Abstract
The first corona-pandemic, coronavirus disease 2019 (COVID-19) caused a huge health crisis and incalculable damage worldwide. Knowledge of how to cure the disease is urgently needed. Emerging immune escaping mutants of the virus suggested that it may be potentially persistent in human society as a regular health threat as the flu virus. Therefore, it is imperative to identify appropriate biomarkers to indicate pathological and physiological states, and more importantly, clinic outcomes. Proteins are the performers of life functions, and their abundance and modification status can directly reflect the immune status. Protein glycosylation serves a great impact in modulating protein function. The use of both unmodified and glycosylated proteins as biomarkers has also been proved feasible in the studies of SARS, Zika virus, influenza, etc. In recent years, mass spectrometry-based glycoproteomics, as well as proteomics approaches, advanced significantly due to the evolution of mass spectrometry. We focus on the current development of the mass spectrometry-based strategy for COVID-19 biomarkers’ investigation. Potential application of glycoproteomics approaches and challenges in biomarkers identification are also discussed.
Similar content being viewed by others
1 Introduction
In 2019, an unknown serious acute respiratory disease broke out. The coronavirus responsible was soon isolated and sequenced by Chinese scientists [1,2,3], and was named SARS-CoV-2 as its highly homological sequence to SARS virus by World Health Organization [4, 5]. The disease caused, COVID-19, became the first corona pandemic in human history and caused tremendous losses worldwide, with over 161 million infected cases and 3.35 million deaths by May 15, 2021 [6].
SARS-CoV-2 primarily transmits through the droplets to respiratory systems, although other dissemination paths and system injuries are reported as well [7,8,9]. As a typical β coronavirus as SARS and MERS, it binds to angiotensin-converting enzyme II (ACE2) in host cells, conducting penetration, biosynthesis, maturation in several organs [10,11,12,13]. It causes a multitude of symptoms mainly including dry cough, fever, etc. [14] Besides lower respiratory tract infection, other symptoms including diarrhea, headache, etc., have also been observed [15, 16]. Some symptom-free coronavirus carriers remain symptom-free after 14 days of observation, but still carry around substantial amounts of the virus [17, 18]. Moreover, with the emerging immune escaping viral strains, COVID-19 might become a common health threat to humans like the seasonal flu [19,20,21]. It is therefore urgent to comprehend the association between molecular fingerprints and clinical outcomes [22,23,24].
Biomarkers are bio-molecules that correlated to particular biological or disease states [25]. Viruses are known to affect the host proteome to achieve optimal growth [26]. The formation of protein interaction between a virus and the host is the determinant factor for the host adaptation process of the virus [27, 28]. More importantly, the abundance and modification status of immune response proteins will directly reflect the immune status of the host [29, 30]. Due to the crucial roles of proteins in the biological and pathological process, protein biomarkers have been identified in Zika, Ebola, swine flu, etc. [31,32,33] Meanwhile, glycosylation arguably serves as one of the top factors in modulating protein location, binding affinity, activity, and fate [34]. Glycosylation also serves critical roles in the life cycle of the virus, including protein folding, receptor binding, adjusting degradation rates, controlling tropisms, and shielding immunogenic epitopes from the immune system [35]. Virus-induced diseases significantly affect the glycosylation status in the host, impacting the abundance and species of glycans, increasing or decreasing their expression, or even leaving the glycan structure incomplete [36, 37]. The level and species of glycans of protein glycosylation are closely related to the chemical status in the cellular microenvironment, particularly sugar levels and species that might be changed by disease, thus having long served as a source of disease biomarkers [38].
Around 20% of COVID-19 cases are reported to develop serious symptoms and even lead to death [39]. Although age, gender, pre-existing diseases, and unhealthy lifestyle are reported to be significant factors in influencing the disease severity of COVID-19 patients, they could not be used to forecast prognostic outcomes directly [40]. Based on extensive studies on COVID-19 pathogenesis and clinic features, it is now clear that the disease progression is closely related to the host’s immune response [41,42,43,44]. Pathogen infection is fundamentally manipulated by eliciting host metabolism: viruses adapt to the host metabolic environment and begin to replicate, thrive, and continue to invade neighboring cells. Therefore, understanding the pathogenic role of viruses using host metabolism is a possible future therapy to stop and defeat viruses [45, 46]. A thorough and detailed analysis of the protein, as well as the metabolome status of COVID-19 patients, should reveal important knowledge of host immune responses to infection, shedding light on their potential links with the clinical outcome of the patients [47].
In another aspect, mass spectrometry (MS)-based approaches, particularly MS-based proteomics and glycoproteomic analysis, are widely applied to explore host response to the pathogen, including infection, invasion, persistence, and pathogenesis, and can initially guide diagnosis to prevent the disease from turning into severity [48, 49]. A unique strength of MS-based strategies lies in the fact that it reveals a non-biased profile of protein/metabolite status of patients in a high-throughput manner [50]. By comparing longitudinal or cross-sectional molecular profiles of COVID-19 patients, extensive studies have been endeavored to search for biomarkers based on MS analysis [22, 23, 51,52,53,54]. Herein, we will focus on the current progress and knowledge obtained of COVID-19 protein biomarkers, primarily identified by MS-based proteomics approaches. We will also discuss the potential use of glycoproteomics in searching for COVID-19 clinic biomarkers and potential challenges in MS-based protein biomarker investigation.
2 MS-based Proteomics
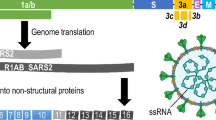
It is well-recognized since the early stage in the biological study that proteins play a fundamental role in life, encompassing metabolism, signaling generation, transduction and regulation, immune response, molecule transportation, structural organization, etc. [55,56,57,58,59]. The proteome is defined as whole proteins translated from the genome [60]. As the systemic study of diverse properties of the proteome [61], proteomics analysis provides a global figure of biological process at the level of protein, including protein identity and abundance, modification status (i.e., post-translational modification, PTM) as well as interactions between each other, thereby indicating the current physiologic and pathological state [62].
Over the past twenty or so years, proteomics has developed a multitude of methods to discover disease biomarkers due to the rapid evolution of MS-based techniques, such as matrix-assisted laser desorption/ionization (MALDI), electrospray ionization (ESI), Orbitrap, Fourier Transform-MS, etc. [62]. Normally, MALDI-MS is applied in relatively simple peptides analysis, while LC–MS/MS is preferred to analyze multi-peptide mixtures. Two overall strategies are commonly applied to identify/characterize proteins in a particular sample: bottom–up or top–down approaches. Top–down proteomics introduces the intact proteins into MS for the analysis of both intact proteins and their fragments, which allows integrated sequence coverage and full characterization of proteoforms but presents defects in sensitivity and throughput [63, 64]. While the most popular strategy to determine protein profiles is the shotgun proteomics attributed to the bottom–up approach (Fig. 1). Shotgun proteomics first uses enzymes for protein mixtures digestion and then the separation of peptides by LC. After that, the peptides were subjected to primary MS to obtain the mass–charge ratio as well as the corresponding signal intensity. Subsequently, a specific peptide ion is delivered for fragmentation to obtain its MS/MS spectrum [65]. The different choices of peptide fragmentation include higher-energy collisional dissociation (HCD), collision-induced dissociation (CID), or electron-transfer dissociation (ETD) which are well covered in other reviews and hence will not be discussed here [66, 67]. What is worth mentioning is that ETD performs the ionization by transferring electrons to a multiply protonated peptide so is widely used to identify PTMs on the peptide, and in certain cases will be combined with HCD to identify modifications [68]. The obtained raw data will then be analyzed to acquire peptide sequence and quantitation in software, such as MaxQuant, Mascot, etc.
A workflow for COVID-19 protein biomarkers identification. The workflow for COVID-19 protein biomarker identification using mass spectrometry (MS)-based proteomics. SARS-CoV-2 infects patients with a different precondition, causing different degrees of symptoms. Urine, plasma or serum, and pharyngeal swab samples were collected from mild and severe patients and healthy controls. As an example, blood samples were subjected to several proteomic processes, including denaturation, thiol-alkylation, digestion, and other steps. MS spectra were compared and then subjected to multivariate statistical analysis, which can analyze protein biomarkers for COVID-19 severity with different levels
The quantitative proteomics methods could be divided into relative and absolute quantification [69]. The core of relative quantification is the comparison of differences, i.e., the quantitative analysis of two or more samples under different physiological and pathological conditions by MS on a large scale and high throughput to obtain precise differences in protein expression, mainly utilizing stable isotope labeling and non-labeling techniques [70, 71]. Absolute quantification is achieved by obtaining the specific amount of protein expression, using MS to monitor the unique peptide of the target protein to obtain the peak area, and comparing it with the known amount of standard peptide (external labeling) or stable isotope-labeled peptide (internal labeling) to determine the specific amount [72, 73]. In contrast, relative quantitation shows the relative fold of change to a common quantitation base and is widely used to indicate the up- or down-regulation of proteins in response to stimuli.
However, when it comes to the choice of quantitation methods, a more important question is whether we want to focus on a particular group of limited candidates (targeted quantitation) or use an unbiased approach to explore as many candidates as we could (non-targeted quantitation, Fig. 2). Traditionally, people tend to follow an “unbiased—biased—validation” route for biomarker discovery [74]. Because the unbiased (non-targeted) quantitation in combination with the profiling proteomics approach can provide a whole picture, and the significantly changed candidates could be further explored in a larger group of clinic samples by the biased (targeted) quantitation method. In the case of COVID-19 biomarkers discovery, given a race with the virus is still ongoing, omitting one step to accelerate the process is reasonable. A variety of different quantitation approaches are then available for choosing, depending on which way we go for: targeted or non-targeted.
A general quantitation method for biomarkers identification. Quantitative approaches for biomarker discovery could be generally divided into non-targeted and targeted methods. Non-targeted proteomics allows a systematic and comprehensive analysis of proteins in samples and is generally used as the first step for biomarker discovery. Depending on whether labeling reagent was used, non-targeted proteomics can be further divided into two categories. The widely used labeling are isobaric tags for relative and absolute quantitation (iTRAQ) and tandem mass tag (TMT), while the data-independent acquisition (DIA) and spectral counting are common methods for quantitation in a labeling free manner. Targeted quantitation, in contrast, is a biased strategy that focused on a small set of biomarker candidates. Common methods of targeted quantitation include well-established methods like multiple reaction monitoring (MRM), selected reaction monitoring (SRM), and newly emerging parallel reaction monitoring (PRM), which provide full MS2 spectrum to confirm the identity of the target in addition to quantitation information
Stable isotope labeling is a classical method and probably still the most common approach for non-target quantitation, involving use of isobaric tags for relative and absolute quantitation (iTRAQ) and tandem mass tag (TMT) [75, 76]. A detailed workflow is shown in Fig. 3 using iTRAQ as an example. In contrast, the label-free method like spectral counting provides an alternative for quantitation. However, all three methods belong to the data-dependent acquisition (DDA) method, where precursor ions are chosen based on signal intensity for MS2 analysis. DDA method is known to be prone to the loss of low-abundance peptides and has certain randomness and an uneven number of scans [77, 78]. The recent development of data-independent acquisition (DIA), from a different aspect, divides the whole scanning range into several windows, each window is selected and fragmented in turn, and all the daughter ions of all parent ions within the window are collected [79]. DIA is subjected to neither specific target peptide nor upper limit of flux and has a uniform number of scanning points. Therefore, DIA can achieve qualitative confirmation and quantitative ion screening, which has advantages over traditional DDA [80]. For targeted quantitation, multiple reaction monitoring (MRM) serves as the gold standard in quantitation for many biomarkers, while newly emerged parallel reaction monitoring (PRM) provides MS2 spectrum in addition to quantitation information, therefore increases the reliability as well as specificity of quantitation [81, 82].
