Abstract
Growing evidence points to the role of epigenetic mechanisms, including DNA methylation, in substance use and addiction. We conducted a systematic review of 47 recent (2012–2015) animal and human studies that investigate DNA methylation and substance use/exposure, spanning preconception to adulthood. The majority of extant studies (i) focused on exposure during adulthood, (ii) examined the effects of alcohol use, (iii) employed a candidate gene approach, and (iv) were cross-sectional. While studies generally support an association between substance use/exposure and DNA methylation and also suggest that developmental context and timing matter, a dearth of longitudinal data and low comparability across studies currently limits the conclusions that can be drawn. Future challenges and directions for the field are discussed.
Similar content being viewed by others
Introduction
Addiction to psychoactive substances (e.g., alcohol, illicit drugs) is a debilitating condition characterised by compulsive drug-seeking and repeated harmful use, despite adverse consequences [1]. Like other complex diseases, addiction results from both genetic and environmental factors, which combine to exert additive, evocative, and interactive effects across the lifespan [2•]. How such gene-environment associations operate at a molecular level, however, remains unclear. In recent years, epigenetic mechanisms have been proposed as a potential candidate, as they respond to both genetic and environmental influences [3, 4••] and are thought to mediate vulnerability to disorders, including addiction [3, 4••].
Epigenetic mechanisms, such as DNA methylation (DNAm) regulate when and where genes are expressed without changing the DNA sequence itself [5]. DNAm refers to the addition of a methyl group, primarily in the context of cytosine-guanine (CpG) dinucleotides. The genome contains an excess of 28 million CpG sites, around 10 % of which cluster into CpG ‘islands’, close to gene promoter regions [6]. Methylated CpG islands impede transcription factors from accessing the DNA sequence. As such, DNAm is typically associated with decreased gene expression, although the functional role of methylation changes within genomic regions other than CpG islands (e.g., intergenic regions) remains unclear [7•]. Importantly, DNAm is dynamic across the lifespan—although patterns are mitotically stable, which can lead to long-term alterations in gene activity, they also show a considerable degree of flexibility over time, enabling cells to respond to changing internal and external inputs [8].
A growing number of studies have begun to clarify the role of DNAm in substance abuse and addiction. Experimental studies in animals have led the way, documenting a number of important findings. First, substance use can alter DNAm—for example, repeated administration of substances (e.g., alcohol, cocaine) has been found to modify methylation patterns in the reward regions of the brain (e.g., striatum [9]). Second, DNAm contributes to the pathophysiology of addiction. Specifically, drug-induced methylation changes have been shown to influence the expression of genes involved in synaptic plasticity and memory consolidation, which in turn drive long-term neuroadaptations underlying the onset and persistence of addictive behaviours [4••]. Third, animal studies have begun to shed light on the role of developmental context on DNAm and addiction risk. For example, alcohol intake during the first half of pregnancy has been found to alter epigenetic patterns in the developing embryo, leading to reduced fetal growth and congenital abnormalities similar to those observed in human fetal alcohol syndrome, as well as subsequent risk for addiction [10].
So far, studies in humans have provided initial support for animal findings, reporting methylomic differences between substance abusers and drug-free controls across a number of substances and tissue types [9, 11••]. However, unlike animal studies that make it possible to experimentally manipulate the type, extent, and timing of substance exposure, studies in humans have been primarily cross-sectional and correlational, making the causal links between epigenetic changes and subsequent addiction more problematic to draw.
The aim of this systematic review is three-fold: (i) to collate findings from recent animal and human research investigating the link between substance exposure, DNAm, and addiction; (ii) to consider the relevance of timing of substance exposure, beginning in preconception through to adulthood; and (iii) to outline future directions for the field.
Methods
Inclusion Criteria
We included studies that investigated associations between DNAm and substance use/exposure. In line with the journal’s focus on current research, we only included articles published during the past 3 years (1 January 2012 to 31 February 2015). No restriction was applied regarding (i) species (e.g., human, mouse), (ii) period of exposure (e.g., prenatal, adulthood), (iii) substance (e.g., alcohol, cocaine), (iv) tissue (e.g., blood, brain), (v) approach (e.g., candidate vs genome-wide), and (vi) design (e.g., cross-sectional vs longitudinal).
Search Strategy
PubMed and PsychInfo were searched for relevant studies written in English. Search terms were applied in MeSH or index terms, as well as text words. Included terms related to either (i) DNA methylation (e.g., methylat*; epigen*), or (ii) substance (e.g., substance use, abuse, dependence; drug; addiction; cocaine; heroin; cannabis; alcohol; opiate*; smoking; tobacco). ‘Cancer’ and ‘medication’ were specified as exclusion terms to avoid studies investigating DNAm in relation to clinical drugs.
Study Selection
Our search yielded 621 records, with 381 remaining after filtering out duplicates (see Fig. 1). Titles and abstracts were screened, and studies were excluded if they were not empirical (e.g., reviews), focussed on epigenetic mechanisms other than DNAm (e.g., histone modifications), or examined drugs other than the ones specified above (e.g., clinical drugs). Given that the majority of DNAm studies on tobacco use examined medical diseases (e.g., cancer) as opposed to addiction-relevant phenotypes, studies with tobacco were not included in the review. Sixty-one studies were retained, and their full text articles were assessed for eligibility. Sixteen articles were removed due to the following reasons: (i) six did not include DNAm data; (ii) six did not report direct associations between DNAm and substance use/exposure; (iii) two did not include substance data; (iv) one was published before 2012; and (v) one was based on cell culture data. A total of 45 original reports were therefore included in the systematic review.
