Abstract
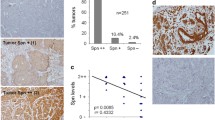
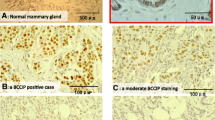
Expression of survivin, a member of the inhibitor-of-apoptosis (IAP) family, is elevated in fetal tissues and in various human cancers originating in the breast, lung, prostate, colon, pancreas, and stomach. Since overexpression of the survivin gene has been linked to poor patient survival in several cancers, survivin may be an important prognostic marker. Mechanisms up-regulating survivin gene expression in cancer are poorly understood. Recently, wild-type p53 was found to repress expression of the survivin gene by binding to the survivin promoter, thereby inhibiting promoter activity. Further, loss of heterozygosity (LOH) at 17p13 distal to the p53 gene is associated with more aggressive behavior of breast cancers. We therefore tested the hypothesis that not only p53 gene mutation but also LOH at 17p13 can up-regulate survivin expression in breast cancer. Survivin mRNA expression was greater in cancers than in uninvolved tissues (p < 0.0001). Mutations of the p53 gene were detected in 5 of 25 tumors; higher survivin gene expression was evident in these. LOH at the D17S938 locus (17p13.1) was found in 10 of 25 tumors, and most of these also showed increased survivin gene expression. Thus expression of survivin may be regulated not only by p53 but additionally by a putative tumor suppressor gene located at 17p13 distal to the p53 gene.
Similar content being viewed by others
References
Saikumar P, Dong Z, Mikhailov V, Denton M, Weinberg JM, Venkatachalam MA: Apoptosis: definition, mechanisms, and relevance to disease. Am J Med 107: 489–506, 1999
Jaattela M: Escaping cell death: survival proteins in cancer. Exp Cell Res 248: 30–43, 1999
Tamm I, Wang Y, Sausville E, Scudiero DA, Vigna N, Oltersdorf T, Reed JCL: IAP-family protein survivin inhibits caspase activity and apoptosis induced by Fas (CD95), Bax, caspases, and anticancer drugs. Cancer Res 58: 5315–5320, 1998
Shin S, Sung BJ, Cho YS, Kim HJ, Ha NC, Hwang JI, Chung CW, Jung YK, Oh BH: An anti-apoptotic protein human survivin is a direct inhibitor of caspase-3 and-7. Biochemistry 40: 1117–1123, 2001
Ambrosini G, Adida C, Altieri DC: A novel anti-apoptosis gene, survivin, expressed in cancer and lymphoma. Nat Med 3: 917–921, 1997
Tanaka K, Iwamoto S, Gon G, Nohara T, Iwamoto M, Tanigawa N: Expression of survivin and its relationship to loss of apoptosis in breast carcinomas. Clin Cancer Res 6: 127–134, 2000
Monzo M, Rosell R, Felip E, Astudillo J, Sanchez JJ, Maestre J, Martin C, Font A, Barnadas A, Abad A: A novel anti-apoptosis gene: re-expression of survivin messenger RNA as a prognosis marker in non-small-cell lung cancers. J Clin Oncol 17: 2100–2104, 1999
Kawasaki H, Altieri DC, Lu CD, Toyoda M, Tenjo T, Tanigawa N: Inhibition of apoptosis by survivin predicts shorter survival rates in colorectal cancer. Cancer Res 58: 5071–5074, 1998
Lu CD, Altieri DC, Tanigawa N: Expression of a novel antiapoptosis gene, survivin, correlated with tumor cell apoptosis and p53 accumulation in gastric carcinomas. Cancer Res 58: 1808–1812, 1998
Sarela AI, Macadam RC, Farmery SM, Markham AF, Guillou PJ: Expression of the antiapoptosis gene, survivin, predicts death from recurrent colorectal carcinoma. Gut 46: 645–650, 2000
Hoffman WH, Biade S, Zilfou JT, Chen J, Murphy M: Transcriptional repression of the anti-apoptotic survivin gene by wild type p53. J Biol Chem 277: 3247–3257, 2002
Seitz S, Poppe K, Fischer J, Nothnagel A, Estevez-Schwarz L, Haensch W, Schlag PM, Scherneck S: Detailed deletion mapping in sporadic breast cancer at chromosomal region 17p13 distal to the TP53 gene: association with clinicopathological parameters. J Pathol 194: 318–326, 2001