A workflow for iTRAQ-based protein quantification. iTRAQ-based protein quantification (in the case of 4-plex) is based on qualitative analysis, i.e., after protein extraction, thiol-alkylation and digestion, the resulting peptides are labeled and mixed with iTRAQ reagent, followed by liquid chromatography separation and analysis using tandem mass spectrometry. A database search of the peptides fragments allows the identification of the labeled peptides and thus the corresponding proteins. The reporter ions generated by the fragments can be used to quantify the peptides and the proteins from which they originate
Body fluids are considered to be the most promising material for the search of disease biomarkers because of their direct association with various tissues, where proteins are found to be secreted or emitted [83]. Back in 2006, Mann et al. constructed the first protein map of body fluids originated from human beings using high-precision, high-resolution MS for large-scale analysis of a variety of body fluids [84,85,86]. In 2014, Kim et al. presented two large-scale human proteome map drafts using high-resolution Fourier transform MS for body fluids (including serum, saliva, urine, etc.) [87]. Among all kinds of body fluids, plasma is the most widely used material for biomarker profiling, largely because it influenced other body fluids to a certain extent, and it can be obtained by simple, non-invasive methods [88, 89]. Plasma is extremely complex, and dynamic ranges of plasma protein are the highest among all body fluids. It is estimated to be over 10 orders of magnitude, and high abundance proteins need to be masked to reduce background complexity before MS analysis [89]. Commonly used methods for processing plasma samples include antibody-depletion followed by fractionation, sometimes target proteins need to be isolated specifically [90].
Gordon et al. used plasma samples to systematically investigate the host interacting factors of SARS-CoV-2 while explored potential pharmacological compounds inhibiting viral replication [91]. For further apprehension of the interaction profile of proteins from both SARS-CoV-2 and host cells, they labeled and expressed 26 viral proteins. Because interaction profiles require the use of immunoprecipitation-based purification techniques, affinity purification MS was applied to identify host proteins that directly interacted with each target, resulting in the identification of over 300 protein–protein interactions. Ultimately it leads to the discovery of potential antiviral drugs, which may spawn a therapeutic regimen to conquer COVID-19. A similar investment conducted by Bojkova et al. [51] encourages the development of translation inhibitors to prevent virus replication. Several phosphoproteomics studies were also conducted by different research groups to identify key kinases in response to infection [52,53,54]. All these studies provide clues for biomarker identification in patient samples.
We searched the entire literature included in Web of Science using COVID-19, proteomics, and biomarker as keywords (Table 1). Except for the study by Wallentin et al. [92] which could obtain a great number of samples directly from the hospital because of support from the ARISTOTLE trial, the majority of the studies used plasma as the sample for the study, and the cohort size was no more than 50 cases. The protein biomarkers uncovered were mainly related to inflammation caused by the disease. For example, Shen et al. worked on selecting potential blood biomarkers for COVID-19 severity assessment [23]. They performed proteomic profiling of serum of 46 COVID-19 and 53 healthy subjects, each sampled at no more than 2 time-points. To obtain higher relative protein quantification accuracy, the study employed a stable isotope-labeled proteomics strategy (TMT) cooperating with orbitrap machine in DDA mode. A total of 93 differential expressed proteins in severe patient sera were finally identified. The study by Shu et al. focused on the host response to COVID-19 pathophysiology as well [22]. To investigate the immune response of distinct clinic outcomes and thus uncover protein markers associated with disease progression and tissue-specific protein alterations, this study performed proteomic analysis of 22 clinically diagnosed COVID-19 patients versus 8 healthy controls with up to 4 time-points each sample using TMT assisted LC–ESI–MS/MS in DDA mode. Validation by machine learning and enzyme-linked immunosorbent assay (ELISA) ultimately identified several altered plasma features. Intriguingly, the plasma proteins identified with significant alterations also included CRP besides acute-phase proteins (APPs), cholesteryl ester transfer protein (CETP), peptidase inhibitor 16 (PI16), etc. The results share commons with respective studies by Shen et al. [23] and by Gordon et al. [91], which implies a pathological association with inflammation and predicts proteinase receptors as pharmacological targets. Although the reviewed studies analyzing in DDA mode achieved promising results, DIA mode is considered to be better than DDA in accuracy and reproducibility due to the unfiltered detection of peptide mixtures, which can be deployed as an improvement.
3 Mass Spectrometry-based Glycoproteomics
3.1 The Overview and Techniques of Glycoproteomics
Glycome refers to the entire complement of carbohydrates produced by cells or tissues, which is composed of varied sequences and conjugated to proteins and lipids forming glycoproteins and glycolipids, respectively [93]. Glycomics is the systematic study on the comprehensive intra/extra-cellular glycome in specific spatiotemporal conditions and environments, indicating the cellular processes governed by interactions among glycans, proteins, and lipids [94, 95]. In many cases, bacteria and viruses infect the host via carbohydrate–carbohydrate interactions, which correlates to the ongoing COVID-19 pandemic as well [95,96,97].
Glycoproteomics is of great importance in the identification of bacteria and viruses as well as in immunology and pharmaceuticals, which is an emerging frontier in biomarker discovery. Although many glycoproteomics studies have been reported concerning the glycosylated spike (S) protein and the infection mechanism it caused between SARS-CoV-2 and host cell receptors, there is still no substantial discovery of a biomarker that can be applied in pathology and drug development [98,99,100,101,102]. However, based on previous studies on the glycoproteomics of SARS and MERS, glycoproteins would not be trivial modifications on the surface of SARS-CoV-2 and deserve further research [103, 104]. To profile the glycosylation status in COVID-19 patient plasma, the process can be enrichment, digestion, MS analysis, and data acquisition in that order. Enrichment strategies for glycoproteins must be following the target that aims to enrich either a certain protein or a broad pack of proteins. Based on the clear experimental target, a series of enrichment methods can be applied for glycoprotein fractionation and enrichment, including differential centrifugation, two-phase separation, biotinylation, etc [105,106,107,108]. Glycoproteomics analysis can be performed separately using different techniques, with structural glycoprotein characterization being the basis and their release from the glycan carrier being the key step. Unlike the linkage of the peptide backbone, the N- and O-linkage of monosaccharides and glycosaminoglycans (GAGs) require the choice of the corresponding release method. For example, for N-glycans with a common structure (Man3GlcNAc2), the use of Peptide-N-glycosidase F can divide the bond between asparagine and core GlcNAc of the N-glycan [109]. Chemical methods, e.g., hydrazinolysis [110] and β-elimination [111], can also be deployed to release the N-glycans, but are not as efficient as enzymatic cleavage. While for the complete release of O-glycans, no single enzyme is competent due to the diversity of Ser/Thr-linked glycans, preferring to use chemical methods.
Similar to proteomics, MS has become a major tool for both glycan and glycopeptides analysis. Compared to other means (e.g., nuclear magnetic resonance, exoglycosidase treatment, or lectin analysis), MS-based glycan analysis can yield a tremendous amount of structural information: MS detects glycan components, while MS/MS or multilevel MS can elucidate details of positional and linkage isomers [112, 113]. MALDI and ESI methods are the primary ionization methods [114] while LC–ESI–MS has the features of high sensitivity, low ion suppression, and provides a resolution of position and linkage isomers [115]. To homogenize the physicochemical properties of the polysaccharide pool and thus reduce ion suppression, glycans are often derivatized, such as stable isotope permethylation [116]. Glycoproteomics analysis by MALDI has the advantages of easy sample preparation, automation, fast data acquisition, and the ability to observe single ions [117]. And MALDI combined with derivatization techniques such as methylation can avoid the loss of acidic groups during ionization, making it the most used method for qualitative analysis. As the detailed methodology for glycan and glycopeptides is well reviewed particularly regarding its application in SARS-CoV-2 [118], we will not cover the details in experimental considerations and design in this mini-review.
3.2 Current Glycoproteomics Discovery Status
Glycosylation of viral envelope proteins is known as one way in which viruses mask epitopes of associated proteins [119]. The pathogen uses glycosylation to evade recognition of the host immune system, making it undetectable and possibly interfering with the host's adaptive immunity and even enhancing the infectivity of the virus [120, 121]. Several pathogenic viruses including HIV, influenza virus, SARS, and Zika virus, are known to use host-derived glycosylation events to build their glycoproteins on the virion surface, both N-linked and O-linked, to infect their target host cells [119]. Similar to other coronaviruses, the differential organization of glycosylation in SARS-CoV-2 affects not only the composition of individual glycans but also the immune stress on the entire viral protein surface, thereby exposing vulnerable areas in the dense carbohydrate layer of the surface [95, 103].
Glycoproteins not only remarkably behave in the viral surface envelope but also the binding effect of the receptor. This role of glycoproteins explains the mechanism of cross-species transmission of some coronaviruses and provides clues to trace the origin of the virus. Hulswit et al. reported that the glycosylated 9-O-Ac-Sia-specific receptor-binding site on β1-coronavirus binds to the glycoprotein of the receptor [122]. Qing et al. also reported that two receptor-binding sites of coronaviruses, one S1A would bind to host sialic acid, and the other S1B would recognize host transmembrane proteins, thus potentially triggering infections from zoonotic to human-to-human transmission [123]. However, due to the relatively low abundance of glycosylation events as well as the technical challenge to enrich and analyze glycopeptides, all the current findings are still in an early stage in terms of biomarker discovery.
