Abstract
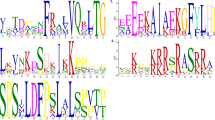
R2R3-MYB transcription factors have been implicated in a diversity of plant-specific processes. Among the functions attributed to myb factors is the determination of cell shape, including regulation of trichome length and density. Because myb transcription factors are likely to play a role in cotton fiber development, the molecular evolutionary properties of six MYB genes previously shown to be expressed in cotton fiber initiation were examined. In accordance with their presumed central role, each of the genes display conservative substitution patterns and limited sequence divergence in diploid members of the genus Gossypium, and this pattern is conserved in allotetraploid cottons. In contrast to highly reiterated rDNA repeats, GhMYB homologues (duplicated gene pairs) exhibit no evidence of concerted evolution, but instead appear to evolve independently in the allopolyploid nucleus. Expression patterns for the MYB genes were examined in several organs to determine if there have been changes in expression patterns between the diploids (G. raimondii and G. arboreum) and the tetraploid (G. hirsutum) or between the duplicated copies in the tetraploid. Spatial and temporal expression patterns appear to have been evolutionarily conserved, both during divergence of the diploid parents of allopolyploid cotton and following polyploid formation. However, the duplicated copies of MYB1 in the tetraploid are not expressed at equal levels or equivalently in all organs, suggesting possible functional differentiation.
Similar content being viewed by others
References
Applequist, W.L., Cronn, R.C., Wendel, J.F., 2001. Co0mparative development of fiber in wild and cultivated cotton. Evol. Dev. 3, 1–15.
Baum, D.A., 1998. The evolution of plant development. Curr. Opin. Plant Biol. 1, 79–86.
Bender, J., Fink, G., 1998. A MYB homolog, ATR1, activates tryptophan gene expression in Arabidopsis. Proc. Natl. Acad. Sci. USA 95, 5655–5660.
Borevitz, J.O., Xia, Y.J., Dolan L., Schiefelbein, J., 1998. Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 12, 2383–2393.
Braun, E.L., Grotewold, E., 1999. Newly discovered plant c-MYBlike genes rewrite the evolution of plant myb gene family. Plant Physio. 121, 21–24.
Brubaker, C.L., Wendel, J.F., 1994. Reevaluating the origin of domesticated cotton (Gossypium hirsutum: Malvaceae) using nuclear restriction fragment length polymorphisms (RFLPs). Amer. J. Bot. 81, 1309–1326.
Brubaker, C.L., Bourland, F.M., Wendel, J.F., 1999. The origin and domestication of cotton. In: Smith WC (ed.) Cotton: Origin, History, Technology, and Production. J. Wiley and Sons, Inc., New York, N.Y.
Brubaker, C.L., Paterson, A.H., Wendel, J.F., 1999. Comparative genetic mapping of allotetraploid cotton and its diploid progenitors. Genome 42, 184–203.
Cronn, R.C., Small, R.L., Wendel, J.F., 1999. Duplicated genes evolve independently following polyploid formation in cotton. Proc. Natl. Acad. Sci. USA 96, 14406–14411.
Cronn, R.C., Small, R.L., Haselkorn, T.S., Wendel, J.F., 2002. Rapid diversification of the cotton genus (Gossypium: Malvaceae) revealed by analysis of sixteen nuclear and chloroplast genes. Amer. J. Bot. 89, 707–725.
Cronn, R., Cedroni, M., Grover, C., Haskelkorn, T., Wendel, J.F., 2002. PCR-mediated recombination in a polyploid plant. Theor. Appl. Genet. 104, 482–489
Cubas, P., Coen, E., Martinez-Zapater, J.M., 2001. Ancient asymmetries in the evolution of flowers. Curr. Biol. 11, 1050–1052.
Doebley, J., Lukens, L., 1998. Transcriptional regulators and the evolution of plant form. Plant Cell 10, 1075–1082.
Force, A., Lynch, M., Pickett, F.B., Amores, A., Yan, Y.L., Postlethwait, J., 1999. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545.
Fryxell, P.A., 1979. The natural history of the cotton tribe. Texas A&M University Press, College Station, T.X.
Gocal, G.F.W., Sheldon, C.C., Gubler, F., Moritz, T., Bagnall, D.J., MacMillan, C.P., Li, S.F., Parish, R.W., Dennis, E.S., Wiegel, D., King, R.W., 2001. GAMYB-like genes, flowering, and gibberellin signaling in Arabidopsis. Plant Physiol. 127, 1682–1693.
