Abstract
Chikungunya is a viral infection transmitted by Aedes mosquitoes in humans. Currently there is no drug available for the treatment of this infection. In absence of a proven therapeutic, vaccines provide an alternative preventive measure for the spread of viral infection in risk prone population. The identification of epitopes is an important step in designing vaccines for viral infections. Traditional methods for vaccine production can be augmented by incorporation of in silico methods at epitopes identification stage. Protein sequences were retrieved for three different viral strains from primary database. Multiple sequence alignment for the retrieved sequences was performed by using MEGA 6.06 to spot the conserved regions. Various immunological databases were utilized to determine the binding affinity with T cells and B cells from structural and non-structural proteins from three different isolates of Chikungunya virus. Five different epitopes were predicted for both structural and non-structural proteins. The epitopes “KKKPGRRERMCMKIE” and “DAEKEAEEEREAELT” and “AEEEREAEL” were predicted as core sequences for class I and class II MHC molecules whereas “SSKYDLECAQ” and “QVLKAKNIGL” were predicted as probable B cell epitopes for structural and non-structural proteins respectively. The identified epitopes can be used for developing a broad spectrum vaccines having effectiveness against different strains of Chikungunya.
Similar content being viewed by others
Introduction
Chikungunya virus belongs to the Togaviridae family transmitted to humans by two mosquitoes Aedes aegypti and Aedes albopictus (Sourisseau et al. 2007). CHIK virus with novel genetic changes was present in the severe Chikungunya outbreaks in 2007 and 2008 in South India. (Sreekumar et al. 2010).
The viral infection was first observed in Tanganyika in the year 1952–1953 (AbuBakar et al. 2007). The symptoms of Chikungunya infection include fever, maculopapular rash and arthralgia (Chhabra et al. 2008). In India, the viral infection was reported during 2006 with more than 1.39 million deaths across 10 states (Pialoux et al. 2007). In 2013, the viral infection was seen in the Caribbean island reporting 355,617 new cases (Susannah 2014). The viral infection had spread across 14 countries and declared as an epidemic infection in few places by the end of April 2014 (WHO 2014).
The Chikungunya viral genome is composed of single stranded positive sense RNA as the genetic material. The genome is 12 kb in size comprising of the 5′-UTR region, coding region of non-structural proteins (nsP1, nsP2, nsP3 and nsP4), coding region for the structural proteins (Capsid, E3, E2, 6 K, E1) and the 3′-poly A tail (Hussain and Chu 2011). The non-structural proteins are responsible for the viral replication. The structural proteins are composed of five subunits translated as a single polyprotein. The capsid protein helps in the viral assembly in host cells and cleaved off to encapsulate the protein. The other proteins E3, E2, 6 K and E1 are translocated to endoplasmic reticulum (ER). The polypeptide is processed further at the N and C terminals resulting in the precursor E2, E3 protein, and 6 K and E1 protein. Heterodimers are formed between E2, E3 and E1 protein at the Golgi complex. The E2, E3 complex undergoes maturation by the action of furin. The heterotrimers comprising envelope proteins E1 and E2, which helps in budding of the virus (Schlesinger and Schlesinger 2001). Both E1 and E2 proteins are responsible for attachment of the virus to the host cell receptor and mediating the immune response in the host cell. E2 gene of CHIKV was composed of 3 domains and the major adaptive mutations were detected in domain B revealed by Structural modelling, which can modulate binding of CHIKV to host cells, while the transmembrane domain in domain C and the epitopes were located in domain A, which was found to be most conserved. (Sahu et al. 2013).
Vaccines are used to provide protection against any infection by stimulating the immune response (Grammatikos et al. 2009). In silico approaches seem to play an evident role in vaccine development by reducing the cost and time in screening protocols (Groot et al. 2001).
The study done by Pallavi and Seth 2009 was identified T-cell epitopes for structural proteins of dengue virus using in silico methods. The epitopes were screened against MHC class I and class II peptides (Pallavi and Seth 2009). The neutralizing epitope of H5N1 virus was predicted in other study using molecular dynamics simulation to identify the solvent accessible regions of the antigen (Mulyanto and Saleh 2011). In another study by Xing 2013 where the phylogenetic analysis was performed and B-cell epitopes were predicted for VP1 gene of Duck Hepatitis A virus 1 (Xing 2013). In a study by Subramanian and Chinnappan 2013 the epitopes were predicted for e6 protein of human papilloma virus using in silico approach. The B-cell epitopes were identified from the glycoprotein and envelope protein of Nipah virus using the computational immunology protocols (Sadnam et al. 2014).