The inevitable challenges in the virus glycoprotein investigations are the macro-heterogeneity and the micro-heterogeneity resulting from the mixture of diverse glycosylation at multiple sites of the protein [124, 125]. Macro-heterogeneity concerns the occupancy, presence, or even absence of glycans at glycosites, while micro-heterogeneity refers to the varieties of glycans at a specific glycosite [125]. Both forms of heterogeneity significantly influence the physical and biochemical properties of proteins [125, 126]. N- and O-glycans also exhibit unique heterogeneity. The diversity of N-glycan structures leads to high micro-heterogeneity and low macro-heterogeneity at specific sites [127, 128]. Variations in macro-heterogeneity are also evident for mucin-type O-glycosylation [129]. Mucin-type O-glycosylation usually has high peptide-level valence and micro-heterogeneity, which may facilitate the formation of multiple O-glycosylations in the neighborhood, but the glycans at each site may be different [129, 130]. Although MS-based glycoproteomics is capable of heterogeneity analysis of all classes of glycosylation, the complexity of glycoforms and variability in the chemical properties of glycoproteins relative to non-modified proteins make enrichment a necessary step before MS analysis [131,132,133,134]. Enrichment of glycopeptides isolates the glycans and glycoconjugates from the non-glycosylated background, thus greatly improving the sensitivity of MS analysis. Affinity chromatography is used as the most common enrichment strategy, based on using specific biochemical interactions of analytes with immobilized ligands to enrich substances of interest from background matrices [135, 136]. Lectins are one of affinity chromatography due to their recognition of carbohydrates and their ability to separate glycans and substrates with varying degrees of specificity [137]. Lectins are commonly used for glycoprotein enrichment include concanavalin A, wheat germ agglutinin, Ricinus communis agglutinin, galectins, and siglecs, and are usually combined with supporting materials, such as agarose or polystyrene-divinylbenzene [137, 138]. Two other types of affinity chromatography, immobilized metal affinity chromatography (IMAC) and metal oxide affinity chromatography (MOAC), offer greater advantages in enriching negatively charged glycans. IMAC and MOAC are both derived from phosphoproteomics with refinements [139, 140]. IMAC consists of transition metal cations (e.g., Fe3+, Ga3+, Ti4+, Zr4+) chelated to an immobilized substrate, whereas MOAC is a mix of transition metals in a metal oxide matrix (e.g., TiOx) [141, 142]. Both techniques exploit the affinity of deprotonated carboxyl groups to achieve enrichment and are particularly effective in enriching sialylated glycopeptides [140,141,142,143]. Hydrophilic interaction chromatography (HILIC) is another important tool for glycoprotein enrichment and characterization and can be applied to a wide range of biomass, such as biological fluids, cancer systems, pathogens, and plants [144,145,146,147]. HILIC exploits the hydrophilic nature of glycans to enrich glycopeptides: due to the difference between the semi-aqueous mobile phase and the hydrophilic stationary phase, enrichment is achieved when glycopeptides elute from the organic loading buffer into the hydrophilic environment [147,148,149]. The recent emergence of porous graphitic carbon (PGC), a chromatographic method capable of both polar and hydrophilic properties, overcomes the disadvantages of silica-based stationary phases and has proven to be an effective method for the separation and enrichment of glycopeptides and glycans [150,151,152]. In addition, the chemical method represented by hydrazide, which has been improved over the years, combined with the release of PNGase F, can also selectively react with glycan derivatives, thus becoming one of the methods for glycoprotein enrichment [153]. However, there is no one universal enrichment strategy for glycoproteomics so far. Different approaches can be adapted for the glycans of interest, which means that the experimental design needs to start from a practical goal and base on the experimental data to prove which approach is applicable.
Glycoproteomics requires much higher overall throughput, data quality, and accessibility for complete glycopeptide identification than conventional proteomics, posing new challenges for both algorithms and search engines. When analyzing intact glycopeptides, it is often necessary to combine multiple sample processing strategies, different MS/MS fragments, and various software to process data, which affects the throughput and quality of MS acquisitions [154,155,156]. Due to the lack of comprehensive quality control, search engine matches for all three aspects of glycans, peptides, and glycopeptides are prone to high false discovery rates (FDR) and lack of validation of spectral interpretation [154]. However, Liu et al. [157] developed a new MS acquisition method and a specialized search engine to address these limitations. By optimizing MS/MS collision parameters, this MS acquisition method can analyze integrated fragments of intact glycopeptides in a single spectrum. The search engine named pGlyco 2.0 can take full advantage of integrated fragments in a spectrum and thus control the quality of glycopeptide-spectral matches (GPSMs). Daniel et al. [158] developed an MSFragger-based glycoproteomics search engine, MSFragger-Glyco, which can search N- and O-glycopeptides quickly and sensitively. The identification results of this search engine are more than doubled the original search outcomes. Although glycoproteomics still faces many challenges, existing and continuing advances in technology continue to drive glycoproteomics to create great value in many fields, including virology and pharmacology.
3.3 Glycoproteomics Characterization of SARS-CoV-2 Spike Protein
Many studies declared that the SARS-CoV-2 carries out cell invasion through a densely glycosylated S protein [98,99,100,101,102]. As a trimeric class I fusion, S protein consists of two subunits S1 and S2, which are generated via proteolytic cleavage [98, 100]. S1 contains the receptor-binding domain (RBD) and is decisive for receptor recognition, while S2 is responsible for membrane fusion and is essential for cell adhesion and immune protection [159]. When S1 binds to the ACE2 receptor of the host cell, S1 will be shed from the S protein, allowing the virus to fuse to the host cell membrane using S2 [98, 160]. Interestingly, the S protein of SARS-CoV-2 has been reported as a major target for neutralizing antibodies as well [104]. Hence, it remains a critical question whether glycosylation of S proteins in coronaviruses allows adequate exposure of viral protein epitopes or acts as a fundamental role in immune evasion.
Emerging experimental evidence, as well as bioinformatic analysis, has pointed out that spike protein is heavily glycosylated. According to sequence features, S protein accommodates at least 22 potential N-glycosylation sites (hence with each trimer presenting 66 N-linked glycosylation sites) and at least 3 mucin-type O-glycosylation sites [161,162,163,164,165]. Eight sites (17, 61, 74, 122, 149, 165, 234, 282) are in the N-terminal domain of the S protein, two (331 and 343) in the RBD, two (1098 and 1134) in the connector domain, and the other sites are outside the functional domains [163]. Three O-glycosylation sites are located within the S1 subunit at Ser673, Thr678, and Ser686 residues [161, 165]. Notably, Shajahan et al. identified an unreported O-glycosylation site at Thr323 and another possible one at Ser525 of the RBD, which may play a key role in viral binding to its cellular receptor ACE2 [166]. Sanda et al. identified eight additional O-glycopeptides near the furin cleavage site of the spike glycoprotein for the first time [165]. These O-glycosylation sites are thought to protect S protein epitopes or key residues from immune system attack and have an important contribution to the immune escape of viruses [161, 166, 167]. Whether all glycosylation sites are glycosylated simultaneously and constantly, or affected by the host environment and other factors, remains an open question to be explored.
The glycan composition that occurred on glycosylation sites is another critical issue to understand the virus–host interaction. Using mass spectrometry-based glycoproteomics, it is found that the glycans on spike protein mainly are high mannose, hybrid, and complex glycans [162,163,164,165,166, 168]. Specifically, eight N-glycosylation sites are predominantly high-mannose-type and the other sites are primarily complex glycans [163, 166]. The two significant high-mannose-type sites (containing more than 80% high mannose) on S protein are N234 and N709 [163]. The major high-mannose-type glycan structure, except N234 with Man9GlcNAc2, at other seven sites on the S protein is Man5GlcNAc2 (Man, mannose; GlcNAc, N-acetylglucosamine), making these sites susceptible as substrates for α-1,2-mannosidases but not for reaction with GlcNAcT-I, and thus unable to be processed into the hybrid- and complex-type glycans [163]. Moreover, the glycan composition and occupancy of the respective sites may be different when S1 and S2 are expressed separately [166]. What factors contributed to the divergence and how they will affect the molecular function remain a question.
Heterogeneity of glycosylation of SARS-CoV-2 Spike protein is reported [163, 164, 166]. Species and quantities of N-glycan on S protein differ when isolated from different host cells [164]. Zhang et al. predicted that native N-glycosylation processing of the S protein of SARS-CoV-2 produces mature glycans that should be identical to recombinant proteins expressed in human cells [164]. Miller et al. evaluated the glycosylation-related heterogeneity of the N-Glycosylation site and showed heterogeneity in the degree of glycan processing of S proteins, with some trimers being more processed than others. This heterogeneity may have a role in confusing the host immune system [162]. Shajahan et al. characterized quantitative N-glycosylation profiles of S proteins and executed an extensive manual interpretation strategy to enrich the data on N- and O-glycosylation to confirm the complexity of glycosylation in SARS-CoV-2 [166]. Furthermore, several studies indicated the glycosylation sites on S protein are relatively conserved during the rapid global spread [161, 167, 169, 170]. This conservation indicates its critical functions in the virus life cycle and host adaptation. The binding of the S protein to the ACE2 receptor is mainly due to the interaction of polar residues between the RBD and the structural domain of ACE2 [11, 99, 169]. Trimeric, complex glycosylated S proteins give them an advantage over monomeric and immature glycosylated variants for receptor binding [171]. The hinge-like dynamic movement of the RBD on the S protein occurs intensifies the affinity of the RBD to ACE2 up to 10–20 times, which partly explains the high viral transmissibility [98, 99, 172]. In contrast, if the biosynthesis of N-glycans is blocked at the oligomannose stage, or if the synthesis of O-glycans is blocked, this will increase the breakdown of spike-protein and thus reduce the possibility of viral binding to ACE2 [173].
4 Limitations and Perspectives
To extend the comprehension of the SARS-CoV-2 infectious mechanism and pathogenesis for a more rapid and sensitive diagnosis, we reviewed MS-based proteomics and glycoproteomics and their application in finding COVID-19 biomarkers from bodies fluids of patients with different stages or preconditions. In the face of the aggressive outbreak of COVID-19, it is urgent to develop a speedy and precise diagnosis of SARS-CoV-2 so that appropriate medical measures can be deployed at an early stage of infection. Since the outbreak of the COVID-19 pandemic, LC–MS-based proteomics and glycoproteomics have been used primarily to find biomarkers to identify SARS-CoV-2, approve drug targets, assess medical efficacy, or elucidate molecular mechanisms of pathogenesis and disease severity [174]. Although SARS-CoV-2 shows signs of mutation and the complexity of the pathogenesis and symptoms of COVID-19 will change accordingly, MS-based histological techniques still have great advantages to help understand the prevalence of COVID-19 [175].
A great challenge is the limited number of cases used for the biomarker study. Given the great variations of demographics of patients (age, gender) and pre-existing diseases, such as immunosuppression, chronic renal insufficiency, obesity, and diabetes, a larger number of samples should be collected to eliminate the effects of these biases and to reveal a true clinic biomarker. In the research containing 1099 confirmed COVID-19 patients, 23.7% (261) patients were diagnosed with at least one comorbidity, demonstrating the implication of pre-existing conditions for the COVID-19 [176]. Another challenge is the sampling time of the patients. Due to individual differences, the time of disease onset may vary from patient to patient, and consequently, when comparing samples, we may be comparing patients at different stage of disease progression, which makes the results more confusing. Therefore, both cross-sectional and longitudinal analyses of patient samples are needed to have a thorough understanding of the disease progression. Ideally, a combinatory approach comparing different patients at the same stage and the same patient at a different stage of COVID-19 would greatly advance our understanding of which protein(s) decide the disease progression.
In the study of proteomics, contemporary MS has been tremendously developed and its resolution can reach even up to 100,000 [177]. Even so, the dynamic range of MS detection is still limited for the profiling of complex samples. For example, to analyze plasma proteins from COVID-19 patients, the challenges posed by the large range of plasma protein concentrations need to be overcome. Commonly deployed strategies include (I) removal of high amount proteins, especially the albumin and immunoglobulins within the plasma, to avoid covering less abundant ones, (II) fractionation of plasma proteins by chromatography, gel electrophoresis, or other means to reduce complexity, and (III) separation of target groups of proteins or peptides of interest using strategies such as the ELISA. Proteins may contain multiple glycan modification sites, and the glycan chain type (O- or N-glycan) and glycan chain occupancy may be different at each site (macro-heterogeneity), while multiple different glycan chain structures may be contained at a single site (micro-heterogeneity) [178]. The macro- and micro- heterogeneities of glycan chain structures need to be resolved one by one in glycoprotein structural analysis, so the establishment of efficient, highly specific, and sensitive methods for the enrichment and analysis of glycosylation modifications on a scale is the key to the in-depth study of glycosylation.
