Abstract
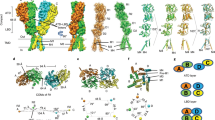
Ionotropic glutamate receptors (iGluRs) mediate excitatory synaptic transmission in vertebrates and invertebrates through ligand-induced opening of transmembrane ion channels. iGluRs are segregated into three subtypes according to their sensitivity to the agonists AMPA (α-amino-3-hydroxy-5-methyl-4-isoxazole propionic acid), kainate (a structural analogue of glutamate) or NMDA (N-methyl-D-aspartate) (Fig. 1). iGluRs are important in the development and function of the nervous system, are essential in memory and learning, and are either implicated in or have causal roles in dysfunctions ranging from Alzheimer's, Parkinson's and Huntington's diseases, schizophrenia, epilepsy and Rasmussen's encephalitis to stroke1,2. Development of iGluR agonists and antagonists has been hampered by a lack of high-resolution structural information. Here we describe the crystal structure of an iGluR ligand-binding region in a complex with the neurotoxin (agonist) kainate. The bilobed structure shows the determinants of receptor–agonist interactions and how ligand-binding specificity and affinity are altered by remote residues and the redox state of the conserved disulphide bond. The structure indicates mechanisms for allosteric effector action and for ligand-induced channel gating. The information provided by this structure will be essential in designing new ligands.

The numbering of substituents on the pyrrolidine ring of kainate is as follows: nitrogen, 1; carboxylate, 2; carboxyl methyl, 3; and isopropenyl, 4.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hollmann, M. & Heinemann, S. Cloned glutamate receptors. Annu. Rev. Neurosci. 17, 31–108 (1994).
Rogers, S. W. et al. Autoantibodies to glutamate receptor GluR3 in Rasmussen's encephalitis. Science 265, 648–651 (1994).
Stern-Bach, Y. et al. Agonist selectivity of glutamate receptors is specified by two domains structurally related to bacterial amino acid-binding proteins. Neuron 13, 1345–1357 (1994).
Kuusinen, A., Arvola, M. & Keinänen, K. Molecular dissection of the agonist binding site of an AMPA receptor. EMBO J. 14, 6327–6332 (1995).
Paas, Y. The macro- and microarchitectures of the ligand-binding domain of glutamate receptors. Trends Neurosci. 21, 117–125 (1998).
Chen, G.-Q., Sun, Y., Jin, R. & Gouaux, E. Probing the ligand binding domain of the GluR2 receptor by proteolysis and deletion mutagenesis defines domain boundaries and yields a crystallizable construct. Protein Sci. (in the press).
Sommer, B. et al. Flip and flop: a cell-specific functional switch in glutamate-operated channels of the CNS. Science 249, 1580–1585 (1990).
Hendrickson, W. A. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254, 51–58 (1991).
Nakanishi, N., Shneider, N. A. & Axel, R. Afamily of glutamate receptor genes: evidence for the formation of heteromultimeric receptors with distinct channel properties. Neuron 5, 569–581 (1990).
Sun, Y.-J., Rose, J., Wang, B.-C. & Hsiao, C.-D. The structure of glutamine-binding protein complexed with glutamine at 1.94 å resolution: comparisons with other amino acid binding proteins. J. Mol. Biol. 278, 219–229 (1998).
Keinänen, K., Arvola, M., Kuusinen, A. & Johnson, M. Ligand recognition in glutamate receptors: insights from mutagenesis of the soluble α-amino-3-hydroxy-5-methyl-4-isoxazole propionic acid (AMPA)-binding domain of glutamate receptor type D (GluR-D). Biochem. Soc. Trans. 25, 835–838 (1997).
Laube, B., Hirai, H., Sturgess, M., Betz, H. & Kuhse, J. Molecular determinants of agonist discrimination by NMDA receptor subunits: analysis of the glutamate binding site on the NR2B subunit. Neuron 18, 493–503 (1997).
Paas, Y., Eisenstein, M., Medevielle, F., Teichberg, V. I. & Devillers-Thiéry, A. Identification of the amino acid subsets accounting for the ligand binding specificity of a glutamate receptor. Neuron 17, 979–990 (1996).
Hirai, H., Kirsch, J., Laube, B., Betz, H. & Kuhse, J. The glycine binding site of the N-methyl-D-aspartate receptor subunit NR1: identification of novel determinants of co-agonist potentiation in the extracellular M3-M4 loop region. Proc. Natl Acad. Sci. USA 93, 6031–6036 (1996).