Results
Descriptive Summary
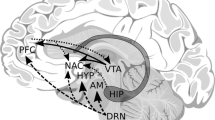
Study characteristics are summarised in Table 1 (see also Fig. 2). Twenty-four studies examined animal samples (n rat = 14 and n mouse = 10) and 21 examined humans. The majority of studies investigated substance exposure during adulthood (n = 33), focused on alcohol (n = 36), involved peripheral samples (n = 25), examined DNAm at a single time point (n = 40), and used a candidate gene approach (n = 26). The most common peripheral tissue examined was blood (n = 18), followed by liver, sperm, pancreas, saliva, placenta, kidney, intestine, and colon. Most commonly examined central tissues were prefrontal cortex (n = 5) and hippocampus (n = 4), followed by nucleus accumbens, hypothalamus, amygdala, cerebellum, ventral tegmental area, striatum, and neocortex. Below, we describe findings first in animals and then in humans, in order of developmental period of substance exposure.
Flowchart of study characteristics. N.B. The same study may fit into multiple categories. Grey-shaded boxes represent developmental periods that have not been investigated. For each level (1–5), boxes are shaded according to study frequency (e.g., the least frequently investigated tissue is represented by lighter shading, while the most commonly investigated tissue is represented by darker shading). Boxes marked with an asterisk indicate presence of longitudinal studies with repeated measures of DNA methylation (here, boxes indicate the number of longitudinal studies out of the total number of studies). Meth methamphetamine, C central tissue (brain), P peripheral tissue, Ca candidate gene, Ew epigenome-wide, Gl global
Animal Studies
Preconception
Three candidate gene studies investigated parental alcohol use prior to conception. In the first, paternal consumption in mice related to decreased DNAm of Bdnf—implicated in stress response and neural development—in paternal sperm cells and offspring ventral tegmental area [12]. In the second, paternal alcohol consumption in rats was associated with increased Pomc methylation (another gene relevant in stress response) within both parental sperm and offspring hypothalamus, although findings were specific to the male germline [13]. In contrast, the third study [14] found that DNAm in H19 CTCF binding sites—involved in imprinting mechanisms—was reduced in offspring tail blood but not in paternal sperm cells.
Prenatal
Three studies from the same working group found that prenatal alcohol exposure associated with increased Pomc methylation in the rat hypothalamus, which in turn related to decreased gene expression [13, 15, 16]. These changes were maintained transgenerationally (up to three generations), but could be rescued by gestational choline supplementation. Epigenome-wide associations between prenatal alcohol exposure and DNAm in brain tissue from adult offspring were identified by one study, particularly within genetic pathways related to nervous system development (including the Cdk5 signalling pathway) and neurological diseases, including the Alzheimer’s disease-linked gene App [17]. Another study found that in utero, exposure to methamphetamine was associated with aberrant hippocampal DNAm in adolescent mice offspring [18]. Hypermethylated genetic pathways related to cerebral cortex GABAergic interneuron differentiation, while hypomethylated pathways related to embryonic development.
Neonatal
Two mouse studies investigated the effect of neonatal alcohol exposure on global methylation within the hippocampus and neocortex [19, 20]. While the first study reported a reduction in global methylation in response to acute alcohol exposure (8 and 24-h postexposure; [20]), the second study observed an increase in global methylation in the exposed group across both regions, which could be partially ameliorated by choline treatment [21]. Although of interest, it is important to note that neonatal substance exposure may be less relevant to human studies compared to other developmental periods, as it is relatively uncommon in humans.
Adulthood
This developmental period received by far the greatest research attention (71 % of animal studies, n = 17) and was primarily examined in relation to alcohol exposure (n = 11). In global methylation studies, exposed mice were found to have lower DNAm in the cerebral cortex [21] but not in liver [22], although reductions were reported in global DNA hydroxymethylation (another type of DNA modification, characterised by the addition of a hydroxymethyl group). Findings from candidate gene studies further indicated that alcohol exposure in adulthood is associated with increased DNAm in multiple genes, including the serotonin receptor Htr3a in blood and hippocampal tissue [23], the sodium transporter Slc5a6 gene in pancreatic tissue [24], and immune function TLR-pathway genes in the liver [25]. Decreased DNAm was instead observed for the glutamate gene Nr2b in the prefrontal cortex [26] and Bdnf gene in motile sperm [12]. Tissue- and gene-specific DNAm alterations were identified by one study in folate-regulating genes [27]. Finally, no associations were found in opioid-related genes Pdny and Pnoc in the rat amygdala [28], as well as imprinting-control genes H19 and Rasgfl in mouse sperm [14].
Seven studies examined DNAm in relation to cocaine and/or opiate exposure. No associations with global methylation were reported in the corpus callosum of cocaine-exposed rats after 1 or 30 days of forced abstinence [29], as well as cocaine or heroin-exposed mice [30]—although a specific reduction in hydroxymethylation in the liver following cocaine administration was reported within the same sample [31]. Drug- and tissue-specific effects were also identified in the study by Tian et al. [32] where global DNAm reductions were evident in the prefrontal cortex (but not in the nucleus accumbens) of mice exposed to cocaine (not heroin)—an effect that was reversible through repeated administration of methionine. With regard to candidate genes, increased Drd2 receptor methylation was observed in the nucleus accumbens of rats exposed to glucocorticoids in utero [33]. This association was specific to morphine administration and reversed by L-dopa treatment. Also in the nucleus accumbens, repeated SAM pretreatment was found to modify cocaine-induced methylation changes in the neuropeptides Cck and Gal, as well as the glutamate transporter Slc17a7 [34]. Pol Bodetto et al. [35] reported that methylation of Pp2cβ, a gene involved in cellular function, was higher in the brain of cocaine-exposed rats versus controls. Finally, in a study investigating myelin-producing genes, reduced mean DNAm of Sox10 was identified in the corpus callosum of cocaine-exposed rats, particularly after a period of forced abstinence [29]. None of the studies examined epigenome-wide alterations in response to adult substance exposure.