Hurst, L.D., Pal, C., 2001. Evidence for purifying selection acting on silent sites in BRCA1. Trends Genet. 17, 62–65.
Jin, H., Martin, C., 1999. Multifunctionality and diversity within the plant MYB gene family. Plant Mol. Biol. 41, 577–585.
Kirik, V., Kolle, K., Wohlfarth, T., Misera, S., Baumlein, H., 1998. Ectopic expression of a novel MYB gene modifies the architecture of the Arabidopsis inflorescence. Plant J. 13, 729–742.
Kranz, H., Denekamp, M., Greco, R., Jin, H., Leyva, A., Meissner, R.C., Petroni, K., Urzainqui, A., Bevan, M., Martin, C., Smeekens, S., Tonelli, C., Paz-Ares, J., Weisshaar, B., 1998. Towards functional characterisation of the members of the R2R3-MYB gene family from Arabidopsis thaliana. Plant J. 16, 263–276.
Kranz, H., Scholz, K., Weisshaar, B., 2000. c-Myb oncogenelike genes encoding three MYB repeats occur in all major plant lineages. Plant J. 21, 231–235.
Loguercio, L.L., Zhang, J.Q., Wilkins, T.A., 1999. Differential regulation of six novel MYB-domain genes defines two distinct expression patterns in allotetraploid cotton (Gossypium hirsutum L.). Mol. Gen. Genet. 261, 660–671.
Luo, D., Carpenter, R., Copsey, L., Vincent, C., Clark, J., Coen, E., 1999. Control of organ asymmetry in flowers of Antirrhinum. Cell 99, 367–376.
Lynch, M. and Force, A., 2000. The probability of duplicate gene preservation by subfunctionalization. Genetics 154, 459–473.
Martin, C. and Paz-Ares, J., 1997. MYB transcription factors in plants. Trends Genet. 13, 67–73.
Meissner, R.C., Hailing, J., Cominelli, E., Denekamp, M., Fuertes, A., Greco, R., Kranz, H.D., Penfield, S., Petroni, K., Urzainqui, A., Martin, C., Paz-Arez, J., Smeekens, S., Tonelli, C., Weisshaar, B., Baumann, E., Klimyuk, V., Marillonnet, S., Patel, K., Speulman, E., Tissier, A.F., Bouchez, D., Jones, J.J.D., Pereira, A., Weisman, E., Bevan, M., 1999. Function search in a large transcription factor gene family in Arabidopsis: Assessing the potential of reverse genetics to identify insertional mutations in R2R3 MYB genes. Plant Cell 11, 1827–1840.
Noda, K Glover, B.J., Linstead, P., Martin, C., 1994. Flower colour intensity depends on specialized cell shape controlled by a Mybrelated transcription factor. Nature 369, 661–664.
Orita, M., Iwahana, H., Kanazawa, H., Hayashi, K., Sekiya, T., 1989. Detection of polymorphisms of human DNA by gel electrophoresis as single-strand conformation polymorphisms. Proc. Natl. Acad. Sci. USA 86, 2766–2770.
Payne, T., Clement, J., Arnold, D., Lloyd, A., 1999. Heterologous myb genes distinct from GL1 enhance trichome production when overexpressed in Nicotiana tabacum. Development 126, 671–682.
Penfield, S., Meissner, R.C., Shoue, D.A., Carpita, N.C., Bevan, M.W., 2001. MYB61 is required for mucilage deposition and extrusion in the Arabidopsis seed coat. Plant Cell 13, 2777–2791.
Purugganan, M.D., 1998 The molecular evolution of development. BioEssays 20, 700–711.
Rabinowicz, P.D., Braun, E.L., Wolfe, A.D., Bowen, B., Grotewold, E., 1999. Maize R2R3 MYB genes: sequence analysis reveals amplification in the higher plants. Genetics 153, 427–444.
Reichmann, J.L., Heard, J., Martin, G., Reuber, L., Jiang, C.Z., Keddie, J., Adam, L., Pineda, O., Ratcliffe, O.J., Samaha, R.R., Creelman, R., Pilgrim, M., Broun, P., Zhang, J.Z., Ghandehari, D., Sherman, B.K., Yu, G.L., 2000. Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290, 2105–2110.