At the time when COVID-19 is ravaging the world, rapid detection of the virus is an elemental strategy to control the sprawl of the disease. Multi-omics for biomarker discovery and altered molecular network investigations are dominant and valuable tools for a more comprehensive overview of COVID-19. In particular, proteomics and the emerging glycoproteomics can provide gene expression and post-transcriptional information that, when applied to the search for biomarkers of COVID-19, can contribute to a deeper exploration of the infection and pathogenesis at molecular levels, and ultimately uncover therapeutic strategies. MS-based proteomics and glycoproteomics have the advantage of being high-throughput and non-biased and are exceptionally promising to open up key strategies to defeat the COVID-19 pandemic.
References
The World Health Organization (WHO). Pneumonia of unknown cause—China. World Health Organization; 2020. http://www.who.int/csr/don/05-january-2020-pneumonia-of-unkown-cause-china/en/. Accessed 22 Mar 2021.2020
Chan JFW, Yuan S, Kok KH, To KKW, Chu H, Yang J, Xing F, Liu J, Yip CCY, Poon RWS, Tsoi HW, Lo SKF, Chan KH, Poon VKM, Chan WM, Ip JD, Cai JP, Cheng VCC, Chen H, Hui CKM, Yuen KY. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 2020;395:514–23.
Zhou P, Lou YX, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Di JR, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL. Discovery of a novel coronavirus associated with the recent pneumonia outbreak in humans and its potential bat origin. bioRxiv. 2020. https://doi.org/10.1101/2020.01.22.914952.
Lai CC, Shih TP, Ko WC, Tang HJ, Hsueh PR. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): the epidemic and the challenges. Int J Antimicrob Agents. 2020;55:105924.
Grifoni A, Sidney J, Zhang Y, Scheuermann RH, Peters B, Sette A. A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2. Cell Host Microbe. 2020;27:671-680.e2.
Dong E, Du H, Gardner L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect Dis. 2020;20:533–4.
Yuki K, Fujiogi M, Koutsogiannaki S. COVID-19 pathophysiology: a review. Clin Immunol. 2020;215:108427.
Battagello DS, Dragunas G, Klein MO, Ayub ALP, Velloso FJ, Correa RG. Unpuzzling COVID-19: tissue-related signaling pathways associated with SARS-CoV-2 infection and transmission. Clin Sci. 2020;134:2137–60.
Woods JA, Hutchinson NT, Powers SK, Roberts WO, Gomez-Cabrera MC, Radak Z, Berkes I, Boros A, Boldogh I, Leeuwenburgh C, Coelho-Júnior HJ, Marzetti E, Cheng Y, Liu J, Durstine JL, Sun J, Ji LL. The COVID-19 pandemic and physical activity. Sport Med Heal Sci. 2020;2:55–64.
Liya G, Yuguang W, Jian L, Huaiping Y, Xue H, Jianwei H, Jiaju M, Youran L, Chen M, Yiqing J. Studies on viral pneumonia related to novel coronavirus SARS-CoV-2, SARS-CoV, and MERS-CoV: a literature review. APMIS. 2020;128:423–32.
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181:271-280.e8.
South AM, Diz DI, Chappell MC. COVID-19, ACE2, and the cardiovascular consequences. Am J Physiol Hear Circ Physiol. 2020;318:H1084–90.
Varga Z, Flammer AJ, Steiger P, Haberecker M, Andermatt R, Zinkernagel AS, Mehra MR, Schuepbach RA, Ruschitzka F, Moch H. Endothelial cell infection and endotheliitis in COVID-19. Lancet. 2020;395:1417–8.
Menni C, Valdes AM, Freidin MB, Ganesh S, El-Sayed Moustafa JS, Visconti A, Hysi P, Bowyer RCE, Mangino M, Falchi M, Wolf J, Steves CJ, Spector TD. Loss of smell and taste in combination with other symptoms is a strong predictor of COVID-19 infection. medRxiv. 2020. https://doi.org/10.1101/2020.04.05.20048421.
Wang HY, Li XL, Yan ZR, Sun XP, Han J, Zhang BW. Potential neurological symptoms of COVID-19. Ther Adv Neurol Disord. 2020;13:175628642091783.
Gu J, Han B, Wang J. COVID-19: gastrointestinal manifestations and potential fecal-oral transmission. Gastroenterology. 2020;158:1518–9.
Backer JA, Klinkenberg D, Wallinga J. Incubation period of 209 novel coronavirus (2019-nCoV) infections among travellers from Wuhan, China, 20 28 January 2020. Eurosurveillance. 2020;25(5):2000062.
Wang W, Tang J, Wei F. Updated understanding of the outbreak of 2019 novel coronavirus (2019-nCoV) in Wuhan. China J Med Virol. 2020;92:441–7.
Davies NG, Barnard RC, Jarvis CI, Kucharski AJ, Munday J, Pearson CAB, Russell TW, Tully DC, Clifford S, et al. Estimated transmissibility and severity of novel SARS-CoV-2 variant of concern 202012/01 in England. medRxiv. 2020. https://doi.org/10.1101/2020.12.24.20248822.
Wibmer CK, Ayres F, Hermanus T, Madzivhandila M, Kgagudi P, Lambson BE, Vermeulen M, van den Berg K, Rossouw T, Boswell M, Ueckermann V, Meiring S, von Gottberg A, Cohen C, Morris L, Bhiman JN, Moore PL. SARS-CoV-2 501Y.V2 escapes neutralization by South African COVID-19 donor plasma. bioRxiv Prepr Serv Biol. 2021;27(4):622–5.
Sabino EC, Buss LF, Carvalho MPS, Prete CA, Crispim MAE, Fraiji NA, Pereira RHM, Parag KV, da Silva PP, Kraemer MUG, Oikawa MK, Salomon T, Cucunuba ZM, Castro MC, de Souza Santos AA, Nascimento VH, Pereira HS, Ferguson NM, Pybus OG, Kucharski A, Busch MP, Dye C, Faria NR. Resurgence of COVID-19 in Manaus, Brazil, despite high seroprevalence. Lancet. 2021;397:452–5.
Shu T, Ning W, Wu D, Xu J, Han Q, Huang M, Zou X, Yang Q, Yuan Y, Bie Y, Pan S, Mu J, Han Y, Yang X, Zhou H, Li R, Ren Y, Chen X, Yao S, Qiu Y, Zhang D-Y, Xue Y, Shang Y, Zhou X. Plasma proteomics identify biomarkers and pathogenesis of COVID-19. Immunity. 2020;53:1108+.
Shen B, Yi X, Sun Y, Bi X, Du J, Zhang C, Quan S, Zhang F, Sun R, Qian L, Ge W, Liu W, Liang S, Chen HH, Zhang Y, Li J, Xu J, He Z, Chen B, Wang J, Yan H, Zheng Y, Wang D, Zhu J, Kong Z, Kang Z, Liang X, Ding X, Ruan G, Xiang N, Cai X, Gao H, Li L, Li S, Xiao Q, Lu T, Zhu Y, Liu H, Chen HH, Guo T. Proteomic and metabolomic characterization of COVID-19 patient sera. Cell. 2020;182:59-72.e15.
Nie X, Qian L, Sun R, Huang B, Dong X, Xiao Q, Zhang Q, Lu T, Yue L, Chen S, Li X, Sun Y, Li L, Xu L, Li Y, Yang M, Xue Z, Liang S, Ding X, Yuan C, Peng L, Liu W, Yi X, Lyu M, Xiao G, Xu X, Ge W, He J, Fan J, Wu J, Luo M, Chang X, Pan H, Cai X, Zhou J, Yu J, Gao H, Xie M, Wang S, Ruan G, Chen H, Su H, Mei H, Luo D, Zhao D, Xu F, Zhu Y, Xia J, Hu Y, Guo T. Multi-organ proteomic landscape of COVID-19 autopsies. Cell. 2021;184:775-791.e14.
Hüttenhain R, Malmström J, Picotti P, Aebersold R. Perspectives of targeted mass spectrometry for protein biomarker verification. Curr Opin Chem Biol. 2009;13:518–25.
Bahir I, Fromer M, Prat Y, Linial M. Viral adaptation to host: a proteome-based analysis of codon usage and amino acid preferences. Mol Syst Biol. 2009;5(1):311.
Long JS, Mistry B, Haslam SM, Barclay WS. Host and viral determinants of influenza A virus species specificity. Nat Rev Microbiol. 2019;17:67–81.
Taubenberger JK, Kash JC. Influenza virus evolution, host adaptation, and pandemic formation. Cell Host Microbe. 2010;7:440–51.
Duyen HTL, Ngoc TV, Ha DT, Hang VTT, Kieu NTT, Young PR, Farrar JJ, Simmons CP, Wolbers M, Wills BA. Kinetics of plasma viremia and soluble nonstructural protein 1 concentrations in dengue: differential effects according to serotype and immune status. J Infect Dis. 2011;203:1292–300.
Braun L, Brenier-Pinchart MP, Yogavel M, Curt-Varesano A, Curt-Bertini RL, Hussain T, Kieffer-Jaquinod S, Coute Y, Pelloux H, Tardieux I, Sharma A, Belrhali H, Bougdour A, Hakimi MA. A Toxoplasma dense granule protein, GRA24, modulates the early immune response to infection by promoting a direct and sustained host p38 MAPK activation. J Exp Med. 2013;210:2071–86.
Scaturro P, Kastner AL, Pichlmair A. Chasing intracellular Zika virus using proteomics. Viruses. 2019;11:878.
García-Dorival I, Wu W, Armstrong SD, Barr JN, Carroll MW, Hewson R, Hiscox JA. Elucidation of the cellular interactome of ebola virus nucleoprotein and identification of therapeutic targets. J Proteome Res. 2016;15:4290–303.
Stalin Raj V, Lamers MM, Smits SL, Demmers JAA, Mou H, Bosch BJ, Haagmans BL. Identification of protein receptors for coronaviruses by mass spectrometry. In: Helena JM, Erica B, Paul B, editors. Coronaviruses: methods and protocols. New York: Springer; 2015. p. 165–82.
Prabakaran S, Lippens G, Steen H, Gunawardena J. Post-translational modification: nature’s escape from genetic imprisonment and the basis for dynamic information encoding. Wiley Interdiscip Rev Syst Biol Med. 2012;4:565–83.