The development of new vaccines is affected by several barriers. The antigens having immunological response in animals may not have the same response as in humans (Dannenberg 2010).
Coming to the anatomy of Chikungunya, the viral surface proteins are antigenic in nature responsible for the pathogenicity whereas the envelope proteins help in viral entry through the cell surface receptors in the host. The envelope proteins are considered to be potential aspirants for vaccine design which acts as a target for the B-cells (Cerdeño-Tárraga et al. 2003). The B-cell and T-cell epitopes must be identified for vaccine design. The binding of antigen to human leukocyte antigen class I (CD8 + T-cells) and class II (CD4 + T-cells) alleles is a measure of the diversity and specificity of immune response (Watts 1997, Germain 1994). Great public health concern is raised by the high morbidity and socio-economic loss associated with the recent CHIKV epidemics worldwide and emphasizes the need to study the immunological basis of CHIKV infection to control the disease. MHC-I restricted epitopes that may help in the advancement of MHC-I restricted epitope based anti-CHIKV immune responses against this infection and this will be useful towards the development of epitope based anti-CHIKV immunotherapy in the future. (Pratheek et al. 2015). These tools offer high specificity and sensitivity and sheds light on the features of immunotherapy against various diseases by designing vaccines based on epitope peptides. This study is an attempt to predict the epitope based peptides against the dangerous Chikungunya virus using immuno-informatics approaches.
Materials and methods
Chikungunya sequence retrieval
The sequences of structural and non-structural proteins from different isolates of Chikungunya virus (CHIKV) were retrieved from UniProt KB (http://www.uniprot.org) and NCBI Protein database (http://www.ncbi.nlm.nih.gov/protein) in FASTA format. The structural protein sequences of different Chikungunya viral isolates namely, S27-African prototype [Q8JUX5], structural polyprotein strain 37997 [Q5XXP3] and structural polyprotein viral strain Nagpur [Q5WQY5] were retrieved from UniProt along with non-structural protein sequences namely, S27-African prototype [Q8JUX6], Non-structural polyprotein strain 37997 [Q5XXP4].
Multiple sequence alignment
The retrieved sequences were subjected to multiple sequence alignment in MEGA 6.06 software. The sequences were aligned to identify the evolutionary origin and the regions of conservation among sequences isolated from geographically diverse locations. This step is necessary so that epitopes can be designed for conserved proteins in virus which will help in making the vaccines of a broad spectrum for their action and specificity. MEGA (Molecular Evolutionary Genetics Analysis) is a web-tool used to perform sequence alignment, understanding phylogenetic trees, and to infer the ancestral relationship among different sequences (http://www.megasoftware.net). ClustalW algorithm was used with default parameters including the length of input sequence, along with maximum number of input sequences to perform the sequence alignment. The sequences were analysed to identify regions which are immunologically competent for prediction of epitope peptides. The peptide must comprise of minimal number of amino acids for behaving as an epitope. Hence the sequences with minimal length having an average length of 15 amino acids (Kringelum et al. 2013) were considered for prediction of epitopes in this study.
Antigenicity prediction and recognition
Antigenicity is defined as the process of recognition of the foreign molecules by the antibodies and the cells of immune system (Cruse and Lewis 1998). The amino acid sequence that was conserved from non-structural and structural proteins was screened for antigenicity prediction. The antigenicity was predicted using VaxiJen v2.0, an open source web server for antigen prediction (Doytchinova and Flower 2007). The T-cell epitopes were predicted by providing the protein sequences as an input. The membrane and soluble regions from the protein sequences were differentiated based on the transmembrane properties. To perform the prediction of transmembrane properties, the identified amino acid sequence was subjected to transmembrane topology estimation in TMHMM v2.0 server (Krogh et al. 2001). The TMHMM v2.0 web server is used to distinguish the surface and intracellular proteins with increased degree of accuracy compared to other available tools.
