Abstract
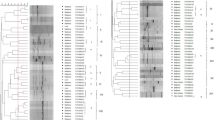
The present study was carried out to evaluate the diversity of rhizobia associated with nodules of mung bean in Pakistan, because this information is necessary for inoculum development. Based on sequence analysis of 16S rRNA gene of thirty-one bacteria, 11 were assigned to genus Bradyrhizobium, 17 to Ensifer, and 3 to Rhizobium. Phylogenetic analyses on the basis of 16S-23S ITS region, atpD, recA, nifH, and nodA of representative strains revealed that B. yuanmingense is the predominant species distributed throughout different mung bean–growing areas. Among the fast-growing rhizobia, Ensifer aridi was predominant in Faisalabad, Layyah, and Rawalpindi, while E. meliloti in Thal desert. Sequence variations and phylogeny of nifH and nodA genes suggested that these genes might have been co-evolved with the housekeeping genes and maintained by vertical gene transfer in rhizobia detected in the present study. Host infectivity assay revealed the successful nodulation of host by rhizobia related to genera Bradyrhizobium, Ensifer and Rhizobium. Among all, Bradyrhizobium and Ensifer spp. inoculation exhibited a significantly higher number of nodules (11–34 nodules plant−1) and nitrogenase activity (nodule ARA 60–110 μmol g−1 h−1). Contrary to the previous studies, our data reveal that B. yuanmingense and E. aridi are predominant species forming effective nodules in mung bean in Pakistan. Furthermore, to the best of our knowledge, this is the first report showing the effective symbiosis of E. aridi, E. meliloti, and Rhizobium pusense with mung bean. The diversity of rhizobia in different habitats revealed in the present study will contribute towards designing site-specific inocula for mung bean.





Similar content being viewed by others
References
Andrews M, Andrews ME (2017) Specificity in legume-rhizobia symbioses. Int J Mol Sci 18(4):705
Lerouge P, Roche P, Faucher C, Maillet F, Truchet G, Promé JC, Dénarié J (1990) Symbiotic host-specificity of Rhizobium meliloti is determined by a sulphated and acylated glucosamine oligosaccharide signal. Nature 344(6268):781–784
Okazaki S, Kaneko T, Sato S, Saeki K (2013) Hijacking of leguminous nodulation signaling by the rhizobial type III secretion system. Proc Natl Acad Sci 110(42):17131–17136
Jansen H, Charnnarongkul S, Kim D (1996) The economics of mungbean cultivation in Thailand. In: Asthana AN, Kim DH (eds) . Recent advances in mungbean research, Indian Society of Pulses Research and Development, pp 6–23
Min L (2001) Research advance in chemical composition and pharmacological action of mung bean. Shanghai J Trad Chin Med 5:18
Lu YL, Chen WF, Wang ET, Han LL, Zhang XX, Chen WX, Han SZ (2009) Mesorhizobium shangrilense sp. nov., isolated from root nodules of Caragana species. Int J Syst Evol Microbiol 59(12):3012–3018
Risal CP, Djedidi S, Dhakal D, Ohkama-Ohtsu N, Sekimoto H, Yokoyama T (2012) Phylogenetic diversity and symbiotic functioning in mungbean (Vigna radiata L. Wilczek) bradyrhizobia from contrast agro-ecological regions of Nepal. Syst Appl Microbiol 35(1):45–53
Yang JK, Yuan TY, Zhang WT, Zhou JC, Li YG (2008) Polyphasic characterization of mung bean (Vigna radiata L.) rhizobia from different geographical regions of China. Soil Biol Biochem 40(7):1681–1688
Zhang YF, Wang ET, Tian CF, Wang FQ, Han LL, Chen WF, Chen WX (2008) Bradyrhizobium elkanii, Bradyrhizobium yuanmingense and Bradyrhizobium japonicum are the main rhizobia associated with Vigna unguiculata and Vigna radiata in the subtropical region of China. FEMS Microbiol Lett 285(2):146–154
Appunu C, N’Zoue A, Moulin L, Depret G, Laguerre G (2009) Vigna mungo, V. radiata and V. unguiculata plants sampled in different agronomical–ecological–climatic regions of India are nodulated by Bradyrhizobium yuanmingense. Syst Appl Microbiol 32(7):460–470