Kirchdoerfer RN, Wang N, Pallesen J, Wrapp D, Turner HL, Cottrell CA, Corbett KS, Graham BS, McLellan JS, Ward AB, Benvenuto D, Giovannetti M, Ciccozzi A, Spoto S, Angeletti S, Ciccozzi M, Jahn R, Lang T, Südhof TC, Lie F, Lite W, Farzan M, Harrison SC, Hyochol A, Michael W, Debra L, Eunyoung Choi RN, Roger Fillingim PB, Nomaguchi M, Fujita M, Miyazaki Y, Adachi A, Shulla A, Gallagher T, Walls AC, Xiong X, Park YJ, Tortorici MA, Snijder J, Quispe J, Cameroni E, Gopal R, Dai M, Lanzavecchia A, Zambon M, Rey FA, Corti D, Veesler D. Unexpected receptor functional mimicry elucidates activation of coronavirus fusion. Cell. 2018;176:1864–8.
Ianni M, Manerba M, Di Stefano G, Porcellini E, Chiappelli M, Carbone I, Licastro F. Altered glycosylation profile of purified plasma ACT from Alzheimer’s disease. Immun Ageing. 2010. https://doi.org/10.1186/1742-4933-7-S1-S6.
De Leoz MLA, Young LJT, An HJ, Kronewitter SR, Kim J, Miyamoto S, Borowsky AD, Chew HK, Lebrilla CB. High-mannose glycans are elevated during breast cancer progression. Mol Cell Proteom. 2011. https://doi.org/10.1074/mcp.M110.002717.
Ptolemy AS, Rifai N. What is a biomarker? Research investments and lack of clinical integration necessitate a review of biomarker terminology and validation schema. Scand J Clin Lab Invest. 2010;70:6–14.
Liao J, Fan S, Chen J, Wu J, Xu S, Guo Y, Li C, Zhang X, Wu C, Mou H, Song C, Li F, Wu G, Zhang J, Guo L, Liu H, Lv J, Xu L, Lang C. Epidemiological and clinical characteristics of COVID-19 in adolescents and young adults. Innov. 2020;1:100001.
Clark A, Jit M, Warren-Gash C, Guthrie B, Wang HHX, Mercer SW, Sanderson C, McKee M, Troeger C, Ong KL, Checchi F, Perel P, Joseph S, Gibbs HP, Banerjee A, Eggo RM. Global, regional, and national estimates of the population at increased risk of severe COVID-19 due to underlying health conditions in 2020: a modelling study. Lancet Glob Heal. 2020;8:e1003–17.
Ong EZ, Chan YFZ, Leong WY, Lee NMY, Kalimuddin S, Haja Mohideen SM, Chan KS, Tan AT, Bertoletti A, Ooi EE, Low JGH. A dynamic immune response shapes COVID-19 progression. Cell Host Microbe. 2020;27:879-882.e2.
Wilk AJ, Rustagi A, Zhao NQ, Roque J, Martínez-Colón GJ, McKechnie JL, Ivison GT, Ranganath T, Vergara R, Hollis T, Simpson LJ, Grant P, Subramanian A, Rogers AJ, Blish CA. A single-cell atlas of the peripheral immune response in patients with severe COVID-19. Nat Med. 2020;26:1070–6.
Catanzaro M, Fagiani F, Racchi M, Corsini E, Govoni S, Lanni C. Immune response in COVID-19: addressing a pharmacological challenge by targeting pathways triggered by SARS-CoV-2. Signal Transduct Target Ther. 2020;5(1):1.
Zhu L, Yang P, Zhao Y, Zhuang Z, Wang Z, Song R, Zhang J, Liu C, Gao Q, Xu Q, Wei X, Sun HX, Ye B, Wu Y, Zhang N, Lei G, Yu L, Yan J, Diao G, Meng F, Bai C, Mao P, Yu Y, Wang M, Yuan Y, Deng Q, Li Z, Huang Y, Hu G, Liu Y, Wang X, Xu Z, Liu P, Bi Y, Shi Y, Zhang S, Chen Z, Wang J, Xu X, Wu G, Wang FS, Gao GF, Liu L, Liu WJ. Single-cell sequencing of peripheral mononuclear cells reveals distinct immune response landscapes of Covid-19 and influenza patients. Immunity. 2020;53:685-696.e3.
Freyberg Z, Harvill ET. Pathogen manipulation of host metabolism: A common strategy for immune evasion. PLoS Pathog. 2017;13:e1006669.
Eisenreich W, Rudel T, Heesemann J, Goebel W. How viral and intracellular bacterial pathogens reprogram the metabolism of host cells to allow their intracellular replication. Front Cell Infect Microbiol. 2019;9:42.
Kim CH. Immune regulation by microbiome metabolites. Immunology. 2018;154:220–9.
Nicora CD, Sims AC, Bloodsworth KJ, Kim YM, Moore RJ, Kyle JE, Nakayasu ES, Metz TO. Metabolite, protein, and lipid extraction (MPLEx): a method that simultaneously inactivates middle east respiratory syndrome coronavirus and allows analysis of multiple host cell components following infection. In: Rahul V, editor. Methods in molecular biology. Humana Press Inc.; 2020. p. 173–94.
Byers NM, Fleshman AC, Perera R, Molins CR. Metabolomic insights into human arboviral infections: dengue, chikungunya, and zika viruses. Viruses. 2019. https://doi.org/10.3390/v11030225.
de Haan N, Wuhrer M, Ruhaak LR. Mass spectrometry in clinical glycomics: the path from biomarker identification to clinical implementation. Clin MASS Spectrom. 2020;18:1–12.
Bojkova D, Klann K, Koch B, Widera M, Krause D, Ciesek S, Cinatl J, Münch C. Proteomics of SARS-CoV-2-infected host cells reveals therapy targets. Nature. 2020;583:469–72.
Bouhaddou M, Memon D, Meyer B, White KM, Rezelj VV, Correa Marrero M, Polacco BJ, Melnyk JE, Krogan NJ, et al. The global phosphorylation landscape of SARS-CoV-2 infection. Cell. 2020;182:685-712.e19.
Liu S, Zhu L, Xie G, Mok BW-Y, Yang Z, Deng S, Lau S-Y, Chen P, Wang P, Chen H, Cai Z. Potential antiviral target for SARS-CoV-2: a key early responsive kinase during viral entry. CCS Chem. 2021;3:559–68.
Klann K, Bojkova D, Tascher G, Ciesek S, Münch C, Cinatl J. Growth factor receptor signaling inhibition prevents SARS-CoV-2 replication. Mol Cell. 2020;80:164-174.e4.
Greenlund LJS, Sreekumaran Nair K. The Effect of Insulin on Protein Metabolism. In: Sudhesh K, Stephen OR, editors. Insulin resistance: insulin action and its disturbances in disease. UK: Wiley; 2005:105–32.
Cohen P. The regulation of protein function by multisite phosphorylation—a 25 year update. Trends Biochem Sci. 2000;25:596–601.
Pockley AG. Heat shock proteins as regulators of the immune response. Lancet. 2003;362:469–76.
Colas P, Cohen B, Ferrigno PK, Silver PA, Brent R. Targeted modification and transportation of cellular proteins. Proc Natl Acad Sci USA. 2000;97:13720–5.
Crippen GM. The tree structural organization of proteins. J Mol Biol. 1978;126:315–32.
Wasinger VC, Cordwell SJ, Poljak A, Yan JX, Gooley AA, Wilkins MR, Duncan MW, Harris R, Williams KL, Humphery-Smith I. Progress with gene-product mapping of the mollicutes: mycoplasma genitalium. Electrophoresis. 1995;16:1090–4.
Jungblut PR, Holzhütter HG, Apweiler R, Schlüter H. The speciation of the proteome. Chem Cent J. 2008;2:16.
Aebersold R, Mann M. Mass-spectrometric exploration of proteome structure and function. Nature. 2016;537:347–55.
Catherman AD, Skinner OS, Kelleher NL. Top Down proteomics: facts and perspectives. Biochem Biophys Res Commun. 2014;445:683–93.
Kelleher NL. Peer reviewed: top-down proteomics. Anal Chem. 2004;76:196–203.
Wu CC, MacCoss MJ. Shotgun proteomics: tools for the analysis of complex biological systems. Curr Opin Mol Ther. 2002;4:242–50.
Gault J, Donlan JAC, Liko I, Hopper JTS, Gupta K, Housden NG, Struwe WB, Marty MT, Mize T, Bechara C, Zhu Y, Wu B, Kleanthous C, Belov M, Damoc E, Makarov A, Robinson CV. High-resolution mass spectrometry of small molecules bound to membrane proteins. Nat Methods. 2016;13:333–6.
Mitchell Wells J, McLuckey SA. Collision-induced dissociation (CID) of peptides and proteins. Methods Enzymol. 2005;402:148–85.
Yu Q, Shi X, Feng Y, Kent KC, Li L. Improving data quality and preserving HCD-generated reporter ions with EThcD for isobaric tag-based quantitative proteomics and proteome-wide PTM studies. Anal Chim Acta. 2017;968:40–9.
Zhou Y, Shan Y, Zhang L, Zhang Y. Progress in stable isotope labeled quantitative proteomics methods. Chin J Chromatogr (Se Pu). 2013;31:496–502.
Bantscheff M, Schirle M, Sweetman G, Rick J, Kuster B. Quantitative mass spectrometry in proteomics: a critical review. Anal Bioanal Chem. 2007;389:1017–31.
Zhu JL, Zhang K, He XW, Zhang YK. New developments of quantitative mass spectrometry-based proteomics. Chin J Anal Chem. 2010;38:434–41.
Brönstrup M. Absolute quantification strategies in proteomics based on mass spectrometry. Expert Rev Proteomic. 2004;1:503–12.
Brun V, Masselon C, Garin J, Dupuis A. Isotope dilution strategies for absolute quantitative proteomics. J Proteomic. 2009;72:740–9.
Rifai N, Gillette MA, Carr SA. Protein biomarker discovery and validation: the long and uncertain path to clinical utility. Nat Biotechnol. 2006;24:971–83.
Thompson A, Schäfer J, Kuhn K, Kienle S, Schwarz J, Schmidt G, Neumann T, Hamon C. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal Chem. 2003;75:1895–904.
Ross PL, Huang YN, Marchese JN, Williamson B, Parker K, Hattan S, Khainovski N, Pillai S, Dey S, Daniels S, Purkayastha S, Juhasz P, Martin S, Bartlet-Jones M, He F, Jacobson A, Pappin DJ. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol Cell Proteomic. 2004;3:1154–69.
Neilson KA, Ali NA, Muralidharan S, Mirzaei M, Mariani M, Assadourian G, Lee A, Van Sluyter SC, Haynes PA. Less label, more free: approaches in label-free quantitative mass spectrometry. Proteomic. 2011;11:535–53.
Tabb DL, Vega-Montoto L, Rudnick PA, Variyath AM, Ham AJL, Bunk DM, Kilpatrick LE, Billheimer DD, Blackman RK, Cardasis HL, Carr SA, Clauser KR, Jaffe JD, Kowalski KA, Neubert TA, Regnier FE, Schilling B, Tegeler TJ, Wang M, Wang P, Whiteaker JR, Zimmerman LJ, Fisher SJ, Gibson BW, Kinsinger CR, Mesri M, Rodriguez H, Stein SE, Tempst P, Paulovich AG, Liebler DC, Spiegelman C. Repeatability and reproducibility in proteomic identifications by liquid chromatography-tandem mass spectrometry. J Proteome Res. 2010;9:761–76.