Identification of T-cell epitopes from conserved regions
The epitopes were identified from cytotoxic T lymphocytes (CTL) based on their conserved regions using the web server, NetCTL v1.2 (Larsen et al. 2005). The binding of antigenic peptides to the MHC class I molecules initiates an immune response and plays an important role in identifying potential molecules for vaccine design (Menaka et al. 2007). This web-server is used for prediction of MHC class I peptides bound to 12 different HLA super types using neural network design. The sensitivity and specificity were set at 0.89 and 0.94 and the threshold score is fixed at 0.5. The total score of antigenicity was computed on the basis of binding of the given peptide to MHC class I alleles, terminal cleavage by proteosomal C and efficiency of transporting of antigenic peptides (TAP) respectively (Larsen et al. 2007). The appropriate epitopes were identified from the scores obtained from the prediction method, which is a function of weighted sum with a relative weight on peptide to the binding of MHC class I molecules. The binding of the peptides to explicit MHC molecules alleles was facilitated to calculate the IC50 values. The Stabilized Matrix Method (SMM) was used for prediction from the Immune Epitope Database (IEDB) (Peter and Sette 2005). The MHC alleles were designated and the length of the peptide sequences was set to default value 9.0 before the prediction. The input parameters used for recognition of immunogenicity were normalized having the range 0–1 for obtaining a weighted score. The conserved peptides were also selected for prediction of epitope interaction with the MHC class II alleles. The prediction was performed using IEDB prediction tool for MHC class II alleles (http://tools.immuneepitope.org/mchii). The SMM prediction method was used to identify the efficient binders for MHC class II alleles by calculating the IC50 values based on the threshold score of 300 nM.
Prediction of B-cell epitopes
The prediction of this B-cell response is crucial in designing a peptide based antigen (Tomar and De 2010). The linear B-cell epitopes were predicted using ABCpred server (Kaur et al. 2007). ABCpred uses a recurrent artificial neural network for prediction of linear B-cell epitopes. The prediction model used by ABCpred has been evaluated on a dataset of 700 epitopes and 700 non-epitope peptides and a five fold cross validation has been performed during model construction (Yasser and Vasant 2010). Before prediction, the conserved sequences were identified from VaxiJen v2.0 with a threshold score of 0.4. The length of the epitope was fixed to nanomer, consensus sequence of nine deoxyribonucleotides. The repeated sequences were filtered out. The variable length peptides from the B-cell epitopes were compared to that of the T-cell epitopes obtained from previous steps. The nanomers having substantial superimposition (≥6 amino acid repeats) on the predicted B-cell epitopes were selected for further interpretation.
EpiTOP 1.0Footnote 1 server predicts log1/IC50 values of MHC class II alleles. It is a QSAR approach based on proteochemometrics. The Peptides of MHC class II alleles are used as an input.
SVMTrip toolFootnote 2 which provide the recommended epitopes. These epitopes are recognised by the sign of a flag. The amino acid sequence of a protein is taken as the input. The web server is constructed for the public use. SVMTrip supports lengthy sequences which may be consist of more tri-peptide detectable frequency tendency. SVMTrip are applied to virus and human peptides, and top ranked peptides are returned (Table 1).
EPMLRFootnote 3 web-server which helps users to predict Linear B-cell epitope with their interested protein or peptide. Users need to paste input sequence and the predicted results will be ranked in decreasing order of predicted score (Supplementary Table). Higher the score of a peptide, higher will be its probability of the epitope (Yao et al. 2014).
3D structure prediction for the predicted epitopes
The structure of the predicted epitopes was determined using Pepstr server. The 3D structures of peptides were predicted to perform future molecular docking studies with 3D structure of HLA molecule. The peptide sequences predicted from B-cell and T-cell was given as input to the Pepstr web server. This tool predicts the protein structures on the basis of secondary structure information derived from PSIPRED and the β turns information is predicted by Beta Turns tool (Saha and Raghava 2006). The peptides having higher probability to act as an epitope were filtered out based on the scores obtained from VaxiJen, ABCpred and MHC class I, class II web servers. The peptide structure obtained from Pepstr server was further visualized using PyMOL 1.7.2 graphics. The flowchart depicting the methodology for epitope prediction is shown in Fig. 1.
Results and discussion
Retrieval of protein sequences and identification of conserved regions
A total of 5 sequences of non-structural and structural proteins were retrieved from different isolates of CHIKV. MEGA software was used for performing multiple sequence alignment. The CLUSTALW software was used to generate conserved sequences from non-structural and structural protein sequences. The conserved sequences generated from the alignment are represented in Table 2.
Prediction of antigenicity and transmembrane characteristics
The conserved regions were identified from both structural and non-structural proteins based on the threshold score (≥0.4) as seen in VaxiJen tool. Furthermore, the transmembrane characteristics were studied using the TMHMM tool. All peptides identified from VaxiJen tool for structural proteins (18 proteins) and non-structural proteins (19 proteins) were identified as either intracellular or surface proteins.
