Abstract
Purpose
Ca2+ homeostasis plays a pivotal role in regulating proliferation and apoptosis during cancer development. This study intended to examine the potential tumor-suppressing role of ZNF503 antisense RNA 1 (ZNF503-AS1) in bladder cancer, which may be implicated in the regulation of Ca2+ homeostasis.
Methods
Differentially expressed long non-coding RNAs (lncRNAs) related to bladder cancer were identified using microarray analysis, followed by the verification of transcription factors to which they bind. The relationship between ZNF503-AS1, GATA6 and SLC8A1 was assessed using dual luciferase reporter, RIP and ChIP assays. The expression levels of ZNF503-AS1, GATA6 and SLC8A1 were modulated to examine their effects on the tumorigenic potential, intracellular Ca2+ concentration and Ca2+-ATPase activity in bladder cancer cells. The in vivo tumorigenic ability was validated in nude mice.
Results
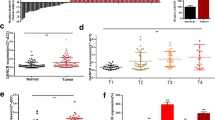
Microarray-based expression profile analysis of the GEO GSE61615 dataset revealed that the expression of ZNF503-AS1 was decreased in bladder cancer. Subsequently, we found that ZNF503-AS1 can bind to the transcription factor GATA6 to up-regulate the expression of SLC8A1. ZNF503-AS1 and SLC8A1 were found to be down-regulated in both primary bladder cancer tissues and cells. Exogenous overexpression of ZNF503-AS1 or SLC8A1 attenuated bladder cancer cell proliferation, invasion and migration, but promoted their apoptosis, accompanied by decreased Ca2+-ATPase activities and increased intracellular Ca2+ concentrations. Additional in vivo experiments validated the inhibitory effect of ZNF503-AS1 overexpression on the tumorigenic capacity of bladder cancer cells in nude mice.
Conclusion
ZNF503-AS1 can recruit transcription factor GATA6 to up-regulate SLC8A1 expression, thereby increasing the intracellular Ca2+ concentration and repressing the proliferation, invasion and migration, and enhancing the apoptosis of bladder cancer cells.







Similar content being viewed by others
References
D.S. Kaufman, W.U. Shipley, A.S. Feldman, Bladder cancer. Lancet 374, 239–249 (2009). https://doi.org/10.1016/S0140-6736(09)60491-8
S. Antoni, J. Ferlay, I. Soerjomataram, A. Znaor, A. Jemal, F. Bray, Bladder cancer incidence and mortality: A global overview and recent trends. Eur Urol 71, 96–108 (2017). https://doi.org/10.1016/j.eururo.2016.06.010
H.L. Roderick, S.J. Cook, Ca2+ signalling checkpoints in cancer: Remodelling Ca2+ for cancer cell proliferation and survival. Nat Rev Cancer 8, 361–375 (2008). https://doi.org/10.1038/nrc2374
S.W. Ip, Y.L. Chu, C.S. Yu, P.Y. Chen, H.C. Ho, J.S. Yang, H.Y. Huang, F.S. Chueh, T.Y. Lai, J.G. Chung, Bee venom induces apoptosis through intracellular Ca2+ −modulated intrinsic death pathway in human bladder cancer cells. Int J Urol 19, 61–70 (2012). https://doi.org/10.1111/j.1442-2042.2011.02876.x
M.G.K. Cumberbatch, A.P. Noon, Epidemiology, aetiology and screening of bladder cancer. Transl Androl Urol 8, 5–11 (2019). https://doi.org/10.21037/tau.2018.09.11
K.A. Mitchell, H. Williams, Emerging genomic biomarkers for improving kidney, prostate, and bladder cancer health disparities outcomes. Urol Oncol 22, S1078-1439 (2019). https://doi.org/10.1016/j.urolonc.2019.04.024
M. Grayson, Bladder cancer. Nature 551, S33 (2017). https://doi.org/10.1038/551S33a
D. Liu, Y. Li, G. Luo, X. Xiao, D. Tao, X. Wu, M. Wang, C. Huang, L. Wang, F. Zeng, G. Jiang, LncRNA SPRY4-IT1 sponges miR-101-3p to promote proliferation and metastasis of bladder cancer cells through up-regulating EZH2. Cancer Lett 388, 281–291 (2017). https://doi.org/10.1016/j.canlet.2016.12.005
Q. Hua, X. Lv, X. Gu, Y. Chen, H. Chu, M. Du, W. Gong, M. Wang, Z. Zhang, Genetic variants in lncRNA H19 are associated with the risk of bladder cancer in a Chinese population. Mutagenesis 31, 531–538 (2016). https://doi.org/10.1093/mutage/gew018
J. Zhuang, L. Shen, L. Yang, X. Huang, Q. Lu, Y. Cui, X. Zheng, X. Zhao, D. Zhang, R. Huang, H. Guo, J. Yan, TGFbeta1 promotes gemcitabine resistance through regulating the LncRNA-LET/NF90/miR-145 signaling axis in bladder cancer. Theranostics 7, 3053–3067 (2017). https://doi.org/10.7150/thno.19542
