Abstract
Z-DNA is a high energy conformer of B-DNA that forms in vivo during transcription as a result of torsional strain generated by a moving polymerase. An understanding of the biological role of Z-DNA has advanced with the discovery that the RNA editing enzyme double-stranded RNA adenosine deaminase type I (ADAR1) has motifs specific for the Z-DNA conformation. Editing by ADAR1 requires a double-stranded RNA substrate. In the cases known, the substrate is formed by folding an intron back onto the exon that is targeted for modification. The use of introns to direct processing of exons requires that editing occurs before splicing. Recognition of Z-DNA by ADAR1 may allow editing of nascent transcripts to be initiated immediately after transcription, ensuring that editing and splicing are performed in the correct sequence. Structural characterization of the Z-DNA binding domain indicates that it belongs to the winged helix-turn-helix class of proteins and is similar to the globular domain of histone-H5.
Article PDF
Similar content being viewed by others
References
Rich A.: DNA comes in many forms, Gene 135 (1993): 99–109.
Wang A.H.-J., Quigley G.J., Kolpak F.J., Crawford J.I., van Boom J.H., van der Marel G. and Rich A.: Molecular structure of a left-handed double helical DNA fragment at atomic resolution, Nature 282 (1979) 680–686.
Klysik J., Stirdivant S.M., Larson J.E., Hart P.A. and Wells R.D.: Left-handed DNA in restriction fragments and a recombinant plasmid, Nature 290 (1981): 672–677.
Haniford D.B. and Pulleybank D.E.: Facile transition of poly[d(TG).d(CA)] into a left-handed helix in physiological conditions, Nature 302 (1983): 632–634.
Peck L.J., Nordheim A., Rich A. and Wang J.C.: Flipping of cloned d(pCpG)n.d(pCpG)n DNA sequences from right-to left-handed helical structure by salt, Co(III), or negative supercoiling, Proc. Natl. Acad. Sci. USA 79 (1982): 4560–4564.
Pohl F.M. and Jovin T.M.: Salt induced co-operative conformational change of a synthetic DNA: Equilibrium and kinetic studies with poly(dG-dC), J. Mol. Biol. 67 (1972): 375–396.
Thamann T.J., Lord R.C., Wang A.H.-J and Rich A.: The high salt form of poly(dG-dC).ploy(dG-dC) is left-handed Z-DNA: Raman spectra of crystals and solutions, Nucleic Acids Res. 9 (1981): 5443–5457.
Behe, M. and Felsenfeld G.: Effects of methylation on a synthetic polynucleotide: The B–Z transition in poly(dG-m5dC)-poly(dG-m5dC), Proc. Natl. Acad. Sci. USA 78 (1981): 1619–1623.
Singleton C.K., Klysik J., Stirdivant S.M. and Wells R.D.: Left-handed Z-DNA is induced by supercoiling in physiological ionic conditions, Nature 299 (1982): 312–316.
Peck L. and Wang J.C.: Energetics of B-to-Z transition in DNA, Proc. Natl. Acad. Sci. USA 80 (1983): 6206–6210.
Ellison M.J., Kelleher III R.J, Wang A.H.-J., Habener J.F. and Rich A.: Sequence-dependent energetics of the B–Z transition in supercoiled DNA containing nonalternating purine-pyrimidine sequences, Proc. Natl. Acad. Sci. USA 82 (1985): 8320–8324.
McLean M.J., Blaho J.A., Kilpatrick M.W. and Wells R.D.: Consecutive A–T pairs can adopt a left-handed DNA structure, Proc. Natl. Acad. Sci. USA 83 (1986): 5884–5888.
Zacharias W., O'Connor T.R. and Larson J.E.: Methylation of cytosine in the 5-position alters the structural and energetic properties of the supercoil-induced Z-helix and of B–Z junctions, Biochemistry 27 (1988): 2970–2978.
Ho P.S., Ellison M.J., Quigley G.J. and Rich A.: A computer aided thermodynamic approach for predicting the formation of Z-DNA in naturally occurring sequences, EMBO J. 5 (1986): 2737–2744.
Ellison M.J., Feigon J., Kelleher III R.J., Wang A.H.-J., Habener J.F. and Rich A.: An assessment of the Z-DNA forming potential of alternating dA–dT stretches in supercoiled plasmids, Biochemistry 25 (1986): 3648–3655.
Liu L.F. and Wang J.C.: Supercoiling of the DNA template during transcription, Proc. Natl. Acad. Sci. USA 84(20) (1987): 7024–7027.
