Abstract
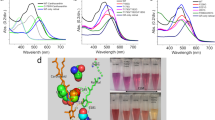
A putative xanthorhodopsin-encoding gene, XR34, was found in the genome of the moderately halophilic gammaproteobacterium Salinivibrio socompensis S34, isolated from modern stromatolites found on the shore of Laguna Socompa (3570 m), Argentina Puna. XR-encoding genes were clustered together with genes encoding X-carotene, retinal (vitamin-A aldehyde), and carotenoid biosynthesis enzymes while the carotene ketolase gene critical for the salinixanthin antenna compound was absent. To identify its functional behavior, we herein overexpressed and characterized this intriguing microbial rhodopsin. Recombinant XR34 showed all the salient features of canonical microbial rhodopsin and covalently bound retinal as a functional chromophore with λmax = 561 nm (εmax ca. 60,000 M−1 cm−1). Two canonical counterions with pK values of around 6 and 3 were identified by pH titration of the recombinant protein. With a recovery time of approximately half an hour in the dark, XR34 shows light–dark adaptation shifting the absorption maximum from 551 to 561 nm. Laser-flash induced photochemistry at pH 9 (deprotonated primary counterion) identified a photocycle starting with a K-like intermediate, followed by an M-state (λmax ca. 400 nm, deprotonated Schiff base), and a final long wavelength-absorbing N- or O-like intermediate before returning to the parental 561 nm-state. Initiating the photocycle at pH 5 (protonated counterion) yields only bathochromic intermediates, due to the lacking capacity of the counterion to accept the Schiff base proton. Illumination of the membrane-embedded protein yielded a capacitive transport current. The presence of the M-intermediate under these conditions was demonstrated by a blue light-induced shunt process.
Graphical abstract









Similar content being viewed by others
Data availability
The sequence of the xantorhodopsin protein-coding gene from Salinivibrio socompensis sp. S34 is available at GenBank Acc. AGR66236.1 together with all sequences of rhodopsin-related proteins retrieved from public databases via Uniprot (https://www.uniprot.org/blast/) and MicRhoDE (http://application.sb-roscoff.fr/micrhode/). All other raw data from results presented in the manuscript are available upon request.
References
Spudich, J. L. (2006). The multitalented microbial sensory rhodopsins. Trends in Microbiology, 14, 480–487. https://doi.org/10.1016/j.tim.2006.09.005.
Gómez-Consarnau, L., Raven, J. A., Levine, N. M., et al. (2019). Microbial rhodopsins are major contributors to the solar energy captured in the sea. Science Advances, 5, 1–7. https://doi.org/10.1126/sciadv.aaw8855.
Oesterhelt, D., & Stoeckenius, W. (1971). Rhodopsin-like protein from the purple membrane of Halobacterium halobium. Nature New Biology, 233, 149–152. https://doi.org/10.1038/newbio233149a0.
Bamann, C., Bamberg, E., Wachtveitl, J., & Glaubitz, C. (2014). Proteorhodopsin. Biochimica et Biophysica Acta (BBA) Bioenergetics, 1837, 614–625. https://doi.org/10.1016/j.bbabio.2013.09.010.
Balashov, S. P., Imasheva, E. S., Boichenko, V. A., et al. (2005). Xanthorhodopsin: A proton pump with a light-harvesting carotenoid antenna. Science (80-), 309, 2061–2064. https://doi.org/10.1126/science.1118046.
Imasheva, E. S., Balashov, S. P., Choi, A. R., et al. (2009). Reconstitution of gloeobacter violaceus rhodopsin with a light-harvesting carotenoid antenna. Biochemistry, 48, 10948–10955. https://doi.org/10.1021/bi901552x.
Balashov, S. P., Imasheva, E. S., Choi, A. R., et al. (2010). Reconstitution of gloeobacter rhodopsin with echinenone: Role of the 4-Keto group. Biochemistry, 49, 9792–9799. https://doi.org/10.1021/bi1014166.
Vollmers, J., Voget, S., Dietrich, S., et al. (2013). Poles apart: Arctic and Antarctic octadecabacter strains share high genome plasticity and a new type of xanthorhodopsin. PLoS ONE, 8, e63422.
Mongodin, E. F., Nelson, K. E., Daugherty, S., et al. (2005). The genome of Salinibacter ruber: Convergence and gene exchange among hyperhalophilic bacteria and archaea. Proceedings of the National Academy of Sciences United States of America, 102, 18147–18152. https://doi.org/10.1073/pnas.0509073102.
Albarracín, V. H., Kurth, D., Ordoñez, O. F., et al. (2015). High-up: A remote reservoir of microbial extremophiles in Central Andean wetlands. Frontiers in Microbiology, 6, 1404. https://doi.org/10.3389/fmicb.2015.01404.
Albarracín, V. H., Gärtner, W., & Farias, M. E. (2016). Forged under the sun: Life and art of extremophiles from Andean Lakes. Photochemistry and Photobiology, 92, 14–28. https://doi.org/10.1111/php.12555.