Determination of T cell epitopes
The T cell receptors are represented on the antigen presenting cells (APC) facilitated by the class I and class II molecules. These T cell receptors are used for recognition of epitopes expressed on the T cells. The T cell epitopes were identified and further evaluated based on the results from VaxiJen. The results were iterated twice due to the loss of antigenicity observed in some T cell epitopes. The iteration enabled to achieve a good rate of T cell epitope prediction. The T cell epitopes were predicted as shown in Tables 2 and 3.
Prediction of MHC class I epitopes and identification
The NetCTL prediction tool was used to identify the conserved regions from structural and non-structural proteins. 307 and 316 nonamers were identified from the conserved regions of structural and non-structural proteins. The MHC class I alleles were predicted using IEDB database using SMM align method. A total of 54 T cell epitopes from the structural proteins interacted with 26 predicted MHC class I alleles having the IC50 value <300 nM. Using the same limit for IC50 value the conserved regions of non-structural proteins revealed a binding interaction with 46 possible MHC class I alleles. The alleles were tested for their antigenicity using VaxiJen. The next step was to identify the peptides having interaction with more than four MHC class I alleles having the VaxiJen score greater than 0.5. Based on the criteria 22 nonamers from structural proteins and 18 nonamers from non-structural proteins were obtained as shown in Table 3. Further analysis revealed that the predicted epitope sequence “KYDLECAQI” derived from structural proteins had the highest antigenicity score of 2.1584 from VaxiJen. The epitopes “RAVPQQKPR”, “ERMCMKIEN” and “AEEEREAEL” having scores of 1.1329, 1.5827 and 1.0952 respectively from structural proteins and non-structural protein interacted with 6 different MHC class I alleles.
Prediction of MHC class II alleles from the conserved regions
There is a direct relationship between SMM-align prediction scores and log-transformed IC50 binding affinity. The IC50 value of 300 nM was used as criteria to increase the confidence level of the prediction of MHC class II alleles. The MHC class II alleles were predicted from structural and non-structural proteins. A total of 68 and 21 peptides were predicted from structural and non-structural proteins having IC50 ranging from 23 to 300 nM. A good epitope must interact with many MHC alleles. Among the total peptides predicted, only 11 peptides from 68 peptides structural proteins and 7 peptides from 21 non-structural peptides had interactions with more than three MHC alleles. The peptides were further evaluated using the antigenicity scores obtained from VaxiJen. The peptides having score ≥0.5 were selected. The epitopes “KKKPGRRERMCMKIE” and “DAEKEAEEEREAELT” were having highest VaxiJen scores of 1.2794 and 0.8668 respectively. The predicted MHC class II alleles for structural and non-structural proteins were shown in Table 4.
Prediction and selection of B-cell epitopes
The B-cell epitopes were identified from ABCpred server for structural and non-structural proteins on the basis of VaxiJen scores. The prediction resulted in 16 epitopes for structural proteins having scores in the range 0.50–2.14 along with 20 epitopes from non-structural proteins having the scores in the range 0.50–2.63 (Table 5). The peptide sequence “SSKYDLECAQ” and “QVLKAKNIGL” from structural and non-structural protein had highest scores from VaxiJen (2.0390 and 2.3633 respectively). The decamer sequences from structural and non-structural proteins had sequence similarity with the T cell nonameric predicted epitopes. EpiTOP identifies 89 % of known epitopes within the top 20 % of predicted binders, reducing laboratory labour, materials and time by 80 %. EpiTop gives comprehensive quantitative predictions and will be expanded and updated with new quantitative matrices over time (Dimitrov et al. 2010).
SVMTrip achieves a sensitivity of 80.1 % and a precision of 55.2 % for sequences with 20 amino acids which are higher than those of AAP and BCPred (Yao et al. 2012).
EPMLR produced better performance in comparison with ABCpred, AAP and BCPred. EPMLR offers new insights into the Linear B cell epitopes prediction and a new option for scientists to do their prediction (Yao et al. 2014).
Prediction of structure for the identified epitopes
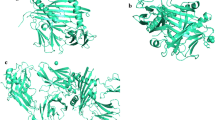
The epitopes predicted for B-cell and T-cell were utilized for prediction of structure using Pepstr web server. Five potential epitopes were identified and their structures were generated using Pepstr. The structures of peptides were further visualized in PyMOL graphics (Fig. 2).