Schroth G.P., Chou P.J. and Ho P.S.: Mapping Z-DNA in the human genome. Computer-aided mapping reveals a nonrandom distribution of Z-DNA-forming sequences in human genes, J. Biol. Chem. 267 (1992): 11846–11855.
Jiang H., Zacharias W. and Amirhaeri S.: Potassium permanganate as an in situ probe for B–Z and Z–Z junctions, Nucleic Acids Res. 19 (1991): 6943–6948.
Palacek E., Rasvoka E. and Boublikova P.: Probing of DNA polymorphic structure in the cell with osmium tetraoxide, Biochem. Biophys. Res. Commun. 150 (1988): 731–738.
Zheng G., Kochel T., Hoepfner R.W., Timmons S.E. and Sinden R.R.: Torsionally tuned cruciform and Z-DNA probes for measuring unrestrained supercoiling at specific sites in DNA of living cells, J. Mol. Biol. 221 (1991): 107–129.
Jaworski A., Hsieh W.-T., Blaho J.A. and Larson J.E.: Left-handed DNA in vivo, Science 238 (1987): 773–777.
Rahmouni A.R. and Wells R.D.: Stabilization of Z-DNA in vivo by localized supercoiling, Science 246 (1989): 358–363.
Jaworski A., Higgins N.P., Wells R.D. and Zacharias W.: Topoisomerase mutants and physiological conditions control supercoiling and Z-DNA formation in vivo, J. Biol. Chem. 266 (1991): 2576–2581.
Lafer E.M., Valle R.P.C., Moller A., Nordheim A., Schur P.H., Rich A. and Stollar B.D.: Z-DNA-specific antibodies in Human systemic lupus erythematosis, J. Clin. Invest. 71 (1983): 314–321.
Lafer E.M., Sousa R., Ali R., Rich A. and Stollar B.D.: The effect of anti-Z-DNA antibodies on the B-DNA-Z-DNA equilibrium, J. Biol. Chem. 261 (1986): 6438–6443.
Nordheim A., Pardue M.L., Lafer E.M., Moller A., Stollar B.D. and Rich A.: Antibodies to left handed Z-DNA bind to interband regions of Drosophila polytene chromosomes, Nature 294 (1981): 417–422.
Lancillotti F., Lopez M.C., Arias P. and Alonco C.: Z-DNA in transcriptionally active chromosomes, Proc. Natl. Acad. Sci. USA 84 (1987): 1560–1564.
Hill R.J.: Z-DNA; A prodrome for the 1990, J. Cell Sci. 99 (1991): 675–680.
Lipps H.J., Nordheim A., Lafer E.M., Ammermann D., Stollar B.D. and Rich A.: Antibodies against Z-DNA react with the macronucleus but not the micronucleus of the hypotrichous ciliate Stylonychia mytilus, Cell 32 (1983): 435–441.
Jackson D.A. and Cook P.R.: A general method for preparing chromatin containing intact DNA, EMBO J. 4 (1985): 913–918.
Jackson D.A., Yuan J. and Cook P.R.: A gentle method for preparing cyto-and nucleo-skeletons and associated chromatin, J. Cell Sci. 90 (1988): 365–378.
Wittig B., Dorbic T. and Rich A.: The level of Z-DNA in the metabolically active, permeabilized mammalian cell nuclei is regulated by torsional strain, J. Cell Biol. 108 (1989): 755–764.
Wittig B., Dorbic T. and Rich A.: Transcription is associated with Z-DNA formation in metabolically active permeabilized mammalian cell nuclei, Proc. Natl. Acad. Sci. USA 88 (1991): 2259–2263.
Wittig B., Wolfl S., Dorbic T., Vahrson W. and Rich A.: Transcription of human c-myc in permeabilized nuclei is associated with formation of Z-DNA in three discrete regions of the gene, EMBO J. 11 (1992): 4653–4663.
Wolfl S., Wittig B. and Rich A.: Identification of transcriptionally induced Z-DNA segments in the human c-myc gene, Biochim. Biophys. Acta 1264 (1995): 294–302.
Wolfl S., Martinez C., Rich A. and Majzoub J.A.: Transcription of the human corticotropin-releasing hormone gene in NPLC cells is correlated with Z-DNA formation, Proc. Natl. Acad. Sci. USA 93 (1996).
Peck L.J. and Wang J.C.: Transcriptional block caused by a negative supercoiling induced structural change in an alternating CG sequence, Cell 40 (1985): 129–137.
Pohl F.M.: Ein Modell der DNS-struktur. Naturwis-senschaften, 1967. 54: p. 616.