Willems A, Coopman R, Gillis M (2001) Phylogenetic and DNA-DNA hybridization analyses of Bradyrhizobium species. Int J Syst Evol Microbiol 51(1):111–117
Gürtler V, Stanisich VA (1996) New approaches to typing and identification of bacteria using the 16S-23S rDNA spacer region. Microbiology 142(1):3–16
Sarr PS, Yamakawa T, Saeki Y, Guisse A (2011) Phylogenetic diversity of indigenous cowpea bradyrhizobia from soils in Japan based on sequence analysis of the 16S-23S rRNA internal transcribed spacer (ITS) region. Syst Appl Microbiol 34(4):285–292
Zinga MK, Jaiswal SK, Dakora FD (2017) Presence of diverse rhizobial communities responsible for nodulation of common bean (Phaseolus vulgaris) in South African and Mozambican soils. FEMS Microbiol Ecol 93(2)
Stępkowski T, Watkin E, McInnes A, Gurda D, Gracz J, Steenkamp ET (2012) Distinct Bradyrhizbium communities nodulate legumes native to temperate and tropical monsoon Australia. Mol Phylogenet Evol 63(2):265–277
Martens M, Dawyndt P, Coopman R, Gillis M, De Vos P, Willems A (2008) Advantages of multilocus sequence analysis for taxonomic studies: a case study using 10 housekeeping genes in the genus Ensifer (including former Sinorhizobium). Int J Syst Evol Microbiol 58(1):200–214
Ahmad M, Zahir ZA, Nazli F, Akram F, Arshad M, Khalid M (2013) Effectiveness of halo-tolerant, auxin producing Pseudomonas and Rhizobium strains to improve osmotic stress tolerance in mung bean (Vigna radiata L.). Braz J Microbiol 44(4):1341–1348
Anjum MA, Zahir ZA, Arshad M, Ashraf M (2011) Isolation and screening of rhizobia for auxin biosynthesis and growth promotion of mung bean (Vigna radiata L.) seedlings under axenic conditions. Soil Environ 30(1):18–26
Hakim S, Mirza BS, Imran A, Zaheer A, Yasmin S, Mubeen F, Mclean JE, Mirza MS (2020) Illumina sequencing of 16S rRNA tag shows disparity in rhizobial and non-rhizobial diversity associated with root nodules of mung bean (Vigna radiata L.) growing in different habitats in Pakistan. Microbiol Res 231:126356
Hakim S, Mirza BS, Zaheer A, Mclean JE, Imran A, Yasmin S, Mirza MS (2018) Retrieved 16S rRNA and nifH sequences reveal co-dominance of Bradyrhizobium and Ensifer (Sinorhizobium) strains in field-collected root nodules of the promiscuous host Vigna radiata (L.) R. Wilczek. Appl Microbiol Biotechnol 102(1):485–497
Ball D (1964) Loss-on-ignition as an estimate of organic matter and organic carbon in non-calcareous soils. J Soil Sci 15(1):84–92
Kjeldahl C (1883) A new method for the determination of nitrogen in organic matter. Z Anal Chem 22:366–382
Bray RH, Kurtz L (1945) Determination of total, organic, and available forms of phosphorus in soils. Soil Sci 59(1):39–46
Knudsen D, Peterson G, Pratt P (1982) Lithium, sodium, and potassium. In: Page A (ed) Methods of soil analysis. Part 2. Chemical and microbiological properties. Soil Science Society of America, Madison, pp 225–246
David K, Apte S, Banerji A, Thomas J (1980) Acetylene reduction assay for nitrogenase activity: gas chromatographic determination of ethylene per sample in less than one minute. Appl Environ Microbiol 39(5):1078–1080
Tak N, Bissa G, Gehlot HS (2020) Methods for isolation and characterization of nitrogen-fixing legume-nodulating bacteria. In: Gupta KJ (ed) Nitrogen metabolism in plants. Springer, New York, pp 119–143
Wilson K (1987) Preparation of genomic DNA from bacteria. In: Ausubel FM (ed) Current protocols in molecular biology. John Wiley&Sons, New York p unit 2.4
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, Thompson JD (2003) Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res 31(13):3497–3500