Vidova V, Spacil Z. A review on mass spectrometry-based quantitative proteomics: targeted and data independent acquisition. Anal Chim Acta. 2017;964:7–23.
Law KP, Lim YP. Recent advances in mass spectrometry: data independent analysis and hyper reaction monitoring. Expert Rev Proteomic. 2013;10:551–66.
Peterson AC, Russell JD, Bailey DJ, Westphall MS, Coon JJ. Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol Cell Proteomic. 2012;11:1475–88.
Gianazza E, Tremoli E, Banfi C. The selected reaction monitoring/multiple reaction monitoring-based mass spectrometry approach for the accurate quantitation of proteins: clinical applications in the cardiovascular diseases. Expert Rev Proteomic. 2014;11:771–88.
Shao C, Li M, Li X, Wei L, Zhu L, Yang F, Jia L, Mu Y, Wang J, Guo Z, Zhang D, Yin J, Wang Z, Sun W, Zhang Z, Gao Y. A tool for biomarker discovery in the urinary proteome: a manually curated human and animal urine protein biomarker database. Mol Cell Proteomics. 2011;10:M111–010975.
de Souza GA, Godoy LMF, Mann M. Identification of 491 proteins in the tear fluid proteome reveals a large number of proteases and protease inhibitors. Genome Biol. 2006. https://doi.org/10.1186/gb-2006-7-8-r72.
Adachi J, Kumar C, Zhang Y, Olsen JV, Mann M. The human urinary proteome contains more than 1500 proteins, including a large proportion of membrane proteins. Genome Biol. 2006. https://doi.org/10.1186/gb-2006-7-9-r80.
Pilch B, Mann M. Large-scale and high-confidence proteomic analysis of human seminal plasma. Genome Biol. 2006. https://doi.org/10.1186/gb-2006-7-5-r40.
Kim MS, Pinto SM, Getnet D, Nirujogi RS, Manda SS, Chaerkady R, Madugundu AK, Kelkar DS, Isserlin R, Jain S, Thomas JK, Muthusamy B, Leal-Rojas P, Kumar P, Sahasrabuddhe NA, Balakrishnan L, Advani J, George B, Renuse S, Selvan LD, Patil AH, Nanjappa V, Radhakrishnan A, Prasad S, Subbannayya T, Raju R, Kumar M, Sreenivasamurthy SK, Marimuthu A, Sathe GJ, Chavan S, Datta KK, Subbannayya Y, Sahu A, Yelamanchi SD, Jayaram S, Rajagopalan P, Sharma J, Murthy KR, Syed N, Goel R, Khan AA, Ahmad S, Dey G, Mudgal K, Chatterjee A, Huang TC, Zhong J, Wu X, Shaw PG, Freed D, Zahari MS, Mukherjee KK, Shankar S, Mahadevan A, Lam H, Mitchell CJ, Shankar SK, Satishchandra P, Schroeder JT, Sirdeshmukh R, Maitra A, Leach SD, Drake CG, Halushka MK, Prasad TS, Hruban RH, Kerr CL, Bader GD, Iacobuzio-Donahue CA, Gowda H, Pandey A. A draft map of the human proteome. Nature. 2014;509:575–81.
Zhao M, Yang Y, Guo Z, Shao C, Sun H, Zhang Y, Sun Y, Liu Y, Song Y, Zhang L, Li Q, Liu J, Li M, Gao Y, Sun W. A comparative proteomics analysis of five body fluids: plasma, urine, cerebrospinal fluid, amniotic fluid, and saliva. Proteomic Clin Appl. 2018;12:1800008.
Anderson NL. The clinical plasma proteome: a survey of clinical assays for proteins in plasma and serum. Clin Chem. 2010;56:177–85.
Lee HJ, Lee EY, Kwon MS, Paik YK. Biomarker discovery from the plasma proteome using multidimensional fractionation proteomics. Curr Opin Chem Biol. 2006;10:42–9.
Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O'Meara MJ, Rezelj VV, Guo JZ, Swaney DL, Tummino TA, Hüttenhain R, Kaake RM, Richards AL, Tutuncuoglu B, Foussard H, Batra J, Haas K, Modak M, Kim M, Haas P, Polacco BJ, Braberg H, Fabius JM, Eckhardt M, Soucheray M, Bennett MJ, Cakir M, McGregor MJ, Li Q, Meyer B, Roesch F, Vallet T, Mac Kain A, Miorin L, Moreno E, Naing ZZC, Zhou Y, Peng S, Shi Y, Zhang Z, Shen W, Kirby IT, Melnyk JE, Chorba JS, Lou K, Dai SA, Barrio-Hernandez I, Memon D, Hernandez-Armenta C, Lyu J, Mathy CJP, Perica T, Pilla KB, Ganesan SJ, Saltzberg DJ, Rakesh R, Liu X, Rosenthal SB, Calviello L, Venkataramanan S, Liboy-Lugo J, Lin Y, Huang XP, Liu Y, Wankowicz SA, Bohn M, Safari M, Ugur FS, Koh C, Savar NS, Tran QD, Shengjuler D, Fletcher SJ, O'Neal MC, Cai Y, Chang JCJ, Broadhurst DJ, Klippsten S, Sharp PP, Wenzell NA, Kuzuoglu-Ozturk D, Wang HY, Trenker R, Young JM, Cavero DA, Hiatt J, Roth TL, Rathore U, Subramanian A, Noack J, Hubert M, Stroud RM, Frankel AD, Rosenberg OS, Verba KA, Agard DA, Ott M, Emerman M, Jura N, von Zastrow M, Verdin E, Ashworth A, Schwartz O, d'Enfert C, Mukherjee S, Jacobson M, Malik HS, Fujimori DG, Ideker T, Craik CS, Floor SN, Fraser JS, Gross JD, Sali A, Roth BL, Ruggero D, Taunton J, Kortemme T, Beltrao P, Vignuzzi M, García-Sastre A, Shokat KM, Shoichet BK, Krogan NJ. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature. 2020 ;583:459–68.
Wallentin L, Lindback J, Eriksson N, Hijazi Z, Eikelboom JW, Ezekowitz MD, Granger CB, Lopes RD, Yusuf S, Oldgren J, Siegbahn A. Angiotensin-converting enzyme 2 (ACE2) levels in relation to risk factors for COVID-19 in two large cohorts of patients with atrial fibrillation. Eur Heart J. 2020;41:4037–46.
Varki A. Biological roles of glycans. Glycobiology. 2017;27:3–49.
Raman R, Raguram S, Venkataraman G, Paulson JC, Sasisekharan R. Glycomics: an integrated systems approach to structure-function relationships of glycans. Nat Methods. 2005;2:817–24.
Adamo R, Sonnino S. Impact of glycoscience in fighting Covid-19. Glycoconj J. 2020;37:511–2.
Roberts PC, Garten W, Klenk HD. Role of conserved glycosylation sites in maturation and transport of influenza A virus hemagglutinin. J Virol. 1993;67:3048–60.
Taube S, Jiang M, Wobus CE. Glycosphingolipids as receptors for non-enveloped viruses. Viruses. 2010;2:1011–49.
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367:1260–3.
Zhao P, Praissman JL, Grant OC, Cai Y, Xiao T, Rosenbalm KE, Aoki K, Kellman BP, Bridger R, Barouch DH, Brindley MA, Lewis NE, Tiemeyer M, Chen B, Woods RJ, Wells L. Virus-receptor interactions of glycosylated SARS-CoV-2 spike and human ACE2 receptor. Cell Host Microbe. 2020;28:586-601.e6.
Romeo A, Iacovelli F, Falconi M. Targeting the SARS-CoV-2 spike glycoprotein prefusion conformation: virtual screening and molecular dynamics simulations applied to the identification of potential fusion inhibitors. Virus Res. 2020;286:198068.
Poh CM, Carissimo G, Wang B, Amrun SN, Lee CYP, Chee RSL, Fong SW, Yeo NKW, Lee WH, Torres-Ruesta A, Leo YS, Chen MIC, Tan SY, Chai LYA, Kalimuddin S, Kheng SSG, Thien SY, Young BE, Lye DC, Hanson BJ, Wang CI, Renia L, Ng LFP. Two linear epitopes on the SARS-CoV-2 spike protein that elicit neutralising antibodies in COVID-19 patients. Nat Commun. 2020. https://doi.org/10.1038/s41467-020-16638-2.
Zhang BZ, Hu YF, Chen LL, Yau T, Tong YG, Hu JC, Cai JP, Chan KH, Dou Y, Deng J, Wang XL, Hung IFN, To KKW, Yuen KY, Huang JD. Mining of epitopes on spike protein of SARS-CoV-2 from COVID-19 patients. Cell Res. 2020;30:702–4.
Watanabe Y, Berndsen ZT, Raghwani J, Seabright GE, Allen JD, McLellan JS, Wilson IA, Bowden TA, Ward AB, Crispin M, Pybus OG, McLellan JS, Wilson IA, Bowden TA, Ward AB, Crispin M. Vulnerabilities in coronavirus glycan shields despite extensive glycosylation. Nat Commun. 2020. https://doi.org/10.1101/2020.02.20.957472.
Pinto D, Park Y-J, Beltramello M, Walls A, Tortorici MA, Bianchi S, Jaconi S, Culap K, Zatta F, De Marco A, Peter A, Guarino B, Spreafico R, Cameroni E, Case JB, Chen R, Havenar-Daughton C, Snell G, Telenti A, Virgin H, Lanzavecchia A, Diamond M, Fink K, Veesler D, Corti D. Structural and functional analysis of a potent sarbecovirus neutralizing antibody. bioRxiv Prepr Serv Biol. 2020. https://doi.org/10.1101/2020.04.07.023903.
Chandler KB, Costello CE. Glycomics and glycoproteomics of membrane proteins and cell-surface receptors: present trends and future opportunities. Electrophoresis. 2016;37:1407–19.
Cordwell SJ, Thingholm TE. Technologies for plasma membrane proteomics. Proteomics. 2010;10:611–27.
Mechref Y, Madera M, Novotny MV. Glycoprotein enrichment through lectin affinity techniques. Methods Mol Biol. 2008;424:373–96.
Wollscheid B, Bausch-Fluck D, Henderson C, O’Brien R, Bibel M, Schiess R, Aebersold R, Watts JD. Mass-spectrometric identification and relative quantification of N-linked cell surface glycoproteins. Nat Biotechnol. 2009;27:378–86.
Waddling CA, Plummer TH, Tarentino AL, Van Roey P. Structural basis for the substrate specificity of endo-β-N- acetylglucosaminidase F3. Biochemistry. 2000;39:7878–85.
Patel T, Bruce J, Merry A, Bigge C, Parekh R, Wormald M, Jaques A. Use of hydrazine to release in intact and unreduced form both N- and O-linked oligosaccharides from glycoproteins. Biochemistry. 1993;32:679–93.
Aminoff D, Gathmann WD, McLean CM, Yadomae T. Quantitation of oligosaccharides released by the β-elimination reaction. Anal Biochem. 1980;101:44–53.
Wuhrer M. Glycomics using mass spectrometry. Glycoconj J. 2013;30:11–22.
Zaia J. Mass spectrometry and glycomics. Omi A J Integr Biol. 2010;14:401–18.
