Abstract
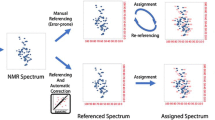
A suite of programs called CAMRA (Computer Aided Magnetic Resonance Assignment) has been developed for computer assisted residue-specific assignments of proteins. CAMRA consists of three units: ORB, CAPTURE and PROCESS. ORB predicts NMR chemical shifts for unassigned proteins using a chemical shift database of previously assigned homologous proteins supplemented by a statistically derived chemical shift database in which the shifts are categorized according to their residue, atom and secondary structure type. CAPTURE generates a list of valid peaks from NMR spectra by filtering out noise peaks and other artifacts and then separating the derived peak list into distinct spin systems. PROCESS combines the chemical shift predictions from ORB with the spin systems identified by CAPTURE to obtain residue specific assignments. PROCESS ranks the top choices for an assignment along with scores and confidence values. In contrast to other auto-assignment programs, CAMRA does not use any connectivity information but instead is based solely on matching predicted shifts with observed spin systems. As such, CAMRA represents a new and unique approach for the assignment of protein NMR spectra. CAMRA will be particularly useful in conjunction with other assignment methods and under special circumstances, such as the assignment of flexible regions in proteins where sufficient NOE information is generally not available. CAMRA was tested on two medium-sized proteins belonging to the chemokine family. It was found to be effective in predicting the assignment providing a database of previously assigned proteins with at least 30% sequence identity is available. CAMRA is versatile and can be used to include and evaluate heteronuclear and three-dimensional experiments.
Similar content being viewed by others
References
Baggiolini, M., Dewald, B. and Moser, B. (1996) Annu. Rev. Immunol., 15, 675–705.
Bartels, C., Billeter, M., Güntert, P. and Wüthrich, K. (1996) J. Biomol. NMR, 7, 207–213.
Bax, A. and Grzesiek, S. (1993) Acc. Chem. Res., 26, 131–138.
Bigam, C., Jellard, T., Gronwald, W. and Sykes, B.D. (1998) Magnetic Moments, 9, 9–12.
Clore, G.M. and Gronenborn, A.M. (1991) Science, 252, 1390–1399.
Croft, D., Kemmink, J., Neidig, K.-P. and Oschkinat, H. (1997) J. Biomol. NMR, 10, 207–219.
Crump, M.P., Gong, J.-H., Loetscher, P., Rajarathnam, K., Arenzana-Seisdedos, F., Virelizier, J.-L., Baggioloini, M., Sykes, B. D. and Clark-Lewis, I. (1997) EMBO J., 16, 6996–7007.
de Dios, A.C., Pearson, J.G. and Oldfield, E. (1993) Science, 5113, 1491–1496.
Fairbrother, W.J. and Skelton, N.J. (1996) Three dimensional structures of the chemokine family. In R. Horuk (ed.), Chemoattractant Ligands and their Receptors, CRC Press, London, pp. 55–86.
Friedrichs, M.S., Mueller, L. and Wittekind, M. (1994) J. Biomol. NMR, 4, 703–726.
Garrett, D.S., Powers, R., Gronenborn, A. and Clore, G.M. (1991) J. Magn. Reson., 94, 214–220.
Gronwald, W., Boyko, R.F., Sönnichsen, F.D., Wishart, D.S. and Sykes, B.D. (1997) J. Biomol. NMR, 10, 165–179.
Gronwald, W., Boyko, R.F. and Sykes, B.D. (1997) CABIOS, 13, 557–558.
Handel, T.M. and Domaille, P.J. (1996) Biochemistry, 33, 15283–15292.
Hare, B.J. and Prestegard, J.H. (1994) J. Biomol. NMR, 4, 35–46.
Kim, K.-S., Rajarathnam, K., Clark-Lewis, I. and Sykes, B.D. (1996) FEBS Lett., 395, 277–282.
Kjaer, M., Andersen, K.V. and Poulsen, F.M. (1994) Methods Enzymol., 239, 288–318.
Kleywegt, G.J., Boelens, R., Cox, M., Llinás, M. and Kaptein, R. (1991) J. Biomol. NMRi, 1, 23–47.
Kraulis, P.J. (1994) J. Mol. Biol., 243 696–718.
Lodi P.J., Garret, D.S., Kuszewski, J., Tsang, M.L.-w., Weatherbee, J.A., Leonard, W.J., Gronenborn, A.M. and Clore, G.M. (1994) Science, 263, 1762–1767.
Lukin, J.A., Gove, A.P. Talukdar, S.N. and Ho, C. (1997) J. Biomol. NMR, 9, 151–166.
Meadows, R.P., Olejniczak, E.T. and Fesik, S.W. (1994) J. Biomol. NMR, 4, 79–96.
Morelle, N., Brutscher, B., Simorre, J-P. and Morelle, M.D. (1995) J. Biomol. NMR, 5, 154–160.
Olson, J.B. Jr. and Markley, J.L. (1994) J. Biomol. NMR, 4, 385–410.
Oschkinat, H., Holak, T.A. and Cieslar, C. (1991) Biopolymers, 31, 699–712.
Ousterhout, J.K. (1994) Tcl and the Tk Toolkit, Addison-Wesley Professional Computing Series.
Piotto, M., Saudek, V. and Sklenar, V. (1992) J. Biomol. NMR, 2, 661–665.
Rajarathnam, K., Clark-Lewis, I. and Sykes, B.D. (1994) Biochemistry, 33, 6623–6630.
Rajarathnam, K., Clark-Lewis, I. and Sykes, B.D. (1995) Biochemistry, 34, 12983–12990.
Rajarathnam, K., Clark-Lewis, I., Dewald, B., Baggiolini, M. and Sykes, B.D. (1996) FEBS Lett., 399, 43–46.
Seavey, B.R., Farr, E.A., Westler, W.M. and Markley, J.L. (1991) J. Biomol. NMR, 1, 217–236.
Shaka, A.J., Lee, C.J. and Pines, A. (1988) J. Magn. Reson., 77, 274–293.
Skelton, N.J., Aspiras, F. and Schall, T.J. (1995) Biochemistry, 34, 5329–5342.
Wall, L. and Schwartz, R.L. (1990) Programming perl, O'Reilly and Associates, Inc.
Wüthrich, K. (1986) NMR of Proteins and Nucleic Acids, Wiley, New York, NY.
Xu, J. and Sanctuary, B.C. (1993) J. Chem. Inf. Comput. Sci., 33, 490–500.
Xu, J., Weber, P.L. and Borer, P.N. (1995) J. Biomol. NMR, 5, 183–192.
Zimmermann, D., Kulikowski, C., Wang, L., Lyons, B. and Montelione, G.T. (1994) J. Biomol. NMR, 4, 241–256.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Gronwald, W., Willard, L., Jellard, T. et al. CAMRA: Chemical shift based computer aided protein NMR assignments. J Biomol NMR 12, 395–405 (1998). https://doi.org/10.1023/A:1008321629308
Issue Date:
DOI: https://doi.org/10.1023/A:1008321629308