Human Studies
Prenatal
Two studies examined DNAm in relation to prenatal alcohol exposure. Wilhelm-Benartzi et al. [36] found that maternal alcohol intake positively associated with global LINE-1 (but not with AluYb8) methylation in placental tissue. One candidate gene study found that cord blood methylation of the developmental gene ZAC1 positively correlated with prenatal maternal alcohol intake as well as associating with reduced fetal and postnatal weight [37].
Adolescence
Only one study examined adolescent substance use. Researching the impact of cannabis smoking on whole blood COMT methylation (important for neurotransmitter catalysis), van der Knaap et al. [38] found no main effect of cannabis use. However, a significant methylation by genotype interaction was identified, where Met/Met carriers with higher DNAm were least likely to be frequent cannabis users.
Adulthood
Eighty-five per cent of studies in humans (n = 18) examined adult substance use, again focusing primarily on alcohol exposure (n = 16). One global methylation study found decreased DNAm in the blood of alcohol drinkers (Alu, not LINE-1 [39]). Results contrast those of increased global methylation identified in the frontal cortex of HIV+ methamphetamine users versus non users [40], as well as in the blood of methadone-substituted former opiate addicts, an effect which was also replicated in independent sample of opioid-treated patients [41].
Candidate gene studies focused mainly on genes involved in neural function, most likely guided by existing neurochemical data regarding addiction on animals and humans. Higher DNAm was observed in the blood of alcohol-dependent individuals within the HTR3A serotonin receptor gene [42] and OPRM1 opioid receptor gene [43]—an association that was also identified in opiate addicts [41]. Lower DNAm of the leptin hormone (LEP) gene was instead identified in the blood of patients with stronger alcohol cravings [44]. No significant associations were reported between alcohol use and DNAm in a number of genes, including PDNY and PNOC opioid-related genes (blood; [42]), the serotonin transporter 5-HTT in females exposed to trauma (alcohol, cannabis; [45]), the DAT dopamine transporter in blood [46], and the drug metabolism gene UGT1A1 in human liver [47]. Of the candidate gene studies reviewed, two featured repeated measures of DNAm, comparing alcohol-dependent cases versus controls at baseline, day 7 and day 14 posttreatment admission. While the first found significant differences in DNAm of volume-regulating neuropeptides AVP and ANP both at baseline and between day 7 and 14 of withdrawal [48], the second [49] reported increased nerve growth factor (NGF) methylation in cases versus controls, but only between day 7 and 14.
All epigenome-wide investigations focussed on the effect of alcohol in blood. Generally, results were mixed, depending on sample characteristics and methods. In terms of specific genes, two epigenome-wide association studies (EWASs) confirmed associations with alcohol metabolism-related genes, including alcohol and aldehyde dehydrogenases (ADH1A, ADH7, ALDH3B2, ALDH1A2) and cytochrome P450 2A13 [50, 51]. In one study [52], the tumour suppressor gene BLCAP and ABR—involved in vestibular morphogenesis—were hypomethylated in heavy alcohol drinkers versus abstinent controls, suggesting one mechanism by which tumour risk may be higher in alcohol drinkers. In another study, alcohol-dependent discordant siblings showed hypomethylation of SSTR4—an important gene for hormonal function—and hypermethylation of the GABA receptor gene GABRP [53]. Finally, two EWASs measured DNAm at multiple time points: before and after a 25-day treatment programme [54], or a 12-year interim period [55]. While the former [54] found no significant differences pre-vs-post treatment, the latter [55] observed a general increase in methylation with alcohol consumption over a 12-year period, particularly in CKM, PHOX2A, and NPDC1. With regard to wider biological pathways, EWAS studies indicated that the most common pathways that were hypermethylated in response to alcohol use were those related to G-protein mediated and GTPase signal transduction processes [51, 54, 55], whereas pathways associated with stimulus and stress responses, as well as immune and inflammatory processes, where likely to be hypomethylated [51]. Hypomethylation was also observed in long terminal repeat (LTR) regions of retrotransposons in the superior frontal cortex of postmortem alcohol users [56]. Other important pathways related to apoptosis [52, 54, 55], metabolism [53], as well as GABA and dopamine systems [40, 53].
Discussion
The aim of the present review was to summarise the latest animal and human research investigating the association between substance use, DNA methylation, and addiction risk, spanning preconception to adulthood. Based on the 45 reports included, we may conclude that there is preliminary support for a link between substance exposure, DNAm, and addiction. However, findings are often mixed and have limited comparability. In this section, we review key similarities and differences across studies, evaluate evidence for the importance of timing of substance exposure, and outline future directions for the field.
Summary of Study Characteristics and Findings
The majority of studies across species focused on substance exposure during adulthood, examined the effects of alcohol, employed a candidate gene approach, and were cross-sectional. One key difference related to tissue, with animal studies most often investigating brain samples and human studies examining DNAm in blood. Global methylation studies were more common in rodents, while epigenome-wide studies were more frequently carried out in humans. Although prospective longitudinal designs were more common in animal studies, the only four reports to include repeated DNAm measures were based on humans.