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173(2):697–703
Fossou RK, Ziegler D, Zézé A, Barja F, Perret X (2016) Two major clades of bradyrhizobia dominate symbiotic interactions with pigeonpea in fields of Côte d’Ivoire. Front Microbiol 7:1793
Vinuesa P, Silva C, Werner D, Martínez-Romero E (2005) Population genetics and phylogenetic inference in bacterial molecular systematics: the roles of migration and recombination in Bradyrhizobium species cohesion and delineation. Mol Phylogenet Evol 34(1):29–54
Poly F, Monrozier LJ, Bally R (2001) Improvement in the RFLP procedure for studying the diversity of nifH genes in communities of nitrogen fixers in soil. Res Microbiol 152(1):95–103
Haukka K, Lindström K, Young JPW (1998) Three phylogenetic groups of nodA and nifH genes in Sinorhizobium and Mesorhizobium isolates from leguminous trees growing in Africa and Latin America. Appl Environ Microbiol 64(2):419–426
Lowe TM, Chan PP (2016) tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res 44(W1):W54–W57
Cao Y, Wang E-T, Zhao L, Chen W-M, Wei G-H (2014) Diversity and distribution of rhizobia nodulated with Phaseolus vulgaris in two ecoregions of China. Soil Biol Biochem 78:128–137
Wang L, Cao Y, Wang ET, Qiao YJ, Jiao S, Liu ZS, Zhao L, Wei GH (2016) Biodiversity and biogeography of rhizobia associated with common bean (Phaseolus vulgaris L.) in Shaanxi Province. Syst Appl Microbiol 39(3):211–219
Zhang YM, Li Y, Chen WF, Wang ET, Tian CF, Li QQ, Zhang YZ, Sui XH, Chen WX (2011) Biodiversity and biogeography of rhizobia associated with soybean plants grown in the North China Plain. Appl Environ Microbiol 77(18):6331–6342
Li QQ, Wang ET, Zhang YZ, Zhang YM, Tian CF, Sui XH, Chen WF, Chen WX (2011) Diversity and biogeography of rhizobia isolated from root nodules of Glycine max grown in Hebei Province, China. Microb Ecol 61(4):917–931
Gehlot HS, Panwar D, Tak N, Tak A, Sankhla IS, Poonar N, Parihar R, Shekhawat NS, Kumar M, Tiwari R (2012) Nodulation of legumes from the Thar desert of India and molecular characterization of their rhizobia. Plant Soil 357(1–2):227–243
Rathi S, Tak N, Bissa G, Chouhan B, Ojha A, Adhikari D, Barik SK, Satyawada RR, Sprent JI, James EK (2018) Selection of Bradyrhizobium or Ensifer symbionts by the native Indian caesalpinioid legume Chamaecrista pumila depends on soil pH and other edaphic and climatic factors. FEMS Microbiol Ecol 94(11):fiy180
Rajendhran J, Gunasekaran P (2011) Microbial phylogeny and diversity: small subunit ribosomal RNA sequence analysis and beyond. Microbiol Res 166(2):99–110
Islam MS, Kawasaki H, Muramatsu Y, Nakagawa Y, Seki T (2008) Bradyrhizobium iriomotense sp. nov., isolated from a tumor-like root of the legume Entada koshunensis from Iriomote Island in Japan. Biosci Biotechnol Biochem 72(6):1416–1429
Delamuta JRM, Ribeiro RA, Menna P, Bangel EV, Hungria M (2012) Multilocus sequence analysis (MLSA) of Bradyrhizobium strains: revealing high diversity of tropical diazotrophic symbiotic bacteria. Braz J Microbiol 43(2):698–710
Nzoué A, Miché L, Klonowska A, Laguerre G, De Lajudie P, Moulin L (2009) Multilocus sequence analysis of bradyrhizobia isolated from Aeschynomene species in Senegal. Syst Appl Microbiol 32(6):400–412
Doignon-Bourcier F, Willems A, Coopman R, Laguerre G, Gillis M, de Lajudie P (2000) Genotypic characterization of Bradyrhizobium strains nodulating small Senegalese legumes by 16S-23S rRNA intergenic gene spacers and amplified fragment length polymorphism fingerprint analyses. Appl Environ Microbiol 66(9):3987–3997
Willems A, Munive A, de Lajudie P, Gillis M (2003) In most Bradyrhizobium groups sequence comparison of 16S-23S rDNA internal transcribed spacer regions corroborates DNA-DNA hybridizations. Syst Appl Microbiol 26(2):203–210