Discussion
New infections are emerging at a faster pace in the current scenario. In particular, the zoonotic diseases are spreading among humans over the recent years. Medical science has tried to improve the current suppositories for enhanced treatment. Chikungunya viral infection is one of the emerging disease transmitted to humans by the Aedes mosquito. The infection has been spreading to various regions across the world at a faster pace. Although, research is been conducted to develop vaccines, currently there is no vaccine developed for utility in humans. An attempt is made in this study to predict and design the epitopes which can be tested to generate an immune response. The structural and non-structural proteins were targeted to design the epitopes using the in silico methods. The conserved regions were identified from these sequences as shown in Table 2. Peptide antigens which are generally 8–10 amino acids in length have potency for binding to both MHC and T-cell receptor (TCR). The peptides were predicted using IEDB database followed by having a sequence match of 100 %. The VaxiJen scores were predicted for each peptide to detect the antigenicity. The peptides having the scores greater than the threshold value are selected for T-cell and B-cell epitope prediction. The MHC class I and class II alleles are predicted for each of the peptides (Table 3). The B cell epitopes were predicted using ABCpred and the scores were predicted using VaxiJen score. The epitopes having high scores of antigenicity from both the tools were further used for prediction of the structure. This approach of computational prediction of epitopes can be utilized as a starting point to develop vaccines against Chikungunya virus. The predicted epitopes can be tested on animal models for the development of vaccine. The method reduces time and cost involved in screening of T-cell and B-cell epitopes implemented using experimental approaches. Aptness of the predicted peptides as a probable vaccine can be investigated based on the experimental approaches.
Notes
EpiTOP 1.0 is available at http://www.pharmfac.net/EpiTOP/.
SVMTrip tool is available at http://sysbio.unl.edu/SVMTrip.
EPMLR is available at http://www.bioinfo.tsinghua.edu.cn/epitope/EPMLR/.
References
AbuBakar S, Sam IC, Wong PF, MatRahim N, Hooi PS, Roslan N (2007) Reemergence of endemic chikungunya, Malaysia. Emerg Infect Dis 13:147–149
Cerdeño-Tárraga AM, Efstratiou A, Dover LG et al (2003) The complete genome sequence and analysis of Corynebacterium diphtheriae NCTC13129. Nucleic Acids Res 31(22):6516–6523
Chhabra M, Mittal V, Bhattacharya D, Rana U, Lal S (2008) Chikungunya fever: a re-emerging viral infection. Indian J Med Microbiol 26(1):5–12
Cruse Julius M, Lewis Robert E (1998) Atlas of immunology. CRC Press, Boca Raton
Dannenberg AM (2010) Perspectives on clinical and preclinical testing of new tuberculosis vaccines. Clin Microbiol 23:781–794
Dimitrov I, Panayot G, Darren RF, Irini D (2010) EpiTOP—a proteochemometric tool for MHC class II binding prediction. Bioinformatics 26(16):2066–2068
Doytchinova IA, Flower DR (2007) VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform 8:4
Germain RN (1994) MHC-dependent antigen processing and peptide presentation: providing ligands for T lymphocyte activation. Cell 76(2):287–299
Grammatikos Alexandros P, Mantadakis Elpis, Falagas Matthew E (2009) Meta-analyses on pediatric infections and vaccines. Infect Dis Clin North Am 23(2):431–457
Groot De, Bosma AS, Chinai N (2001) From genome to vaccine: In silico predictions, ex vivo verification. Vaccine 19:4385–4395
Hussain KM, Chu JJH (2011) Insights into the interplay between chikungunya virus and its human host. Future Virol 6(10):1211–1223
Kaur H, Garg A, Raghava GPS (2007) PEPstr: a de novo method for tertiary structure prediction of small bioactive peptides. Protein Pept Lett 14:626–630
Kringelum JV, Nielsen M, Padkjaer SB, Lund O (2013) Structural analysis of B-cell epitopes in antibody: protein complexes. Mol Immunol 53:24–34
Krogh A, Larsson B, Von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305(3):567–580
Larsen MV, Lundegaard C, Lamberth K et al (2005) An integrative approach to CTL epitope prediction: a combined algorithm integrating MHC class I binding, TAP transport efficiency, and proteasomal cleavage predictions. Eur J Immunol 35(8):2295–2303