Kurth, D., Belfiore, C., Gorriti, M. F., et al. (2015). Genomic and proteomic evidences unravel the UV-resistome of the poly-extremophile Acinetobacter sp. Ver3. Frontiers in Microbiology, 6, 1–18. https://doi.org/10.3389/fmicb.2015.00328.
Farías, M. E., Rascovan, N., Toneatti, D. M., et al. (2013). The discovery of stromatolites developing at 3570 m above sea level in a high-altitude Volcanic Lake Socompa, Argentinean Andes. PLoS ONE 8, e53497. https://doi.org/10.1371/journal.pone.0053497.
Bequer Urbano, S., Albarracín, V. H., Ordoñez, O. F., et al. (2013). Lipid storage in high-altitude Andean Lakes extremophiles and its mobilization under stress conditions in Rhodococcus sp. A5, a UV-resistant actinobacterium. Extremophiles, 17, 217–227. https://doi.org/10.1007/s00792-012-0508-2.
Farías, M. E., Contreras, M., Rasuk, M. C., et al. (2014). Characterization of bacterial diversity associated with microbial mats, gypsum evaporites and carbonate microbialites in thalassic wetlands: Tebenquiche and La Brava, Salar de Atacama, Chile. Extremophiles, 18, 311–329. https://doi.org/10.1007/s00792-013-0617-6.
Rasuk, M. C., Kurth, D., Flores, M. R., et al. (2014). Microbial characterization of microbial ecosystems associated to evaporites domes of gypsum in Salar de Llamara in Atacama Desert. Microbial Ecology, 68, 483–494. https://doi.org/10.1007/s00248-014-0431-4.
Ordoñez, O., Lanzarotti, E., Kurth, D., et al. (2015). Genome comparison of two exiguobacterium strains from high altitude Andean Lakes with different arsenic resistance: Identification and 3D modeling of the Acr3 efflux pump. Frontiers in Environmental Science, 3, 50.
Toneatti, D. M., Albarracín, V. H., Flores, M. R., et al. (2017). Stratified bacterial diversity along physico-chemical gradients in high-altitude modern stromatolites. Frontiers in Microbiology, 8, 1–12. https://doi.org/10.3389/fmicb.2017.00646.
Alonso-Reyes, D. G., Farias, M. E., & Albarracín, V. H. (2020). Uncovering cryptochrome/photolyase gene diversity in aquatic microbiomes exposed to diverse UV-B regimes. Aquatic Microbial Ecology, 85, 141–154. https://doi.org/10.3354/AME01947.
Alonso-Reyes, D. G., Galván, S., Irazoqui, J. M., et al. (2022). Dissecting light sensing and metabolic pathways on the millimeter scale in high-altitude modern stromatolites. Microbial Ecology. https://doi.org/10.1007/s00248-022-02112-7.
Albarracín, V. H., Kraiselburd, I., Bamann, C., et al. (2016). Functional green-tuned proteorhodopsin from modern stromatolites. PLoS ONE, 11, 1–18. https://doi.org/10.1371/journal.pone.0154962.
Galisteo, C., Sánchez-Porro, C., de la Haba, R. R., et al. (2019). Characterization of Salinivibrio socompensis sp. nov., A New Halophilic Bacterium Isolated from the High-Altitude Hypersaline Lake Socompa, Argentina. Microorganisms. 7, 241. https://doi.org/10.3390/microorganisms7080241.
de la Haba, R. R., López-Hermoso, C., Sánchez-Porro, C., et al. (2019). Comparative genomics and phylogenomic analysis of the genus Salinivibrio. Frontiers in Microbiology, 10, 1–15. https://doi.org/10.3389/fmicb.2019.02104.
Gorriti, M. F., Dias, G. M., Chimetto, L. A., et al. (2014). Genomic and phenotypic attributes of novel salinivibrios from stromatolites, sediment and water from a high altitude lake. BMC Genomics, 15, 473. https://doi.org/10.1186/1471-2164-15-473.
Scharf, B., Hess, B., & Engelhard, M. (1992). Chromophore of sensory rhodopsin II from Halobacterium halobium. Biochemistry, 31, 12486–12492. https://doi.org/10.1021/bi00164a027.
Rozewicki, J., Li, S., Amada, K. M., et al. (2019). MAFFT-DASH: Integrated protein sequence and structural alignment. Nucleic Acids Research, 47, W5–W10. https://doi.org/10.1093/nar/gkz342.
Drozdetskiy, A., Cole, C., Procter, J., & Barton, G. J. (2015). JPred4: A protein secondary structure prediction server. Nucleic Acids Research, 43, W389–W394. https://doi.org/10.1093/nar/gkv332.
Stamatakis, A. (2014). RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30, 1312–1313. https://doi.org/10.1093/bioinformatics/btu033.
Pettersen, E. F., Goddard, T. D., Huang, C. C., et al. (2004). UCSF Chimera—A visualization system for exploratory research and analysis. Journal of Computational Chemistry, 25, 1605–1612. https://doi.org/10.1002/jcc.20084.
Berman, H. M., Bhat, T. N., Bourne, P. E., et al. (2000). The Protein Data Bank and the challenge of structural genomics. Natural Structural Biology, 7, 957–959. https://doi.org/10.1038/80734.