Zaia J. Mass spectrometry of oligosaccharides. Mass Spectrom Rev. 2004;23:161–227.
Vanderschaeghe D, Festjens N, Delanghe J, Callewaert N. Glycome profiling using modern glycomics technology: technical aspects and applications. Biol Chem. 2010;391:149–61.
Ciucanu I, Kerek F. A simple and rapid method for the permethylation of carbohydrates. Carbohydr Res. 1984;131:209–17.
Harvey DJ. Matrix-assisted laser desorption/ionization mass spectrometry of carbohydrates. Mass Spectrom Rev. 1999;18:349–450.
Pujić I, Perreault H. Recent advancements in glycoproteomic studies: glycopeptide enrichment and derivatization, characterization of glycosylation in SARS CoV2, and interacting glycoproteins. Mass Spectrom Rev. 2021. https://doi.org/10.1002/mas.21679.
Watanabe Y, Bowden TA, Wilson IA, Crispin M. Exploitation of glycosylation in enveloped virus pathobiology. Biochim Biophys Acta Gen Subj. 2019;1863:1480–97.
Samuel MA, Diamond MS. Pathogenesis of west nile virus infection: a balance between virulence, innate and adaptive immunity, and viral evasion. J Virol. 2006;80:9349–60.
Gerlach D, Guo Y, De Castro C, Kim SH, Schlatterer K, Xu FF, Pereira C, Seeberger PH, Ali S, Codée J, Sirisarn W, Schulte B, Wolz C, Larsen J, Molinaro A, Lee BL, Xia G, Stehle T, Peschel A. Methicillin-resistant Staphylococcus aureus alters cell wall glycosylation to evade immunity. Nature. 2018;563:705–9.
Hulswit RJG, Lang Y, Bakkers MJG, Li W, Li Z, Schouten A, Ophorst B, Van Kuppeveld FJM, Boons GJ, Bosch BJ, Huizinga EG, De Groot RJ. Human coronaviruses OC43 and HKU1 bind to 9-O-acetylated sialic acids via a conserved receptor-binding site in spike protein domain A. Proc Natl Acad Sci USA. 2019;116:2681–90.
Qing E, Hantak M, Perlman S, Gallagher T. Distinct roles for sialoside and protein receptors in coronavirus infection. MBio. 2020. https://doi.org/10.1128/mBio.02764-19.
West CM, Malzl D, Hykollari A, Wilson IBH. Glycomics, glycoproteomics, and glycogenomics: an inter-taxa evolutionary perspective. Mol Cell Proteomic. 2021. https://doi.org/10.1074/MCP.R120.002263.
Čaval T, Heck AJR, Reiding KR. Meta-heterogeneity: Evaluating and describing the diversity in glycosylation between sites on the same glycoprotein. Mol Cell Proteomic. 2021;20: 100010. https://doi.org/10.1074/MCP.R120.002093.
Struwe WB, Robinson CV. Relating glycoprotein structural heterogeneity to function—insights from native mass spectrometry. Curr Opin Struct Biol. 2019;58:241–8.
Wu D, Struwe WB, Harvey DJ, Ferguson MAJ, Robinson CV. N-glycan microheterogeneity regulates interactions of plasma proteins. Proc Natl Acad Sci USA. 2018;115:8763–8.
Blazev R, Ashwood C, Abrahams JL, Chung LH, Francis D, Yang P, Watt KI, Qian H, Quaife-Ryan GA, Hudson JE, Gregorevic P, Thaysen-Andersen M, Parker BL. Integrated glycoproteomics identifies a role of N-glycosylation and galectin-1 on myogenesis and muscle development. Mol Cell Proteomic. 2021;20:100030.
Maynard JC, Chalkley RJ. Methods for enrichment and assignment of N-acetylglucosamine modification sites. Mol Cell Proteomic. 2021. https://doi.org/10.1074/mcp.R120.002206.
Riley NM, Bertozzi CR, Pitteri SJ. A pragmatic guide to enrichment strategies for mass spectrometry-based glycoproteomics. Mol Cell Proteomic. 2021;20:100029.
Xiao H, Sun F, Suttapitugsakul S, Wu R. Global and site-specific analysis of protein glycosylation in complex biological systems with mass spectrometry. Mass Spectrom Rev. 2019;38:356–79. https://doi.org/10.1002/mas.21586.
Yu A, Zhao J, Peng W, Banazadeh A, Williamson SD, Goli M, Huang Y, Mechref Y. Advances in mass spectrometry-based glycoproteomics. Electrophoresis. 2018;39:3104–22.
Martinez M, Renuse S, Kreimer S, O’Meally R, Natov P, Madugundu AK, Nirujogi RS, Tahir R, Cole R, Pandey A, Zachara NE. Quantitative proteomics reveals that the OGT interactome is remodeled in response to oxidative stress. Mol Cell Proteomic. 2021. https://doi.org/10.1016/J.MCPRO.2021.100069.
Hackett WE, Zaia J. Calculating glycoprotein similarities from mass spectrometric data. Mol Cell Proteomic. 2021;20: 100028. https://doi.org/10.1074/MCP.R120.002223.
Hage DS, Anguizola JA, Li R, Matsuda R, Papastavros E, Pfaunmiller E, Sobansky M, Zheng X. Affinity chromatography. In: Paul R, Poole C, Riekkola ML, editors. Liquid chromatography: fundamentals and instrumentation. 2nd edn. Amsterdam, Netherlands: Elsevier; 2017. p. 319–41.
Lee WC, Lee KH. Applications of affinity chromatography in proteomics. Anal Biochem. 2004;324:1–10.
McDowell CT, Klamer Z, Hall J, West CA, Wisniewski L, Powers TW, Angel PM, Mehta AS, Lewin DN, Haab BB, Drake RR. Imaging mass spectrometry and lectin analysis of n-linked glycans in carbohydrate antigen-defined pancreatic cancer tissues. Mol Cell Proteomic. 2021;20: 100012. https://doi.org/10.1074/MCP.RA120.002256.
Johannes L, Jacob R, Leffler H. Galectins at a glance. J Cell Sci. 2018. https://doi.org/10.1242/jcs.208884.
Low TY, Mohtar MA, Lee PY, Omar N, Zhou H, Ye M. Widening the bottleneck of phosphoproteomics: evolving strategies for phosphopeptide enrichment. Mass Spectrom Rev. 2021;40:309–33.
Riley NM, Coon JJ. Phosphoproteomics in the age of rapid and deep proteome profiling. Anal Chem. 2016;88:74–94.
Melo-Braga MN, Ibáñez-Vea M, Larsen MR, Kulej K. Comprehensive protocol to simultaneously study protein phosphorylation, acetylation, and N -linked sialylated glycosylation. Methods Mol Biol. 2014;1295:275–92.
Leney AC, El Atmioui D, Wu W, Ovaa H, Heck AJR. Elucidating crosstalk mechanisms between phosphorylation and O-GlcNAcylation. Proc Natl Acad Sci USA. 2017;114:E7255–61.
Diez IA, Govender I, Naicker P, Stoychev S, Jordaan J, Jensen O. Zirconium(IV)-IMAC for phosphopeptide enrichment in phosphoproteomics. BioRxiv. 2020. https://doi.org/10.1101/2020.04.13.038810.
Chen Z, Yu Q, Yu Q, Johnson J, Shipman R, Zhong X, Huang J, Asthana S, Carlsson C, Okonkwo O, Li L. In-depth site-specific analysis of N-glycoproteome in human cerebrospinal fluid and glycosylation landscape changes in Alzheimer’s disease. Mol Cell Proteomic. 2021;20: 100081. https://doi.org/10.1016/J.MCPRO.2021.100081.
Sun N, Wu H, Chen H, Shen X, Deng C. Advances in hydrophilic nanomaterials for glycoproteomics. Chem Commun. 2019;55:10359–75.
Jandera P, Janás P. Recent advances in stationary phases and understanding of retention in hydrophilic interaction chromatography. A review. Anal Chim Acta. 2017;967:12–32.
Jandera P. Stationary and mobile phases in hydrophilic interaction chromatography: A review. Anal Chim Acta. 2011;692:1–25.
Kozlik P, Goldman R, Sanda M. Hydrophilic interaction liquid chromatography in the separation of glycopeptides and their isomers. Anal Bioanal Chem. 2018;410:5001–8.
Ikegami T. Hydrophilic interaction chromatography for the analysis of biopharmaceutical drugs and therapeutic peptides: a review based on the separation characteristics of the hydrophilic interaction chromatography phases. J Sep Sci. 2019;42:130–213.
Ashwood C, Pratt B, Maclean BX, Gundry RL, Packer NH. Standardization of PGC-LC-MS-based glycomics for sample specific glycotyping. Analyst. 2019;144:3601–12.
Bapiro TE, Richards FM, Jodrell DI. Understanding the complexity of porous graphitic carbon (PGC) chromatography: modulation of mobile-stationary phase interactions overcomes loss of retention and reduces variability. Anal Chem. 2016;88:6190–4.
Wei J, Tang Y, Bai Y, Zaia J, Costello CE, Hong P, Lin C. Toward automatic and comprehensive glycan characterization by online PGC-LC-EED MS/MS. Anal Chem. 2020;92:782–91.
de Haan N, Yang S, Cipollo J, Wuhrer M. Glycomics studies using sialic acid derivatization and mass spectrometry. Nat Rev Chem. 2020;4:229–42.
Wu SW, Pu TH, Viner R, Khoo KH. Novel LC-MS2 product dependent parallel data acquisition function and data analysis workflow for sequencing and identification of intact glycopeptides. Anal Chem. 2014;86:5478–86.
Shah P, Wang X, Yang W, Eshghi ST, Sun S, Hoti N, Chen L, Yang S, Pasay J, Rubin A, Zhang H. Integrated proteomic and glycoproteomic analyses of prostate cancer cells reveal glycoprotein alteration in protein abundance and glycosylation. Mol Cell Proteomic. 2015;14:2753–63.
Park GW, Kim JY, Hwang H, Lee JY, Ahn YH, Lee HK, Ji ES, Kim KH, Jeong HK, Yun KN, Kim YS, Ko JH, An HJ, Kim JH, Paik YK, Yoo JS. Integrated glycoproteome analyzer (I-GPA) for automated identification and quantitation of site-specific N-glycosylation. Sci Rep. 2016. https://doi.org/10.1038/srep21175.
Liu MQ, Zeng WF, Fang P, Cao WQ, Liu C, Yan GQ, Zhang Y, Peng C, Wu JQ, Zhang XJ, Tu HJ, Chi H, Sun RX, Cao Y, Dong MQ, Jiang BY, Huang JM, Shen HL, Wong CCL, He SM, Yang PY. PGlyco 2.0 enables precision N-glycoproteomics with comprehensive quality control and one-step mass spectrometry for intact glycopeptide identification. Nat Commun. 2017;8:1–14.
Polasky DA, Yu F, Teo GC, Nesvizhskii AI. Fast and comprehensive N- and O-glycoproteomics analysis with MSFragger-Glyco. Nat Methods. 2020;17:1125–32.