As a symptom of how young the field is, there are not sufficient data to assess how DNAm of specific genes relates to exposure to specific substances across developmental periods and tissue types. We do, however, highlight five genes that were investigated by multiple studies and, promisingly, showed a consistent direction of associations. In three rodent studies from the same working group [13, 15, 16], increased methylation and decreased expression of Pomc—a gene implicated in stress response, metabolism, and immune function—was observed in response to prenatal alcohol exposure across multiple tissues. These results highlight one mechanism through which fetal alcohol programming can occur, contributing to HPA axis dysregulation and increased addiction risk [16]. In two other studies, the opioid receptor mu 1 (OPRM1)—a gene important for mediating drug-induced activation of reward pathways—was hypermethylated in the blood of former opiate addicts [41] and alcohol-dependent individuals [43]. It was not possible to establish, however, whether higher methylation was a predisposing factor for addiction and/or a consequence of long-term substance use. Furthermore, hypermethylation of the serotonin receptor 3A (HTR3A) was identified in relation to alcohol exposure across both humans [42] and rodents [23]. Finally, a null association between alcohol exposure and methylation in the opioid signaling genes PDNY and PNOC was reported in human blood [42] and brain tissue in rats [28]. Despite these consistent findings, it is noteworthy that genes investigated by candidate studies did not typically converge with those identified by studies using hypothesis-free, epigenome-wide analyses. Instead, EWAS studies more often reported significant associations with drug metabolizing genes, as well as highlighting wider biological pathways linked to substance exposure, including signal transduction, inflammation, and apoptosis, in addition to stress response and neurotransmission. How these pathways specifically contribute to addiction, however, remains unclear.
Given the limited comparability across studies, we were not able to systematically assess the importance of developmental context in the relationship between substance use, DNAm, and addiction risk. However, the studies reviewed did provide preliminary support for the relevance of timing of substance exposure on DNAm. For example, evidence from animal models demonstrated that substance exposure can influence DNAm even prior to conception, supporting the existence of transgenerational effects [12–14]. Studies also pointed to the prenatal period as a particularly sensitive developmental window. For example, in utero substance exposure influenced DNAm of developmental genes, which in turn affected postnatal outcomes (e.g., reduced postnatal weight [37]), although the relevance of these changes for addiction risk is yet to be characterised. It is important to note that the period between birth and adulthood received very little attention. In fact, none of the studies investigated childhood and only one examined adolescence—a key period of vulnerability for the development of substance use disorders [57].
Current Challenges for the Field
Despite considerable advances in epigenetic research, studies investigating the role of DNAm in substance use and addiction continue to face a number of key challenges [11••, 58••]. Firstly, our knowledge of the methylome is still limited. Because we know little about ‘typical’ methylation patterns in humans, it is difficult to establish when such patterns deviate to contribute to diseased states. This is complicated by the fact that DNAm patterns can vary across multiple factors, including tissue, cell type, sex, and age [59•]. In general, the compilation of reference datasets will be important for providing a ‘typical’ benchmark against which to compare epigenetic findings. Knowledge is also limited regarding the relative contribution of genetic and environmental influences on observed methylation patterns, which will require the use of genetically informative designs, such as twin studies and studies identifying methylation quantitative trait loci [60, 61]. More work will also be needed to determine the functional significance of identified DNAm changes at transcriptomic, metabolomic, proteomic, and neural biological levels.
A second set of challenges relates to research methodology. Methods have varied widely across studies, including differences in preprocessing, quality control, genomic coverage, data analysis, choice of covariates and significance thresholds used for detecting effects. Together, these sources of variability have limited comparability across studies and complicated efforts to replicate findings—a necessary step for weeding out false positives. The increased availability of standardised pipelines will considerably help in this respect [62•]. Furthermore, the integration of discovery and replicate samples will become increasingly important, as was the case for genomic studies. More research will also be needed to determine what sample sizes are required to reach appropriate statistical power, although simulation-based studies are beginning to provide recommendations [63].
A third issue relates to difficulties in establishing causal relationships between substance use, DNAm, and addiction. Most of the studies reviewed adopted a cross-sectional approach with DNAm data sampled at only one time point. Human studies, in particular, focussed primarily on adults who had already been exposed to substances. As such, it remains unclear whether DNAm can be a risk factor for, as well as a consequence of substance use, and how substance exposure and DNAm interrelate over time to influence addiction risk.
Below, we propose a number of ways in which future research may strengthen causal inferences and improve understanding of the role of DNAm in substance exposure and addiction.
A Proposed Model for Conducting Research on DNA Methylation, Substance Use, and Addiction
Firstly, it will be important to capitalise on the strengths of animal models to clearly delineate the mechanisms linking substance exposure, DNAm, and addiction. Systematic investigations will need to be conducted within a given substance, across multiple tissues, over developmental periods, and in different strains. Importantly, it will be necessary to collect prospective, repeated measures of DNAm pre- and post-substance exposure, in order to (i) investigate whether preexposure DNAm predicts individual differences in drug-seeking behaviours, (ii) trace the timing and stability of DNAm changes following exposure, and (iii) clarify whether DNAm mediates the effect of substance use on the onset and persistence of addiction. The sampling of multiple tissues over time will also make it possible to establish cross-tissue variability and locate peripheral biomarkers that most closely resemble DNAm changes in neural networks underlying addiction. Furthermore, incorporating additional omics data, such as gene expression, serum levels, protein content, and enzymatic activity, will be useful for clarifying the functional relevance of observed DNAm changes at multiple biological levels [e.g., 13, 16]. Importantly, the use of methyl-modifying agents (e.g., methionine, choline [15, 32]) will offer valuable opportunities for testing the reversibility of drug-induced DNAm changes, identifying whether certain developmental periods are more sensitive to intervention, and examining whether normalisation of DNAm patterns parallel changes in addiction-relevant phenotypes. Finally, the availability of methylomic data in relation to different substances will make it possible to disentangle DNAm markers that are common to multiple substances (perhaps reflecting a general liability to addiction) as opposed to substance-specific markers.