Webb, B., & Sali, A. (2016). Comparative protein structure modeling using MODELLER. Current Protocols in Bioinformatics, 54, 5.6.1-5.6.37. https://doi.org/10.1002/cpbi.3.
Morizumi, T., Ou, W. L., Van Eps, N., et al. (2019). X-ray crystallographic structure and oligomerization of gloeobacter rhodopsin. Science and Reports, 9, 1–14. https://doi.org/10.1038/s41598-019-47445-5.
Hartmut, L., Brigitte, S., Jason, S., et al. (2008). Crystallographic structure of xanthorhodopsin, the light-driven proton pump with a dual chromophore. Proceedings of the National Academy of Sciences USA, 105, 16561–16565. https://doi.org/10.1073/pnas.0807162105.
Gushchin, I., & Gordeliy, V. (2018). Microbial rhodopsins. SubCellular Biochemistry, 87, 19–56. https://doi.org/10.1007/978-981-10-7757-9_2.
Kato, H. E., Inoue, K., Abe-Yoshizumi, R., et al. (2015). Structural basis for Na+ transport mechanism by a light-driven Na+ pump. Nature, 521, 48–53. https://doi.org/10.1038/nature14322.
Tsukamoto, T., Mizutani, K., Hasegawa, T., et al. (2016). X-ray crystallographic structure of thermophilic rhodopsin: Implications for high thermal stability and optogenetil function. Journal of Biological Chemistry, 291, 12223–12232. https://doi.org/10.1074/jbc.M116.719815.
Tamogami, J., Kikukawa, T., Nara, T., et al. (2012). Photoinduced proton release in proteorhodopsin at low pH: The possibility of a decrease in the pKa of Asp227. Biochemistry, 51, 9290–9301. https://doi.org/10.1021/bi300940p.
Balashov, S. P., & Lanyi, J. K. (2007). Xanthorhodopsin: Proton pump with a carotenoid antenna. Cellular and Molecular Life Sciences, 64, 2323–2328. https://doi.org/10.1007/s00018-007-7167-y.
Balashov, S. P., Imasheva, E. S., Govindjee, R., & Ebrey, T. G. (1996). Titration of aspartate-85 in bacteriorhodopsin: What it says about chromophore isomerization and proton release. Biophysical Journal, 70, 473–481. https://doi.org/10.1016/S0006-3495(96)79591-7.
Bamberg, E., Apell, H.-J., Dencher, N. A., et al. (1979). Photocurrents generated by bacteriorhodopsin on planar bilayer membranes. Biophysics of Structure and Mechanism, 5, 277–292. https://doi.org/10.1007/BF02426663.
Karvaly, B., & Dancsházy, Z. (1977). Bacteriorhodopsin: A molecular photoelectric regulator Quenching of photovoltaic effect of bimolecular lipid membranes containing bacteriorhodopsin by blue light. FEBS Letters, 76, 36–40. https://doi.org/10.1016/0014-5793(77)80115-4.
Balashov, S. P., Petrovskaya, L. E., Lukashev, E. P., et al. (2012). Aspartate-histidine interaction in the retinal Schiff base counterion of the light-driven proton pump of Exiguobacterium sibiricum. Biochemistry, 51, 5748–5762. https://doi.org/10.1021/bi300409m.
Vignale, F. A., Lencina, A. I., Stepanenko, T. M., et al. (2022). Lithifying and non-lithifying microbial ecosystems in the Wetlands and Salt Flats of the Central Andes. Microbial Ecology, 83, 1–17. https://doi.org/10.1007/s00248-021-01725-8.
Acknowledgements
The authors acknowledge the generous financial support by PICT RAICES 2019–3216 projects. CB thanks Anja Becker for technical assistance and is grateful for the continuous support from the Max-Planck-Society. WG thanks the University of Leipzig for hosting him as a visitor. VHA is a staff researcher from the National Research Council (CONICET) in Argentina. DA and MG were the recipients of doctoral/postdoctoral fellowships from CONICET.
Author information
Authors and Affiliations
Contributions
MG, DA, PW, VHA, and CB performed the experiments. VHA, MEF, WG, and EB designed the project. All authors contributed to the generation of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare to have no competing interests regarding this manuscript.
Additional information
This paper is dedicated to Silvia Braslavsky, a pioneer in photobiology and photobiophysics, on the occasion of her 80th birthday. The authors want to congratulate Silvia on her anniversary and like to acknowledge her great contribution to the establishment of a German-Argentinean research group in the photo-microbiology of Andean extremophiles. Likewise, we are thankful for her warm hospitality in her house in Mulheim an der Ruhr, where delicious food, perfect wine, and science tertulia were always a lovely blend.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gorriti, M.F., Bamann, C., Alonso-Reyes, D.G. et al. Functional characterization of xanthorhodopsin in Salinivibrio socompensis, a novel halophile isolated from modern stromatolites. Photochem Photobiol Sci 22, 1809–1823 (2023). https://doi.org/10.1007/s43630-023-00412-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43630-023-00412-6