Treco D. and Arnheim N.: The evolutionary conserved repetitive sequence d(TG.AC)n promotes reciprocal exchange and generates unusual recombinant tetrads during yeast meiosis, Mol. Cell. Biol. 6 (1986): 3934–3947.
Bullock P., Miller J. and Botchan M.: Effects of poly[d(pGpT).d(pApC)] and poly[d(pCpG).d(pCpG)] repeats on homologous recombination in somatic cells, Mol. Cell. Biol. 6(11) (1986): 3948–3953.
Wahls W.P., Wallace L.J. and Moore P.D.: The Z-DNA motif d(TG)30 promotes reception of information during gene conversion while stimulating homologous recombination in human cells in culture, Mol. Cell. Biol. 10(2) (1990): 785–793.
Aplan P.D., Raimondi S.C. and Kirsch I.R.: Disruption of the SCL gene by at(1;3) translocation in a patient with T cell acute lymphoblastic leukemia, EMBO. J. 8(9) (1989): 2621–2631.
Boehm T., Mengle-Gaw L., Kees U.R., Spurr N., Lavenir I., Forster A. and Rabbitts T.H.: Alternating purine-pyrimidine tracts may promote chromosomal translocations seen in a variety of human lymphoid tumours, EMBO J. 8(9) (1989): 2621–2631.
Satyanarayana K. and Strominger J.L.: DNA sequences near a meiotic recombinational breakpoint within the human HLA-DQ region, Immunogenetics 35(4) (1992): 235–240.
Steinmetz M., Stephan D. and Lindahl K.F.: Gene organization and recombinational hotspots in the murine major histocompatibility complex, Cell 44 (1986): 895–904.
Weinreb A., Katzenberg D.R., Gilmore G.L. and Birshtein B.K.: Site of unequal sister chromatid exchange contains a potential Z-DNA forming tract, Proc. Natl. Acad. Sci. USA 85(2) (1991): 529–533.
Garner M.M. and Felsenfeld G.: Effect of Z-DNA on nucleosome placement, J. Mol. Biol. 196 (1987): 581–590.
Soyer-Gobillard M.O., Geraud M.L., Coulaud D., Barray M., Theveny B., Revet B. and Delain E.: Location of Band Z-DNA in the chromosome of a primitive eukaryote dinoflagellate, J. Cell. Biol. 111(2) (1990): 293–304.
Wolfl S., Vahrson W. and Herbert A.G.: Analysis of left handed Z-DNA in vivo. In: Saluz H.P. and Wiebauer K. (eds), DNA and Nucleoprotein Structure in vivo. Saluz H.P. and Wiebauer K. (ed.) Landes Co.: Austin, TX. 1995, p. 137–159.
Krishna P., Kennedy B.P., Waisman D.M., van de Sande J.H. and McGhee J.D.: Are many Z-DNA binding proteins actually phospholipid-binding proteins? Proc. Natl. Acad. Sci. USA 87 (1990): 1292–1295.
Rohner K.J., Hobi R. and Kuenzle C.C.: Z-DNA-binding proteins. Identification critically depends on the proper choice of ligands, J. Biol. Chem. 265 (1990): 19112–19115.
Bass B.L., Nihikura K., Keller W., Seeburg P.H., Emeson R.B., O'Connell M.A., Samuel C.E. and Herbert A.: A standardized nomenclature for adenosine deaminases that act on RNA, RNA 3 (1997): 947–949.
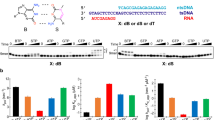
Herbert A.G. and Rich A.: A method to identify and characterize Z-DNA binding proteins using a linear oligodeoxynucleotide, Nucleic Acids Res. 21 (1993): 2669–2672.
Herbert A.G., Spitzner J.R., Lowenhaupt K. and Rich A.: Z-DNA binding protein from chicken blood nuclei, Proc. Natl. Acad. Sci. USA 90 (1993): 3339–3342.
Herbert A.G., Lowenhaupt K., Spitzner J.R. and Rich A.: Chicken double-stranded RNA adenosine deaminase has apparent specificity for Z-DNA, Proc. Natl. Acad. Sci. USA 92 (1995): 7550–7554.
Herbert A., Alfken J., Kim Y.-G., Mian S., Nishikura K. and Rich A.: A Z-DNA binding domain present in the human editing enzyme, double-stranded RNA adenosine deaminase, Proc. Natl. Acad. Sci, USA 94 (1997): 8421–8426.