Zhang S, Xie F, Yang J, Li Y (2011) Phylogeny of bradyrhizobia from Chinese cowpea miscellany inferred from 16S rRNA, atpD, glnII, and 16S–23S intergenic spacer sequences. Can J Microbiol 57(4):316–327
Waleron M, Waleron K, Kamasa J, Przewodowski W, Lojkowska E (2011) Polymorphism analysis of housekeeping genes for identification and differentiation of Clavibacter michiganensis subspecies. Eur J Plant Pathol 131(2):341–354
Suerbaum S (2000) Genetic variability within Helicobacter pylori. Int J Med Microbiol 290(2):175–181
Weir B (2006) Systematics, specificity, and ecology of New Zealand Rhizobia. University of Auckland Auckland, New Zealand
Ojha A, Tak N, Rathi S, Chouhan B, Rao SR, Barik SK, Joshi SR, Sprent JS, James EK, Gehlot HS (2017) Molecular characterization of novel Bradyrhizobium strains nodulating Eriosema chinense and Flemingia vestita, important unexplored native legumes of the sub-Himalayan region (Meghalaya) of India. Syst Appl Microbiol 40(6):334–344
Panday D, Schumann P, Das SK (2011) Rhizobium pusense sp. nov., isolated from the rhizosphere of chickpea (Cicer arietinum L.). Int J Syst Evol Microbiol 61(11):2632–2639
Andrews M, De Meyer S, James E, Stępkowski T, Hodge S, Simon M, Young J (2018) Horizontal transfer of symbiosis genes within and between rhizobial genera: occurrence and importance. Genes 9(7):321
Chang YL, Wang ET, Sui XH, Zhang XX, Chen WX (2011) Molecular diversity and phylogeny of rhizobia associated with Lablab purpureus (Linn.) grown in Southern China. Syst Appl Microbiol 34(4):276–284
Le Quéré A, Tak N, Gehlot HS, Lavire C, Meyer T, Chapulliot D, Rathi S, Sakrouhi I, Rocha G, Rohmer M (2017) Genomic characterization of Ensifer aridi, a proposed new species of nitrogen-fixing rhizobium recovered from Asian, African and American deserts. BMC Genomics 18(1):85
Tak N, Awasthi E, Bissa G, Meghwal RR, James EK, Sprent JS, Gehlot HS (2016) Multi locus sequence analysis and symbiotic characterization of novel Ensifer strains nodulating Tephrosia spp. in the Indian Thar Desert. Syst Appl Microbiol 39(8):534–545
Sakrouhi I, Belfquih M, Sbabou L, Moulin P, Bena G, Filali-Maltouf A, Le Quéré A (2016) Recovery of symbiotic nitrogen fixing acacia rhizobia from Merzouga Desert sand dunes in South East Morocco–identification of a probable new species of Ensifer adapted to stressed environments. Syst Appl Microbiol 39(2):122–131
Missbah El Idrissi M, Lamin H, Alami S, Bouhnik O, ElFaik S, Abdelmoumen H, Bedmar EJ (2019) Nodulation of Retama monosperma by Ensifer aridi in an abandonned lead mine soils in eastern Morocco. Front Microbiol 10:1456
Mnasri B, Badri Y, Saïdi S, de Lajudie P, Mhamdi R (2009) Symbiotic diversity of Ensifer meliloti strains recovered from various legume species in Tunisia. Syst Appl Microbiol 32(8):583–592
Sankhla IS, Tak N, Meghwal RR, Choudhary S, Tak A, Rathi S, Sprent JI, James EK, Gehlot HS (2017) Molecular characterization of nitrogen fixing microsymbionts from root nodules of Vachellia (Acacia) jacquemontii, a native legume from the Thar Desert of India. Plant Soil 410(1–2):21–40
Funding
The study was financially supported by HEC through Ph.D. fellowship to Sughra Hakim.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible Editor: Ieda Carvalho Mendes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 1268 kb)
Rights and permissions
About this article
Cite this article
Hakim, S., Imran, A. & Mirza, M.S. Phylogenetic diversity analysis reveals Bradyrhizobium yuanmingense and Ensifer aridi as major symbionts of mung bean (Vigna radiata L.) in Pakistan. Braz J Microbiol 52, 311–324 (2021). https://doi.org/10.1007/s42770-020-00397-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-020-00397-9