Asanuma K, Moriai R, Yajima T, Yagihashi A, Yamada M, Kobayashi D, Watanabe N: Survivin as a radioresistance factor in pancreatic cancer. Jpn J Cancer Res 91: 1204–1209, 2000
Mahotka C, Wenzel M, Springer E, Gabbert HE, Gerharz CD: Survivin-deltaEx3 and survivin-2B: two novel splice variants of the apoptosis inhibitor survivin with different antiapoptotic properties. Cancer Res 59: 6097–6102, 1999
Tsuji N, Kamagata C, Furuya M, Kobayashi D, Yagihashi A, Morita T, Horita S, Watanabe N: Selection of an internal control gene for quantitation of mRNA in colonic tissues. Anticancer Res 22: 4173–4178, 2002
Holland PM, Abramson RD, Watson R, Gelfand DH: Detection of speci c polymerase chain reaction product by utilizing the 5′-3′ exonuclease activity of Thermus aquaticus DNA polymerase. Proc Natl Acad Sci U S A 88: 7276–7280, 1991
Yajima T, Yagihashi A, Kameshima H, Kobayashi D, Furuya D, Hirata K, Watanabe N: Quantitative reverse transcription-PCR assay of the RNA component of human telomerase using the TaqMan fluorogenic detection system. Clin Chem 44: 2441–2445, 1998
Michalovitz D, Halevy O, Oren M: Conditional inhibition of transformation and of cell proliferation by a temperature-sensitive mutant of p53. Cell 62: 671–680, 1990
Reihsaus E, Kohler M, Kraiss S, Oren M, Montenarh M: Regulation of the level of the oncoprotein p53 in non-transformed and transformed cells. Oncogene 5: 137–145, 1990
Finlay CA, Hinds PW, Tan TH, Eliyahu D, Oren M, Levine AJ: Activating mutations for transformation by p53 produce a gene product that forms an hsc70-p53 complex with an altered half-life. Mol Cell Biol 8: 531–539, 1988
Cesarman E, Inghirami G, Chadburn A, Knowles DM: High levels of p53 protein expression do not correlate with p53 gene mutations in anaplastic large cell lymphoma. Am J Pathol 143: 845–856, 1993
Matsushima AY, Cesarman E, Chadburn A, Knowles DM: Post-thymic T cell lymphomas frequently overexpress p53 protein but infrequently exhibit p53 gene mutations. Am J Pathol 144: 573–584, 1994
Kennedy SM, Macgeogh C, Jaffe R, Spurr NK: Overexpression of the oncoprotein p53 in primary hepatic tumors of childhood does not correlate with gene mutations. Hum Pathol 25: 438–442, 1994
Leahy DT, Salman R, Mulcahy H, Sheahan K, O'Donoghue DP, Parfrey NA: Prognostic signi cance of p53 abnormalities in colorectal carcinoma detected by PCR-SSCP and immunohistochemical analysis. J Pathol 180: 364–370, 1996
Ho WL, Chang JW, Tseng RC, Chen JT, Chen CY, Jou YS, Wang YC: Loss of heterozygosity at loci of candidate tumor suppressor genes in microdissected primary non-small cell lung cancer. Cancer Detect Prev 26: 343–349, 2002
Launonen V, Mannermaa A, Stenback F, Kosma VM, Puistola U, Huusko P, Anttila M, Bloigu R, Saarikoski S, Kauppila A, Winqvist R: Loss of heterozygosity at chromosomes 3, 6, 8, 11, 16, and 17 in ovarian cancer: correlation to clinicopathological variables. Cancer Genet Cytogenet 122: 49–54, 2000
Haupt Y, Maya R, Kazaz A, Oren M: Mdm2 promotes the rapid degradation of p53. Nature 387: 296–299, 1997
Sjostrom J, Blomqvist C, Heikkila P, Boguslawski KV, Raisanen-Sokolowski A, Bengtsson NO, Mjaaland I, Malmstrom P, Ostenstadt B, Bergh J, Wist E, Valvere V, Saksela E: Predictive value of p53, mdm-2, p21, and mib-1 for chemotherapy response in advanced breast cancer. Clin Cancer Res 6: 3103–3110, 2000
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Tsuji, N., Furuse, K., Asanuma, K. et al. Mutations of the p53 Gene and Loss of Heterozygosity at Chromosome 17p13.1 are Associated with Increased Survivin Expression in Breast Cancer. Breast Cancer Res Treat 87, 23–31 (2004). https://doi.org/10.1023/B:BREA.0000041575.73262.aa
Issue Date:
DOI: https://doi.org/10.1023/B:BREA.0000041575.73262.aa