Swanson, G. T., Gereau, R. W. IV, Green, T. & Heinemann, S. F. Identification of amino acid residues that control functional behavior in GluR5 and GluR6 kainate receptors. Neuron 19, 913–926 (1997).
Jacobson, B. L., He, J. J., Lemon, D. D. & Quiocho, F. A. Interdomain salt bridges modulate ligand-induced domain motion of the sulfate receptor protein for active transport. J. Mol. Biol. 223, 27–30 (1992).
Li, F., Owens, N. & Verdoorn, T. A. Functional effects of mutations in the putative agonist binding region of recombinant α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptors. Mol. Pharmacol. 47, 148–154 (1995).
Aizenman, E., Lipton, S. A. & Loring, R. H. Selective modulation of NMDA responses by reduction and oxidation. Neuron 2, 1257–1263 (1989).
Sutcliffe, M. J., Wo, Z. G. & Oswald, R. E. Three-dimensional models of non-NMDA glutamate receptors. Biophys. J. 70, 1575–1589 (1996).
Partin, K. M., Bowie, D. & Mayer, M. L. Structural determinants of allosteric regulation in alternatively spliced AMPA receptors. Neuron 14, 833–843 (1995).
Partin, K. M., Fleck, M. W. & Mayer, M. L. AMPA receptor flip/flop mutants affecting deactivation, desensitization, and modulation by cyclothiazide, aniracetam, and thiocyanate. J. Neurosci. 16, 6634–6647 (1996).
Otwinowsky, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Cowtan, K. D. & Main, P. Phase combination and cross validation in iterated density-modification calculations. Acta Crystallogr. D 42, 43–48 (1996).
Read, R. J. Model phases: probabilities and bias. Methods Enzymol. 277, 110–128 (1997).
Jones, T. A. & Kjeldgaard, M. Electron-density map interpretation. Methods Enzymol. 277, 173–208 (1997).
Brünger, A. T. X-PLOR. Version 3.1. A System for X-ray Crystallography and NMR 1–382 (Yale Univ. Press, New Haven, 1992).
Collaborative Computational Project Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Hollmann, M., Maron, C. & Heinemann, S. N-Glycosylation site tagging suggests a three transmembrane domain topology for the glutamate receptor GluR1. Neuron 13, 1331–1343 (1994).
Lomeli, H. et al. Control of kinetic properties of AMPA receptor channels by nuclear RNA editing. Science 266, 1709–1713 (1994).
Acknowledgements
We thank C. Lima and other members of the Hendrickson laboratory for assistance; J. Lidestri for maintenance of the X-ray laboratory at Columbia University; A. Karlin, J. Javitch and M.Akabas for advice; S. Heinemann and M. Hartley for glutamate-receptor clones; C. Ogata for help with MAD data collection at NSLS X4a; T. Terwilliger for advice on the use of Solve; and W. Minor and Z.Otwinoski for a new version of the HKL suite of programs. Extra MAD data were collected at the Cornell High Energy Synchrotron Source (CHESS), which is supported by the NSF, using the Macromolecular Diffraction at CHESS (MacCHESS) facility, which is supported by an award from the NIH. N.A. was supported by an NIH Ophthalmology training grant. E.G. is an NSF Young Investigator and the recipient of an Alfred P. Sloan Research Fellowship.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
Supplementary Information
Rights and permissions
About this article
Cite this article
Armstrong, N., Sun, Y., Chen, GQ. et al. Structure of a glutamate-receptor ligand-binding core in complex with kainate. Nature 395, 913–917 (1998). https://doi.org/10.1038/27692
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/27692
This article is cited by
-
Streptococcus mutans glutamate binding protein (GlnH) as antigen target for a mucosal anti-caries vaccine
Brazilian Journal of Microbiology (2022)
-
Genome-wide identification, characterization and classification of ionotropic glutamate receptor genes (iGluRs) in the malaria vector Anopheles sinensis (Diptera: Culicidae)
Parasites & Vectors (2018)
-
Mapping the membrane proteome of anaerobic gut fungi identifies a wealth of carbohydrate binding proteins and transporters
Microbial Cell Factories (2016)
-
Architecture of fully occupied GluA2 AMPA receptor–TARP complex elucidated by cryo-EM
Nature (2016)
-
Structural mechanisms of activation and desensitization in neurotransmitter-gated ion channels
Nature Structural & Molecular Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.