Larsen MV, Lundegaard C, Lamberth K, Buus S, Lund O, Nielsen M (2007) NetCTL-1.2: large-scale validation of methods for cytotoxic T-lymphocyte epitope prediction. BMC Bioinform 8:424
Menaka R, Bertil S, Lin F, Vladimir B (2007) Predicting peptides binding to MHC class II molecules using multi-objective evolutionary algorithms. BMC Bioinform 8:459
Mulyanto C, Saleh R (2011) Prediction of a neutralizing epitope of a H5N1 virus hemagglutinin complexed with an antibody variable fragment using molecular dynamics simulation. J Biophys Chem 2:258–267
Pallavi S, Seth PK (2009) Prediction of T cell epitopes for the utility of vaccine development from structural proteins of dengue virus variants using in silico methods. Indian J Biotechnol 8:193–198
Peter B, Sette A (2005) Generating quantitative models describing the sequence specificity of biological processes with the stabilized matrix method. BMC Bioinform 31(6):132
Pialoux G, Gaüzère B, Jauréguiberry S, Strobel M (2007) Chikungunya, an epidemic arbovirus. Lancet Infect Dis 7:319–327
Pratheek BM, Suryawanshi AR, Chattopadhyay S, Chattopadhyay S (2015) In silico analysis of MHC-I restricted epitopes of Chikungunya virus proteins: Implication in understanding anti-CHIKV CD8+ T cell response and advancement of epitope based immunotherapy for CHIKV infection. Infect Genet Evol 31:118–126
Sadnam S, Rehman I, Mahbud HAKM, Nurun NAHM (2014) Prediction of Epitope-based peptides for the utility of vaccine development from fusion and glycoprotein of Nipah virus using in silico approach. Adv Bioinform 2014:1–17
Saha S, Raghava GPS (2006) Prediction of continuous B‐cell epitopes in an antigen using recurrent neural network. Proteins 65(1):40–48
Sahu A, Das B, Das M, Patra A, Biswal S, Kar SK, Hazra RK (2013) Genetic characterization of E2 region of Chikungunya virus circulating in Odisha, Eastern India from 2010 to 2011. Infect Genet Evol 18:113–124
Schlesinger S, Schlesinger MJ (2001) Togaviridae: the viruses and their replication. Fields Virol 1:895–916
Sourisseau M, Schilte C, Casartelli N, Trouillet C, Guivel-Benhassine F, Rudnicka D (2007) Characterization of re-emerging chikungunya virus. PLoS Pathog 3(6):e89
Sreekumar E, Issac A, Nair S, Hariharan R, Janki MB, Arathy DS, Regu R, Mathew T, Anoop M, Niyas KP, Pillai MR (2010) Genetic characterization of 2006-2008 isolates of Chikungunya virus from Kerala, South India, by whole genome sequence analysis. Virus Genes 40:14–27
Subramanian N, Chinnappan S (2013) Prediction of promiscuous epitopes in the e6 protein of three risk human papilloma viruses: a computational approach. Asian Pac J Cancer Prev 14(7):4167–4175
Susannah L (2014) The painful, mosquito borne chikungunya virus has reached the US. http://www.vox.com/2014/7/19/5916249/chikungunya-fever-virus-caribbean-florida-disease-symptoms-explained
Tomar N, De RK (2010) Immunoinformatics: an integrated scenario. Immunology 131:153–168
Watts C (1997) Capture and processing of exogenous antigens for presentation on MHC molecules. Annu Rev Immunol 15:821–850
WHO (2014) Chikungunya—Fact sheet. European Centre for Disease Prevention and Control (ECDC)
Xing JW (2013) sequence analysis and b cell epitope prediction of duck hepatitis a virus 1 vp1 gene. Adv Mater Res 647:217
Yao B, Zhang L, Liang S, Zhang C (2012) SVMTriP: a method to predict antigenic epitopes using support vector machine to integrate tri-peptide similarity and propensity. PLoS One 7(9):e45152
Yao L, Meng G, Xian-Ming P (2014) EPMLR: sequence-based linear B-cell epitope prediction method using multiple linear regression. BMC Bioinform 15:414
Yasser EM, Vasant H (2010) Recent advances in B-cell epitope prediction methods. Immunome Res 6(Suppl2):S2
Acknowledgments
Authors wish to thank the Department of Biotechnology for DBT BIF centre and Department of Science and Technology for DST CURIE project for Women’s University, Ministry of Science and Technology, New Delhi, India, for financial assistance. These authors declare that they have no conflict of interest and this article does not contain any studies with human and animal subjects performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kori, P., Sajjan, S.S. & Madagi, S.B. In silico prediction of epitopes for Chikungunya viral strains. Journal of Pharmaceutical Investigation 45, 579–591 (2015). https://doi.org/10.1007/s40005-015-0205-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40005-015-0205-0