Reinisch, A.R., Dong, J.M., Brubaker, C.L., Stelly, D.M., Wendel, J.F., Paterson, A.H., 1994. A detailed RFLP map of cotton, Gossypium hirsutum ?Gossypium barbadense: chromosome organization and evolution in a disomic polyploid genome. Genetics 138, 829–847.
Romero, I., Fuertes, A., Benito, B.J., Malpica, J.M., Leyva, A., Paz-Ares, J., 1998. More than 80 R2R3-MYB regulatory genes in the genome of Arabidopsis thaliana. Plant J. 14, 273–284.
Rozas, J., Rozas, R., 1999. DnaSP version 3: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15, 174–175.
Seelanan, T., Schnabel, A., Wendel, J.F., 1997. Congruence and consensus in the cotton tribe. Systematic Botany 22, 259–290.
Slabaugh, M.B., Huestis, G.M., Leonard, J., Holloway, J.L., Rosato, C., Hongtrakul, V., Martini, N., Toepfer, R., Voetz, M., Schell, J., Knapp, S.J., 1997. Sequence-based genetic markers for genes and gene families: single-stranded conformational polymorphisms for the fatty acid synthesis genes of Cuphea. Theor. Appl. Genet. 94, 400–408.
Small, R.L., Wendel, J.F., 2000. Copy number lability and evolutionary dynamics of the Adh gene family in diploid and tetraploid cotton (Gossypium). Genetics 155, 1913–1926.
Small, R.L., Ryburn, J.A., Cronn, R.C., Seelanan, T., Wendel, J.F., 1998. The tortoise and the hare: choosing between noncoding plastome and nuclear Adh sequences for phylogeny reconstruction in a recently diverged plant group. Amer. J. Bot. 85, 1301–1315.
Song, K., Osborn, T.C., 1994. A method for examining expression of homologous genes in plant polyploids. Plant Mol. Biol. 26, 1065–1071.
Stracke, R., Weber, M. Weisshaar, B., 2001. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. P. Biol. 4, 447–456.
Swofford, D.L, 1998. PAUP*: phylogenetic anaylsis using parsimony (* and other methods), Version 4.0b1. Sunderland, MA: Sinauer Associates, Inc.
Tajima, F., 1993. Simple methods for testing the molecular evolutionary clock hypothesis. Genetics 135, 599–607.
Wada, T., Tachibana, T., Shimura, Y., Okada, K., 1997. Epidermal cell differentiation in Arabidopsis determined by a MYB homolog, CPC. Science 277, 1113–1116.
Wendel, J.F., 1989 New World tetraploid cottons contain Old World cytoplasm. Proc. Acad. Natl. Sci. USA 86, 4132–4136.
Wendel, J.F., 2000. Genome evolution in polyploids. Pl. Mol. Biol. 42, 225–249.
Wendel, J.F., Olson, P.D., Stewart, JMcD, 1989. Genetic diversity, introgression, and independent domestication of Old World cultivated cottons. Amer. J. Bot. 76, 795–806.
Wendel, J.F., 1995. Cotton. In: N. Simmonds and J. Smartt (eds), Evolution of Crop Plants. Longman, London pp 358–366.
Wendel, J.F., Schnabel, A., Seelanan, T., 1995. Bidirectional interlocus concerted evolution following allopolyploid speciation in cotton (Gossypium). Proc. Natl. Acad. Sci. USA 92, 280–284.
Wilkins, T.A., Jernstedt, J.A., 1999. Molecular genetics of developing cotton fibers. In: Basra AS (ed.), Cotton Fibers. Haworth Press, New York pp. 231–267.
Wilkins, T.A., Smart, L.B., 1996. Isolation of RNA from plant tissue. In: A Laboratory Guide to RNA. Krieg PA (ed.) Wiley-Liss, Inc. pp. 21–40.
Yip, S.P., Hopkinson, D.A., Whitehouse, D.B., 1999. Improvement of SSCP analysis by use of denaturants. Biotechniques 27, 20–24.
Zhang. P., Chopra, S., Peterson, T., 2000. A segmental gene duplication generated differentially expressed myb-homologous genes in maize. Plant Cell 12, 2311–2322.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cedroni, M., Cronn, R., Adams, K. et al. Evolution and expression of MYB genes in diploid and polyploid cotton. Plant Mol Biol 51, 313–325 (2003). https://doi.org/10.1023/A:1022051100610
Issue Date:
DOI: https://doi.org/10.1023/A:1022051100610