Herbert A., Schade M., Lowenhaupt K., Alfken J., Schwartz T., Shlyakhtenko L.S., Lyubchenko Y.L. and Rich A.: The Za domain from human ADAR1 binds to the Z-DNA conformer of many different sequences, Nucleic Acids Res. 26 (1998): 3486–3493.
Schwartz T., Lowenhaupt K., Kim Y.-G., Li L., Brown II B.A., Herbert A. and Rich A.: Proteolytic dissection of Zab, the Z-DNA binding domain of human ADAR1, J. Biol. Chem. (1998): (in press).
Herbert A., Kim Y.-G., Alfken J., Nishikura K. and Rich A.: A Z-DNA binding domain from the human editing enzyme dsRNA adenosine deaminase, Proc. Natl. Acad. Sci. USA 94 (1997): 12875–12879.
Berger I., Manoharan W.W.R., Schwartz T., Alfken J., Kim Y.-G., Lowenhaupt K., Herbert A. and Rich A.: Spectroscopic characterization of Za, a novel DNA binding domain from human ADAR1, Biochemistry 37 (1998): 13313–13321.
Ramakrishnan V., Finch J.T., Graziano V., Lee P.L. and Sweet R.M.: Crystal structure of the globular domain of histone H5 and its implications for nucleosome binding, Nature 362 (1993): 219–223.
Clark K.L., Halay E.D., Lai E. and Burley S.K.: Co-crystal structure of the HNF-3/fork head DNA recognition motif resembles histone H5, Nature 364 (1993): 412–420.
Schade M., Turner C., Lowenhaupt K., Rich A. and Herbert A.: Structure/function analysis of the Z-DNA binding domain Za of ADAR1 reveals similarity to (a + b) family of helixturn-helix proteins, EMBO J. (1998): (in press).
Kim U., Wang Y., Sanford T., Zeng Y. and Nishikura K.: Molecular cloning of a cDNA for double-stranded RNA adenosine deaminase, a candidate enzyme for nuclear RNA editing, Proc. Natl. Acad. Sci. USA 91 (1994): 11457–11461.
O'Connell M., Krause S., Higuchi M., Hsuan J.J., Totty N.F., Jenny A. and Keller W.: Cloning of cDNAs encoding mammalian double-stranded RNA-specific adenosine deaminase, Mol. Cell. Biol. 15(3) (1995): 1389–1397.
Hough R.F. and Bass B.L.: Analysis of Xenopus dsRNA adenosine deaminase cDNAs reveals similarities to DNA methyltransferase, RNA 3 (1997): 356–370.
Liu Y., George C.X., Patterson J.R. and Samuel C.E.: Functionally distinct double-stranded RNA-binding domains associated with alternative splice variants of the interferon-inducible double-stranded RNA-specific adenosine deaminase, J. Biol. Chem. 14 (1997): 4419–4428.
Patterson J.B. and Samuel C.E.: Expression and regulation by interferon of a double-stranded-RNA-specific adenosine deaminase from human cells: Evidence for two forms of the deaminase, Mol. Cell. Biol. 15 (1995): 5376–5388.
Liu Y., Herbert A., Rich, A. and Samuel C.E.: Double-stranded RNA-specific adenosine deaminase nucleic acid binding properties, Methods 15 (1998): 199–205.
Bass B.L. and Weintraub H.: A developmentally regulated activity that unwinds RNA duplexes, Cell 48 (1987): 607–613.
Rebagliati M.R. and Melton D.A.: Antisense RNA injections in fertilized frog eggs reveal an RNA duplex unwinding activity, Cell 48 (1987): 599–605.
Bass B.L. and Weintraub H.: An unwinding activity that covalently modifies its double-stranded RNA substrate, Cell 55 (1988): 1089–1098.
Polson A.G., Crain P.F., Pomerantz S.C., McCloskey J.A. and Bass B.L.: The mechanism of adenosine to inosine conversion by the double-stranded RNA unwinding/modifying activity: A high performance liquid chromatography-mass spectrometry analysis, Biochemistry 30 (1991): 11507–11514.
Sommer B., Kohler M., Sprengel R. and Seeburg P.H.: RNA editing in brain controls a determinant of ion flow in glutamate-gated channels, Cell 67 (1991): 11–19.
Hume R.I., Dingledine R. and Heinemann S.F.: Identification of a site in the glutamate receptor subunits that controls calcium permeability, Science 253 (1991): 1028–1031.
Verdoorn T.A., Burnashev N., Monyer H., Seeburg P.H. and Sakmann B.: Structural determinants of ion flow through recombinant glutamate receptor channels, Science 252 (1991): 1715–1718.