The knowledge generated from animal research could then be used to inform the design of human studies and to map out the most promising DNAm markers for further investigation. This will require, however, the use of strategies to maximise cross-species comparability. For example, the use of data from epidemiological birth cohorts that feature repeated measures of DNA [e.g., 64], could allow researchers to examine whether preexposure versus postexposure DNAm changes identified in longitudinal animal studies extend to humans. Furthermore, analytic methods that make it possible to integrate repeated measures of environmental exposure (e.g., substance use), DNAm, and phenotypic outcomes (e.g., addiction)—such as structural equation modelling—will be particularly useful for tracing longitudinal associations and for testing mediation hypotheses [65]. The development of advanced causal inference methods, such as the two-step epigenetic Mendelian randomisation [66, 67], may also show promise for testing causal pathways documented in animals. As discussed above, the inclusion of additional biological markers (e.g., serum levels) will be necessary for establishing the functional relevance of identified DNAm markers. Furthermore, given the scarce availability of central tissues in human research (i.e., postmortem), it will be important in the future to investigate whether peripheral DNAm markers can be related to in vivo structural and functional brain data (e.g., striatal activity when viewing addiction-related cues). Finally, increased use of approaches that capitalise on co-methylation patterns between CpG sites, such as regional or network-based approaches, will be important for reducing multiple testing and increasing power to detect effects in humans, enabling to move beyond individual methylation sites toward wider biological systems [68].
Conclusions
DNA methylation is emerging as an important molecular mechanism mediating substance use response and addiction risk. However, the limited understanding of the epigenome, heterogeneity across studies, a reliance on cross-sectional designs, and lack of replications make it difficult to interpret the relevance of the extant data for mechanisms of addiction. Rapid developments in knowledge, methodology, and research designs will offer exciting opportunities for delineating the role of DNAm in the pathophysiology of addiction, as well as testing its potential clinical utility as an exposure indicator, disease biomarker, and therapeutic target (Table 2).
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
McQuown SC, Wood MA. Epigenetic regulation in substance use disorders. Curr psychiatry rep. 2010;12(2):145–53. doi:10.1007/s11920-010-0099-5.
Kendler KS, Chen X, Dick D, Maes H, Gillespie N, Neale MC, et al. Recent advances in the genetic epidemiology and molecular genetics of substance use disorders. Nat Neurosci. 2012;15(2):181–9. doi:10.1038/nn.3018. Provides a comprehensive overview of genetic influences on substance use disorders.
Tsankova N, Renthal W, Kumar A, Nestler EJ. Epigenetic regulation in psychiatric disorders. Nat Rev Neurosci. 2007;8(5):355–67. doi:10.1038/nrn2132.
Nestler EJ. Epigenetic mechanisms of drug addiction. Neuropharmacology. 2014;76(Pt B):259–68. doi:10.1016/j.neuropharm.2013.04.004. Important for conveying a mechanistic understanding of the role of epigenetics in drug addiction.
Jaenisch R, Bird A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet. 2003;33(Suppl):245–54. doi:10.1038/ng1089.
Eckhardt F, Lewin J, Cortese R, Rakyan VK, Attwood J, Burger M, et al. DNA methylation profiling of human chromosomes 6, 20, and 22. Nat Genet. 2006;38(12):1378–85. doi:10.1038/ng1909.
Jones PA. Functions of DNA methylation: islands, start sites, gene bodies, and beyond. Nat Rev Genet. 2012;13(7):484–92. doi:10.1038/nrg3230. Comprehensive overview of the epigenetic process of DNA methylation, including physical properties, functions, and role in development and disease.
Alabert C, Groth A. Chromatin replication and epigenome maintenance. Nat Rev Mol Cell Biol. 2012;13(3):153–67. doi:10.1038/nrm3288.
Wong CC, Mill J, Fernandes C. Drugs and addiction: an introduction to epigenetics. Addiction (Abingdon, England). 2011;106(3):480–9. doi:10.1111/j.1360-0443.2010.03321.x.
Kaminen-Ahola N, Ahola A, Maga M, Mallitt KA, Fahey P, Cox TC, et al. Maternal ethanol consumption alters the epigenotype and the phenotype of offspring in a mouse model. PLoS Genet. 2010;6(1), e1000811. doi:10.1371/journal.pgen.1000811.
Harlaar N, Hutchison KE. Alcohol and the methylome: design and analysis considerations for research using human samples. Drug Alcohol Depend. 2013;133(2):305–16. doi:10.1016/j.drugalcdep.2013.07.026. Systematic review of existing human studies examining DNA methylation and alcohol use, including current challenges and considerations for future research.
Finegersh A, Homanics GE. Paternal alcohol exposure reduces alcohol drinking and increases behavioral sensitivity to alcohol selectively in male offspring. PLoS One. 2014;9(6), e99078. doi:10.1371/journal.pone.0099078.
Govorko D, Bekdash RA, Zhang C, Sarkar DK. Male germline transmits fetal alcohol adverse effect on hypothalamic proopiomelanocortin gene across generations. Biol Psychiatry. 2012;72(5):378–88. doi:10.1016/j.biopsych.2012.04.006.
