., Eshed, Y. & Lifschitz, E. The making of a compound leaf: genetic manipulation of leaf architecture in tomato. Cell 84, 735–744 (1996)
Muller, J. et al. In vitro interactions between barley TALE homeodomain proteins suggest a role for protein-protein associations in the regulation of Knox gene function. Plant J. 27, 13–23 (2001)
Byrne, M. E. et al. ASYMMETRIC LEAVES1 mediates leaf patterning and stem cell function in Arabidopsis . Nature 408, 967–971 (2000)
Schneeberger, R., Tsiantis, M., Freeling, M. & Langdale, J. A. The rough sheath2 gene negatively regulates homeobox gene expression during maize leaf development. Development 125, 2857–2865 (1998)
Tsiantis, M., Schneeberger, R., Golz, J. F., Freeling, M. & Langdale, J. A. The maize rough sheath2 gene and leaf development programs in monocot and dicot plants. Science 284, 154–156 (1999)
Timmermans, M. C. P., Hudson, A., Becraft, P. W. & Nelson, T. ROUGH SHEATH2: A myb protein that represses Knox homeobox genes in maize lateral organ primordia. Science 284, 151–153 (1999)
Smith, L. G., Greene, B., Veit, B. & Hake, S. A dominant mutation in the maize homeobox gene, Knotted-1, causes its ectopic expression in leaf cells with altered fates. Development 116, 21–30 (1992)
McHale, N. A. & Koning, R. E. PHANTASTICA regulates development of the adaxial mesophyll in Nicotiana leaves. Plant Cell 16, 1251–1262 (2004)
Wikstrom, N., Savolainen, V. & Chase, M. W. Evolution of the angiosperms: calibrating the family tree. Proc. R. Soc. Lond. B 268, 2211–2220 (2001)
Tsiantis, M. & Hay, A. Comparative plant development: the time of the leaf? Nature Rev. Genet. 4, 169–180 (2003)
Steeves, T. A. & Sussex, I. M. Studies on the development of excised leaves in sterile culture. Am. J. Bot. 44, 665–673 (1957)
Bierhorst, D. W. On the stem apex, leaf initiation and early leaf ontogeny in filicalean ferns. Am. J. Bot. 64, 125–152 (1977)
Bharathan, G. et al. Homologies in leaf form inferred from KNOXI gene expression during development. Science 296, 1858–1860 (2002)
Kim, M., McCormick, S., Timmermans, M. & Sinha, N. The expression domain of PHANTASTICA determines leaflet placement in compound leaves. Nature 424, 438–443 (2003)
Kim, M. et al. Reduced leaf complexity in tomato wiry mutants suggests a role for PHAN and KNOX genes in generating compound leaves. Development 130, 4405–4415 (2003)
Theodoris, G., Inada, N. & Freeling, M. Conservation and molecular dissection of ROUGH SHEATH2 and ASYMMETRIC LEAVES1 function in leaf development. Proc. Natl Acad. Sci. USA 100, 6837–6842 (2003)
Pautot, V. et al. KNAT2: evidence for a link between Knotted-like genes and carpel development. Plant Cell 13, 1719–1734 (2001)
Golz, J. F., Keck, E. J. & Hudson, A. Spontaneous mutations in KNOX genes give rise to a novel floral structure in antirrhinum. Curr. Biol. 12, 515–522 (2002)
Parnis, A. et al. The dominant developmental mutants of tomato, Mouse-Ear and Curl, are associated with distinct modes of abnormal transcriptional regulation of a Knotted gene. Plant Cell 9, 2143–2158 (1997)
Janssen, B.-J., Lund, L. & Sinha, N. Overexpression of a homeobox gene LeT6 reveals indeterminate features in the tomato compound leaf. Plant Physiol. 117, 771–786 (1998)
Acknowledgements
We thank D. Stork for technical help; C. Hammond and T. Gardner for help during their undergraduate degrees (funded by the Wellcome Trust and a Nuffield Bursary, respectively); M. Tsiantis and G. Theodoris for rs2 fusion constructs; M. Tsiantis and M. Knight for criticizing our manuscript; M. Tsiantis, D. Barker and the Oxford Systematics Discussion Group for discussions; and J. Hofer, A. Hudson and N. McHale for the use of unpublished data presented in Fig. 3. D.L.A. was the recipient of a Sainsbury PhD studentship. The work was funded by a grant to J.A.L. and R.W.S. from the Biotechnology and Biological Sciences Research Council.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure S1
Hybridization analysis of KNOX and ARP gene families in S. kraussiana. (PPT 280 kb)
Supplementary Figure S2
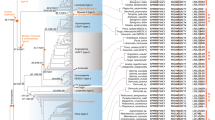
Sequence alignment and phylogenetic relationships of Selaginella class I KNOX and ARP genes. (PPT 1684 kb)
Supplementary Figure S3
Phylogenetic relationships in the KNOX gene family. (PPT 619 kb)
Supplementary Figure S4
Phylogenetic relationships within the MYB gene super-family. (PPT 77 kb)
Supplementary Figure Legends
Legends to accompany the above Supplementary Figures. This file also contains additional references. (DOC 98 kb)
Rights and permissions
About this article
Cite this article
Harrison, C., Corley, S., Moylan, E. et al. Independent recruitment of a conserved developmental mechanism during leaf evolution. Nature 434, 509–514 (2005). https://doi.org/10.1038/nature03410
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03410
This article is cited by
-
Origin and function of stomata in the moss Physcomitrella patens
Nature Plants (2016)
-
What determines a leaf's shape?
EvoDevo (2014)
-
Co-option of the polarity gene network shapes filament morphology in angiosperms
Scientific Reports (2014)
-
The function of the RNA-binding protein TEL1 in moss reveals ancient regulatory mechanisms of shoot development
Plant Molecular Biology (2012)
-
A complex case of simple leaves: indeterminate leaves co-express ARP and KNOX1 genes
Development Genes and Evolution (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.