Ou X, Liu Y, Lei X, Li P, Mi D, Ren L, Guo L, Guo R, Chen T, Hu J, Xiang Z, Mu Z, Chen X, Chen J, Hu K, Jin Q, Wang J, Qian Z. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun. 2020;11:1–12.
Walls AC, Tortorici MA, Snijder J, Xiong X, Bosch BJ, Rey FA, Veesler D. Tectonic conformational changes of a coronavirus spike glycoprotein promote membrane fusion. Proc Natl Acad Sci USA. 2017;114:11157–62. https://doi.org/10.1073/pnas.1708727114.
Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The proximal origin of SARS-CoV-2. Nat Med. 2020;26:450–2.
Miller LM, Barnes LF, Raab SA, Draper BE, El-Baba TJ, Lutomski CA, Robinson CV, Clemmer DE, Jarrold MF. Heterogeneity of glycan processing on trimeric SARS-CoV-2 spike protein revealed by charge detection mass spectrometry. J Am Chem Soc. 2021;143:56. https://doi.org/10.1021/jacs.1c00353.
Watanabe Y, Allen JD, Wrapp D, McLellan JS, Crispin M. Site-specific glycan analysis of the SARS-CoV-2 spike. Science. 2020;369:330–3.
Zhang Y, Zhao W, Mao Y, Chen Y, Wang S, Zhong Y, Su T, Gong M, Du D, Lu X, Cheng J, Yang H. Site-specific N-glycosylation characterization of recombinant SARS-CoV-2 spike proteins. Mol Cell Proteomic. 2021;20: 100058. https://doi.org/10.1074/MCP.RA120.002295.
Sanda M, Morrison L, Goldman R. N-and O-Glycosylation of the SARS-CoV-2 Spike Protein. Anal Chem. 2021;93:2003–9.
Shajahan A, Supekar NT, Gleinich AS, Azadi P. Deducing the N- And O-glycosylation profile of the spike protein of novel coronavirus SARS-CoV-2. Glycobiology. 2020;30:981–8.
Xu W, Wang M, Yu D, Zhang X. Variations in SARS-CoV-2 spike protein cell epitopes and glycosylation profiles during global transmission course of COVID-19. Front Immunol. 2020;11: 565278. https://doi.org/10.3389/fimmu.2020.565278.
Brun J, Vasiljevic S, Gangadharan B, Hensen M, Chandran AV, Hill ML, Kiappes JL, Dwek RA, Alonzi DS, Struwe WB, Zitzmann N. Analysis of SARS-CoV-2 spike glycosylation reveals shedding of a vaccine candidate. bioRxiv. 2020. https://doi.org/10.1101/2020.11.16.384594.
Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. 2020;181:281-292.e6.
Walls AC, Xiong X, Park YJ, Tortorici MA, Snijder J, Quispe J, Cameroni E, Gopal R, Dai M, Lanzavecchia A, Zambon M, Rey FA, Corti D, Veesler D. Unexpected receptor functional mimicry elucidates activation of coronavirus fusion. Cell. 2019;176:1026-1039.e15.
Bouwman KM, Tomris I, Turner HL, van der Woude R, Shamorkina TM, Bosman GP, Rockx B, Herfst S, Snijder J, Haagmans BL, Ward AB, Boons GJ, de Vries RP. Multimerization- And glycosylation-dependent receptor binding of SARS-CoV-2 spike proteins. PLoS Pathog. 2021. https://doi.org/10.1371/JOURNAL.PPAT.1009282.
Yan R, Zhang Y, Li Y, Xia L, Guo Y, Zhou Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science. 2020;367:1444–8.
Yang Q, Hughes TA, Kelkar A, Yu X, Cheng K, Park SJ, Huang WC, Lovell JF, Neelamegham S. Inhibition of SARS-CoV-2 viral entry upon blocking N-and O-glycan elaboration. Elife. 2020;9:1–44.
Mahmud I, Garrett TJ. Mass spectrometry techniques in emerging pathogens studies: COVID-19 perspectives. J Am Soc MASS Spectrom. 2020;31:2013–24.
Grubaugh ND, Hanage WP, Rasmussen AL. Making sense of mutation: what D614G means for the COVID-19 pandemic remains unclear. Cell. 2020;182:794–5.
Mao R, Liang J, Shen J, Ghosh S, Zhu LR, Yang H, Wu KC, Chen MH. Implications of COVID-19 for patients with pre-existing digestive diseases. Lancet Gastroenterol Hepatol. 2020;5:426–8.
Michalski A, Cox J, Mann M. More than 100,000 detectable peptide species elute in single shotgun proteomics runs but the majority is inaccessible to data-dependent LC-MS/MS. J Proteome Res. 2011;10:1785–93.
Baerenfaenger M, Meyer B. Intact human alpha-acid glycoprotein analyzed by ESI-qTOF-MS: simultaneous determination of the glycan composition of multiple glycosylation sites. J Proteome Res. 2018;17:3693–703.
Liou TG, Adler FR, Cahill BC, Cox DR, Cox JE, Grant GJ, Hanson KE, Hartsell SC, Hatton ND, Helms MN, Jensen JL, Kartsonaki C, Li Y, Leung DT, Marvin JE, Middleton EA, Osburn-Staker SM, Packer KA, Shakir SM, Sturrock AB, Tardif KD, Warren KJ, Waddoups LJ, Weaver LJ, Zimmerman E, Paine R. SARS-CoV-2 innate effector associations and viral load in early nasopharyngeal infection. Physiol Rep. 2021;9(4):14761.
Sims JT, Krishnan V, Chang C-Y, Engle SM, Casalini G, Rodgers GH, Bivi N, Nickoloff BJ, Konrad RJ, de Bono S, Higgs RE, Benschop RJ, Ottaviani S, Cardoso A, Nirula A, Corbellino M, Stebbing J. Characterization of the cytokine storm reflects hyperinflammatory endothelial dysfunction in COVID-19. J Allergy Clin Immunol. 2021;147:107–11.
Park J, Kim H, Kim SY, Kim Y, Lee J-S, Dan K, Seong M-W, Han D. In-depth blood proteome profiling analysis revealed distinct functional characteristics of plasma proteins between severe and non-severe COVID-19 patients. Sci Rep. 2020. https://doi.org/10.1038/s41598-020-80120-8.
Chen Y, Zheng Y, Yu Y, Wang Y, Huang Q, Qian F, Sun L, Song ZZ, Chen Z, Feng J, An Y, Yang J, Su Z, Sun S, Dai F, Chen Q, Lu Q, Li P, Ling Y, Yang Z, Tang H, Shi L, Jin L, Holmes EC, Ding C, Zhu TT, Zhang Y. Blood molecular markers associated with COVID-19 immunopathology and multi-organ damage. EMBO J. 2020;39:1–23.
Zeng HL, Chen D, Yan J, Yang Q, Han QQ, Li SS, Cheng L. Proteomic characteristics of bronchoalveolar lavage fluid in critical COVID-19 patients. FEBS J. 2020. https://doi.org/10.1111/febs.15609.
Hou X, Zhang X, Wu X, Lu M, Wang D, Xu M, Wang H, Liang T, Dai J, Duan H, Xu Y, Yu X, Li Y. Serum protein profiling reveals a landscape of inflammation and immune signaling in early-stage COVID-19 infection. Mol Cell Proteomic. 2020;19:1749–59.
Lei Q, Li Y, Hou H, Wang F, Ouyang Z, Zhang Y, Lai D, Banga Ndzouboukou J-L, Xu Z, Zhang B, Chen H, Xue J, Lin X, Zheng Y, Yao Z, Wang X, Yu C, Jiang H, Zhang H, Qi H, Guo S, Huang S, Sun Z, Tao S, Fan X. Antibody dynamics to SARS-CoV-2 in asymptomatic COVID-19 infections. Allergy. 2021;76:551–61.
Wendt R, Kalbitz S, Luebbert C, Kellner N, Macholz M, Schroth S, Ermisch JJJ, Latosisnka A, Arnold B, Mischak H, Hartmut Beige J, Metzger J, Lübbert C, Kellner N, Macholz M, Schroth S, Ermisch JJJ, Latosisnka A, Arnold B, Mischak H, Hartmut Beige J, Metzger J. Urinary peptides significantly associate with COVID-19 severity: pilot proof-of-principle data and design of a multicentric diagnostic study. Proteomics. 2020;20:2000202. https://doi.org/10.1002/pmic.202000202.
Messner CB, Demichev V, Wendisch D, Michalick L, White M, Freiwald A, Textoris-Taube K, Vernardis IS, Egger A-S, Kreidl M, Ludwig D, Kilian C, Agostini F, Zelezniak A, Thibeault C, Pfeiffer M, Hippenstiel S, Hocke A, von Kalle C, Campbell A, Hayward C, Porteous DJ, Marioni RE, Langenberg C, Lilley KS, Kuebler WM, Muelleder M, Drosten C, Suttorp N, Witzenrath M, Kurth F, Sander LE, Ralser M. Ultra-High-Throughput Clinical Proteomics Reveals Classifiers of COVID-19 Infection. CELL Syst. 2020;11:11+.
Haljasmägi L, Salumets A, Rumm AP, Jürgenson M, Krassohhina E, Remm A, Sein H, Kareinen L, Vapalahti O, Sironen T, Peterson H, Milani L, Tamm A, Hayday A, Kisand K, Peterson P. Longitudinal proteomic profiling reveals increased early inflammation and sustained apoptosis proteins in severe COVID-19. Sci Rep. 2020;10:20533.
Liu X, Cao Y, Fu H, Wei J, Chen J, Hu J, Liu B. Proteomics analysis of serum from COVID-19 patients. ACS Omega. 2021;6:7951–8.
D’Alessandro A, Thomas T, Dzieciatkowska M, Hill RC, Francis RO, Hudson KE, Zimring JC, Hod EA, Spitalnik SL, Hansen KC. Serum proteomics in COVID-19 patients: altered coagulation and complement status as a function of IL-6 level. J Proteome Res. 2020;19:4417–27.
Wu M, Chen Y, Xia H, Wang C, Tan CY, Cai X, Liu Y, Ji F, Xiong P, Liu R, Guan Y, Duan Y, Kuang D, Xu S, Cai H, Xia Q, Yang D, Wang MW, Chiu IM, Cheng C, Ahern PP, Liu L, Wang G, Surana NK, Xia T, Kasper DL. Transcriptional and proteomic insights into the host response in fatal COVID-19 cases. Proc Natl Acad Sci USA. 2020;117:28336–43.
Acknowledgements
This study was funded by SZSTC (SGDX20190816230207535) and obtained a donation from Kwok Chung Bo Fun Charitable Fund, which commemorates the establishment of the Kwok Yat Wai Endowed Chair of Environmental and Biological Analysis. All authors of this article contributed separately as follows: Li Zhong draft the manuscript, Lin Zhu revised the manuscript, and Zong-Wei Cai supervised the whole study and provided financial supports.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors declared no interest conflicts.
About this article
Cite this article
Zhong, L., Zhu, L. & Cai, ZW. Mass Spectrometry-based Proteomics and Glycoproteomics in COVID-19 Biomarkers Identification: A Mini-review. J. Anal. Test. 5, 298–313 (2021). https://doi.org/10.1007/s41664-021-00197-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41664-021-00197-6