Knezovich JG, Ramsay M. The effect of preconception paternal alcohol exposure on epigenetic remodeling of the h19 and rasgrf1 imprinting control regions in mouse offspring. Front Genet. 2012;3:10. doi:10.3389/fgene.2012.00010.
Bekdash RA, Zhang C, Sarkar DK. Gestational choline supplementation normalized fetal alcohol-induced alterations in histone modifications, DNA methylation, and proopiomelanocortin (POMC) gene expression in beta-endorphin-producing POMC neurons of the hypothalamus. Alcohol Clin Exp Res. 2013;37(7):1133–42. doi:10.1111/acer.12082.
Gangisetty O, Bekdash R, Maglakelidze G, Sarkar DK. Fetal alcohol exposure alters proopiomelanocortin gene expression and hypothalamic-pituitary-adrenal axis function via increasing MeCP2 expression in the hypothalamus. PLoS One. 2014;9(11), e113228. doi:10.1371/journal.pone.0113228.
Laufer BI, Mantha K, Kleiber ML, Diehl EJ, Addison SM, Singh SM. Long-lasting alterations to DNA methylation and ncRNAs could underlie the effects of fetal alcohol exposure in mice. Dis Model Mech. 2013;6(4):977–92. doi:10.1242/dmm.010975.
Itzhak Y, Ergui I, Young JI. Long-term parental methamphetamine exposure of mice influences behavior and hippocampal DNA methylation of the offspring. Mol Psychiatry. 2015;20(2):232–9. doi:10.1038/mp.2014.7.
Nagre NN, Subbanna S, Shivakumar M, Psychoyos D, Basavarajappa BS. CB1-receptor knockout neonatal mice are protected against ethanol-induced impairments of DNMT1, DNMT3A, and DNA methylation. J Neurochem. 2015;132(4):429–42. doi:10.1111/jnc.13006.
Otero NK, Thomas JD, Saski CA, Xia X, Kelly SJ. Choline supplementation and DNA methylation in the hippocampus and prefrontal cortex of rats exposed to alcohol during development. Alcohol Clin Exp Res. 2012;36(10):1701–9. doi:10.1111/j.1530-0277.2012.01784.x.
Fowler AK, Hewetson A, Agrawal RG, Dagda M, Dagda R, Moaddel R, et al. Alcohol-induced one-carbon metabolism impairment promotes dysfunction of DNA base excision repair in adult brain. J Biol Chem. 2012;287(52):43533–42. doi:10.1074/jbc.M112.401497.
Tammen SA, Dolnikowski GG, Ausman LM, Liu Z, Sauer J, Friso S, et al. Aging and alcohol interact to alter hepatic DNA hydroxymethylation. Alcohol Clin Exp Res. 2014;38(8):2178–85. doi:10.1111/acer.12477.
Barker JM, Zhang Y, Wang F, Taylor JR, Zhang H. Ethanol-induced Htr3a promoter methylation changes in mouse blood and brain. Alcohol Clin Exp Res. 2013;37 Suppl 1:E101–7. doi:10.1111/j.1530-0277.2012.01906.x.
Srinivasan P, Kapadia R, Biswas A, Said HM. Chronic alcohol exposure inhibits biotin uptake by pancreatic acinar cells: possible involvement of epigenetic mechanisms. Am J Physiol Gastrointest Liver Physiol. 2014;307(9):G941–9. doi:10.1152/ajpgi.00278.2014.
Khachatoorian R, Dawson D, Maloney EM, Wang J, French BA, French SW, et al. SAMe treatment prevents the ethanol-induced epigenetic alterations of genes in the Toll-like receptor pathway. Exp Mol Pathol. 2013;94(1):243–6. doi:10.1016/j.yexmp.2012.09.024.
Qiang M, Li JG, Denny AD, Yao JM, Lieu M, Zhang K et al. Epigenetic mechanisms are involved in the regulation of ethanol consumption in mice. Int J Neuropsychopharmacol. 2014;18(2). doi:10.1093/ijnp/pyu072.
Wani NA, Hamid A, Kaur J. Alcohol-associated folate disturbances result in altered methylation of folate-regulating genes. Mol Cell Biochem. 2012;363(1–2):157–66. doi:10.1007/s11010-011-1168-8.
D’Addario C, Caputi FF, Ekstrom TJ, Di Benedetto M, Maccarrone M, Romualdi P, et al. Ethanol induces epigenetic modulation of prodynorphin and pronociceptin gene expression in the rat amygdala complex. J Mol Neurosci. 2013;49(2):312–9. doi:10.1007/s12031-012-9829-y.
Nielsen DA, Huang W, Hamon SC, Maili L, Witkin BM, Fox RG, et al. Forced abstinence from cocaine self-administration is associated with DNA methylation changes in myelin genes in the corpus callosum: a preliminary study. Front Psychiatry. 2012;3:60. doi:10.3389/fpsyt.2012.00060.
Fragou D, Zanos P, Kouidou S, Njau S, Kitchen I, Bailey A, et al. Effect of chronic heroin and cocaine administration on global DNA methylation in brain and liver. Toxicol Lett. 2013;218(3):260–5. doi:10.1016/j.toxlet.2013.01.022.
Chao MR, Fragou D, Zanos P, Hu CW, Bailey A, Kouidou S, et al. Epigenetically modified nucleotides in chronic heroin and cocaine treated mice. Toxicol Lett. 2014;229(3):451–7. doi:10.1016/j.toxlet.2014.07.023.