Melcher T., Maas S., Sprengel R., Higuchi M. and Seeburg P.H.: RED2, a brain-specific member of the RNA-specific adenosine deaminase family, J. Biol. Chem. 271 (1996): 31795–31798.
Kohler M., Burnashev N., Sakmann B. and Seeburg P.H.: Determinants of Ca2+ permeability in both TM1 and TM2 of high affinity kainate receptor channels: Diversity by RNA editing, Neuron 10 (1993): 491–500.
Lomeli H., Mosbacher J., Melcher T., Hoger T., Geiger J.R., Kuner T., Monyer H., Higuchi M., Bach A. and Seeburg P.H.: Control of kinetic properties of AMPA receptor channels by nuclear RNA editing, Science 266 (1994): 1709–1713.
Ma J., Qian R., Rausa III F.M. and Colley K.J.: Two naturally occurring a2,6-Sialyltransferase forms with a single amino acid change in the catalytic domain differ in their catalytic activity and proteolytic processing, J. Biol. Chem. 272 (1997): 672–679.
Burns C.N., Chu H., Rueter S.M., Hutchinson L.K., Canton H., Sanders-Bush E. and Emeson R.: Regulation of serotonin-2C receptor G-protein coupling by RNA editing, Nature 387 (1997): 303–308.
Patton D.E., Silva T. and Bezanilla F.: RNA editing generates a diverse array of transcripts encoding squid Kv2 K+ channels with altered functional properties, Neuron 19 (1997): 711–722.
Paul M.L. and Bass B.L.: Inosine exists in mRNA at tissue-specific levels and is most abundant in brain mRNA, EMBO J. 16 (1998): 1120–1127.
Morse D.P. and Bass B.L.: Detection of inosine in messenger RNA by inosine specific cleavage, Biochemistry 36 (1997): 8429–8434.
Bass B.L.: RNA editing: New uses for old players in the RNA world. In: Gesteland R.F. and Atkins J.F., (eds), The RNA World. Cold Spring Harbor Laboratory Press, Plainview, NY. 1993, 383–418.
Herbert A.G.: RNA editing, introns and evolution, Trends in Genet. 12(1) (1996): 6–9.
Higuchi M., Single F.N., Kohler M., Sommer B., Sprengel R. and Seeburg P.H.: RNA editing of AMPA receptor subunit GluR-B: A base-paired intron-exon structure determines position and efficiency, Cell 75 (1993): 1361–1370.
Herb A., Higuchi M., Sprengel R. and Seeburg P.H.: Q/R site editing in kainate receptor GluR5 and GluR6 pre-mRNAs requires distant intronic sequences, Proc. Natl. Acad. Sci. USA 93 (1996): 1875–1880.
Egebjerg J., Kukekov V. and Heinemann S.F.: Intron sequence directs RNA editing of the glutamate receptor GluR2 coding sequence, Proc. Natl. Acad. Sci. USA 91 (1994): 10270–10274.
Takeuchi H., Hanamura N. and Harada I.: Structural specificity of peptides in Z-DNA formation and energetics of the peptide-induced B–Z transition of poly(dG-m5C), J. Mol. Biol. 236(2) (1994): 610–617.
Arndt-Jovin D.J., Udvardy A., Garner M.M., Ritter S. and Jovin T.: Z-DNA binding and inhibition by GTP of Drosophila Topoisomerase II, Biochemistry 32(18) (1993): 4862–4872.
Bechert T., Diekmann S. and Arndt-Jovin D.J.: Human 170 kDa and 180 kDa topoisomerases II bind preferentially to curved and left-handed linear DNA, J. Biomol. Struct. Dyn. 12(3) (1994): 605–623.
Glikin C.G., Jovin M.T. and Arndt-Jovin D.J.: Interactions of Drosophila DNA topoisomerase II with left-handed Z-DNA in supercoiled minicircles, Nucleic Acids Res. 19 (1991): 7139–7144.
Dawkins R.: The Blind Watchmaker, Harlow: Longman Scientific and Technical, 1986.
Wang J., Zeng Y., Murray J.M. and Nishikura K.: Genomic organization and chromosomal location of the human dsRNA adenosine deaminase gene: The enzyme for glutamate-activated ion channel RNA editing, J. Mol. Biol. 254 (1995): 184–195.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Herbert, A., Rich, A. Left-handed Z-DNA: structure and function. Genetica 106, 37–47 (1999). https://doi.org/10.1023/A:1003768526018
Issue Date:
DOI: https://doi.org/10.1023/A:1003768526018