Tian W, Zhao M, Li M, Song T, Zhang M, Quan L, et al. Reversal of cocaine-conditioned place preference through methyl supplementation in mice: altering global DNA methylation in the prefrontal cortex. PLoS One. 2012;7(3), e33435. doi:10.1371/journal.pone.0033435.
Rodrigues AJ, Leao P, Pego JM, Cardona D, Carvalho MM, Oliveira M, et al. Mechanisms of initiation and reversal of drug-seeking behavior induced by prenatal exposure to glucocorticoids. Mol Psychiatry. 2012;17(12):1295–305. doi:10.1038/mp.2011.126.
Anier K, Malinovskaja K, Pruus K, Aonurm-Helm A, Zharkovsky A, Kalda A. Maternal separation is associated with DNA methylation and behavioural changes in adult rats. Eur Neuropsychopharmacol J Eur College Neuropsychopharmacol. 2014;24(3):459–68. doi:10.1016/j.euroneuro.2013.07.012.
Pol Bodetto S, Carouge D, Fonteneau M, Dietrich JB, Zwiller J, Anglard P. Cocaine represses protein phosphatase-1Cbeta through DNA methylation and methyl-CpG binding protein-2 recruitment in adult rat brain. Neuropharmacology. 2013;73:31–40. doi:10.1016/j.neuropharm.2013.05.005.
Wilhelm-Benartzi CS, Houseman EA, Maccani MA, Poage GM, Koestler DC, Langevin SM, et al. In utero exposures, infant growth, and DNA methylation of repetitive elements and developmentally related genes in human placenta. Environ Health Perspect. 2012;120(2):296–302. doi:10.1289/ehp.1103927.
Azzi S, Sas TC, Koudou Y, Le Bouc Y, Souberbielle JC, Dargent-Molina P, et al. Degree of methylation of ZAC1 (PLAGL1) is associated with prenatal and post-natal growth in healthy infants of the EDEN mother child cohort. Epigenetics Off J DNA Methylation Soc. 2014;9(3):338–45. doi:10.4161/epi.27387.
van der Knaap LJ, Schaefer JM, Franken IH, Verhulst FC, van Oort FV, Riese H. Catechol-O-methyltransferase gene methylation and substance use in adolescents: the TRAILS study. Genes Brain Behav. 2014;13(7):618–25. doi:10.1111/gbb.12147.
Zhu ZZ, Hou L, Bollati V, Tarantini L, Marinelli B, Cantone L, et al. Predictors of global methylation levels in blood DNA of healthy subjects: a combined analysis. Int J Epidemiol. 2012;41(1):126–39. doi:10.1093/ije/dyq154.
Desplats P, Dumaop W, Cronin P, Gianella S, Woods S, Letendre S, et al. Epigenetic alterations in the brain associated with HIV-1 infection and methamphetamine dependence. PLoS One. 2014;9(7), e102555. doi:10.1371/journal.pone.0102555.
Doehring A, Oertel BG, Sittl R, Lotsch J. Chronic opioid use is associated with increased DNA methylation correlating with increased clinical pain. Pain. 2013;154(1):15–23. doi:10.1016/j.pain.2012.06.011.
Zhang H, Herman AI, Kranzler HR, Anton RF, Zhao H, Zheng W, et al. Array-based profiling of DNA methylation changes associated with alcohol dependence. Alcohol Clin Exp Res. 2013;37 Suppl 1:E108–15. doi:10.1111/j.1530-0277.2012.01928.x.
Zhang H, Herman AI, Kranzler HR, Anton RF, Simen AA, Gelernter J. Hypermethylation of OPRM1 promoter region in European Americans with alcohol dependence. J Hum Genet. 2012;57(10):670–5. doi:10.1038/jhg.2012.98.
Hillemacher T, Weinland C, Lenz B, Kraus T, Heberlein A, Glahn A, et al. DNA methylation of the LEP gene is associated with craving during alcohol withdrawal. Psychoneuroendocrinology. 2015;51:371–7. doi:10.1016/j.psyneuen.2014.10.014.
Beach SR, Brody GH, Lei MK, Gibbons FX, Gerrard M, Simons RL, et al. Impact of child sex abuse on adult psychopathology: a genetically and epigenetically informed investigation. J Fam Psychol JFP J Div Fam Psychol Am Psychol Assoc (Div 43). 2013;27(1):3–11. doi:10.1037/a0031459.
Nieratschker V, Grosshans M, Frank J, Strohmaier J, von der Goltz C, El-Maarri O, et al. Epigenetic alteration of the dopamine transporter gene in alcohol-dependent patients is associated with age. Addict Biol. 2014;19(2):305–11. doi:10.1111/j.1369-1600.2012.00459.x.
Yasar U, Greenblatt DJ, Guillemette C, Court MH. Evidence for regulation of UDP-glucuronosyltransferase (UGT) 1A1 protein expression and activity via DNA methylation in healthy human livers. J Pharm Pharmacol. 2013;65(6):874–83. doi:10.1111/jphp.12053.
Glahn A, Riera Knorrenschild R, Rhein M, Haschemi Nassab M, Groschl M, Heberlein A, et al. Alcohol-induced changes in methylation status of individual CpG sites, and serum levels of vasopressin and atrial natriuretic peptide in alcohol-dependent patients during detoxification treatment. Eur Addict Res. 2014;20(3):143–50. doi:10.1159/000357473.
Heberlein A, Muschler M, Frieling H, Behr M, Eberlein C, Wilhelm J, et al. Epigenetic down regulation of nerve growth factor during alcohol withdrawal. Addict Biol. 2013;18(3):508–10. doi:10.1111/j.1369-1600.2010.00307.x.
Harlaar N, Bryan AD, Thayer RE, Karoly HC, Oien N, Hutchison KE. Methylation of a CpG site near the ALDH1A2 gene is associated with loss of control over drinking and related phenotypes. Alcohol Clin Exp Res. 2014;38(3):713–21. doi:10.1111/acer.12312.
Zhang R, Miao Q, Wang C, Zhao R, Li W, Haile CN, et al. Genome-wide DNA methylation analysis in alcohol dependence. Addict Biol. 2013;18(2):392–403. doi:10.1111/adb.12037.
Philibert RA, Plume JM, Gibbons FX, Brody GH, Beach SR. The impact of recent alcohol use on genome wide DNA methylation signatures. Front Genet. 2012;3:54. doi:10.3389/fgene.2012.00054.
Zhao R, Zhang R, Li W, Liao Y, Tang J, Miao Q, et al. Genome-wide DNA methylation patterns in discordant sib pairs with alcohol dependence. Asia-Pacific Psychiatry Off J Pac Rim Coll Psychiatrists. 2013;5(1):39–50. doi:10.1111/appy.12010.
Philibert RA, Penaluna B, White T, Shires S, Gunter T, Liesveld J, et al. A pilot examination of the genome-wide DNA methylation signatures of subjects entering and exiting short-term alcohol dependence treatment programs. Epigenetics Off J DNA Methylation Soc. 2014;9(9):1212–9. doi:10.4161/epi.32252.
Weng JT, Wu LS, Lee CS, Hsu PW, Cheng AT. Integrative epigenetic profiling analysis identifies DNA methylation changes associated with chronic alcohol consumption. Comput Biol Med. 2014. doi:10.1016/j.compbiomed.2014.12.003.
Ponomarev I, Wang S, Zhang L, Harris RA, Mayfield RD. Gene coexpression networks in human brain identify epigenetic modifications in alcohol dependence. J Neurosci Off J Soc Neurosci. 2012;32(5):1884–97. doi:10.1523/jneurosci.3136-11.2012.
Crews F, He J, Hodge C. Adolescent cortical development: a critical period of vulnerability for addiction. Pharmacol Biochem Behav. 2007;86(2):189–99. doi:10.1016/j.pbb.2006.12.001.
Mill J, Heijmans BT. From promises to practical strategies in epigenetic epidemiology. Nat Rev Genet. 2013;14(8):585–94. doi:10.1038/nrg3405. An overview of current methods used in epigenetic epidemiology, challenges for the field and future directions.
Liang L, Cookson WOC. Grasping nettles: cellular heterogeneity and other confounders in epigenome-wide association studies. Human Mol Genet. 2014;23(R1):R83–8. doi:10.1093/hmg/ddu284. An overview of important confounds that need to be accounted for in epigenetic research.
Bell JT, Pai AA, Pickrell JK, Gaffney DJ, Pique-Regi R, Degner JF, et al. DNA methylation patterns associate with genetic and gene expression variation in HapMap cell lines. Genome Biol. 2011;12(1):R10-R. doi:10.1186/gb-2011-12-1-r10.
Bell JT, Saffery R. The value of twins in epigenetic epidemiology. Int J Epidemiol. 2012. doi:10.1093/ije/dyr179.
Morris TJ, Beck S. Analysis pipelines and packages for Infinium HumanMethylation450 BeadChip (450k) data. Methods (San Diego, Calif). 2015;72:3–8. doi:10.1016/j.ymeth.2014.08.011. An overview of currently available pipelines and packages for the preparation and analysis of DNA methylation data quantified using the Illumina 450k platform.
Tsai P-C, Bell JT. Power and sample size estimation for epigenome-wide association scans to detect differential DNA methylation. Int J Epidemiol. 2015. doi:10.1093/ije/dyv041.
Relton CL, Gaunt T, McArdle W, Ho K, Duggirala A, Shihab H, et al. Data resource profile: Accessible Resource for Integrated Epigenomic Studies (ARIES). Int J Epidemiol. 2015. doi:10.1093/ije/dyv072.
Cecil CA, Lysenko LJ, Jaffee SR, Pingault JB, Smith RG, Relton CL, et al. Environmental risk, oxytocin receptor gene (OXTR) methylation and youth callous-unemotional traits: a 13-year longitudinal study. Mol Psychiatry. 2014;19(10):1071–7. doi:10.1038/mp.2014.95.
Relton CL, Davey SG. Two-step epigenetic Mendelian randomization: a strategy for establishing the causal role of epigenetic processes in pathways to disease. Int J Epidemiol. 2012;41(1):161–76. doi:10.1093/ije/dyr233.
Pingault JB, Cecil CAM, Murray J, Munafò MR, Viding E. Causal inference in psychopathology: a systematic review of Mendelian randomisation studies aiming to identify environmental risk factors for psychopathology. Psychopathol Rev. In Press.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 2008;9:559. doi:10.1186/1471-2105-9-559.
Compliance with Ethics Guidelines
Conflict of Interest
Charlotte A.M. Cecil, Esther Walton, and Essi Viding declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the Topical Collection on Transgenerational Considerations in Addictions
Rights and permissions
About this article
Cite this article
Cecil, C.A.M., Walton, E. & Viding, E. DNA Methylation, Substance Use and Addiction: a Systematic Review of Recent Animal and Human Research from a Developmental Perspective. Curr Addict Rep 2, 331–346 (2015). https://doi.org/10.1007/s40429-015-0072-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40429-015-0072-9