X. Agirre, C. Meydan, Y. Jiang, L. Garate, A.S. Doane, Z. Li, A. Verma, B. Paiva, J.I. Martin-Subero, O. Elemento, C.E. Mason, F. Prosper, A. Melnick, Long non-coding RNAs discriminate the stages and gene regulatory states of human humoral immune response. Nat Commun 10, 821 (2019). https://doi.org/10.1038/s41467-019-08679-z
M. Huarte, The emerging role of lncRNAs in cancer. Nat Med 21, 1253–1261 (2015). https://doi.org/10.1038/nm.3981
K. Inamura, Major tumor suppressor and oncogenic non-coding RNAs: Clinical relevance in lung cancer. Cells 6,12 (2017). https://doi.org/10.3390/cells6020012
X. Chen, C. Jiang, B. Qin, G. Liu, J. Ji, X. Sun, M. Xu, S. Ding, M. Zhu, G. Huang, B. Yan, C. Zhao, LncRNA ZNF503-AS1 promotes RPE differentiation by downregulating ZNF503 expression. Cell Death Dis 8, e3046 (2017). https://doi.org/10.1038/cddis.2017.382
P. Shahi, C.Y. Wang, D.A. Lawson, E.M. Slorach, A. Lu, Y. Yu, M.D. Lai, H. Gonzalez Velozo, Z. Werb, ZNF503/Zpo2 drives aggressive breast cancer progression by down-regulation of GATA3 expression. Proc Natl Acad Sci U S A 114, 3169–3174 (2017). https://doi.org/10.1073/pnas.1701690114
K.A. Kwei, M.D. Bashyam, J. Kao, R. Ratheesh, E.C. Reddy, Y.H. Kim, K. Montgomery, C.P. Giacomini, Y.L. Choi, S. Chatterjee, C.A. Karikari, K. Salari, P. Wang, T. Hernandez-Boussard, G. Swarnalata, M. van de Rijn, A. Maitra, J.R. Pollack, Genomic profiling identifies GATA6 as a candidate oncogene amplified in pancreatobiliary cancer. PLoS Genet 4, e1000081 (2008). https://doi.org/10.1371/journal.pgen.1000081
J.J. Munoz, S.A. Drigo, M.C. Barros-Filho, F.A. Marchi, C. Scapulatempo-Neto, G.S. Pessoa, G.C. Guimaraes, J.C. Trindade Filho, A. Lopes, M.A. Arruda, S.R. Rogatto, Down-regulation of SLC8A1 as a putative apoptosis evasion mechanism by modulation of calcium levels in penile carcinoma. J Urol 194, 245–251 (2015). https://doi.org/10.1016/j.juro.2014.11.097
A. Fujita, J.R. Sato, O. Rodrigues Lde, C.E. Ferreira, M.C. Sogayar, Evaluating different methods of microarray data normalization. BMC Bioinformatics 7, 469 (2006). https://doi.org/10.1186/1471-2105-7-469
G.K. Smyth, Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3, Article3 (2004). https://doi.org/10.2202/1544-6115.1027
Y.L. Tuo, X.M. Li, J. Luo, Long noncoding RNA UCA1 modulates breast cancer cell growth and apoptosis through decreasing tumor suppressive miR-143. Eur Rev Med Pharmacol Sci 19, 3403–3411 (2015)
G. Arun, S. Diermeier, M. Akerman, K.C. Chang, J.E. Wilkinson, S. Hearn, Y. Kim, A.R. MacLeod, A.R. Krainer, L. Norton, E. Brogi, M. Egeblad, D.L. Spector, Differentiation of mammary tumors and reduction in metastasis upon Malat1 lncRNA loss. Genes Dev 30, 34–51 (2016). https://doi.org/10.1101/gad.270959.115
S.Y. Kim, D. Yang, J. Myeong, K. Ha, S.H. Kim, E.J. Park, I.G. Kim, N.H. Cho, K.P. Lee, J.H. Jeon, I. So, Regulation of calcium influx and signaling pathway in cancer cells via TRPV6-Numb1 interaction. Cell Calcium 53, 102–111 (2013). https://doi.org/10.1016/j.ceca.2012.10.005
D.T. Miyamoto, K.W. Mouw, F.Y. Feng, W.U. Shipley, J.A. Efstathiou, Molecular biomarkers in bladder preservation therapy for muscle-invasive bladder cancer. Lancet Oncol 19, e683–e695 (2018). https://doi.org/10.1016/S1470-2045(18)30693-4
J.R. Prensner, A.M. Chinnaiyan, The emergence of lncRNAs in cancer biology. Cancer Discov 1, 391–407 (2011). https://doi.org/10.1158/2159-8290.CD-11-0209
Y.P. Zhu, X.J. Bian, D.W. Ye, X.D. Yao, S.L. Zhang, B. Dai, H.L. Zhang, Y.J. Shen, Long noncoding RNA expression signatures of bladder cancer revealed by microarray. Oncol Lett 7, 1197–1202 (2014). https://doi.org/10.3892/ol.2014.1843
R. Flippot, G. Beinse, A. Boileve, J. Vibert, G.G. Malouf, Long non-coding RNAs in genitourinary malignancies: A whole new world. Nat Rev Urol 16, 484–504 (2019). https://doi.org/10.1038/s41585-019-0195-1
W. Zhu, H. Liu, X. Wang, J. Lu, W. Yang, Long noncoding RNAs in bladder cancer prognosis: A meta-analysis. Pathol Res Pract 215, 152429 (2019). https://doi.org/10.1016/j.prp.2019.04.021
G. Hu, Q. Tang, S. Sharma, F. Yu, T.M. Escobar, S.A. Muljo, J. Zhu, K. Zhao, Expression and regulation of intergenic long noncoding RNAs during T cell development and differentiation. Nat Immunol 14, 1190–1198 (2013). https://doi.org/10.1038/ni.2712
A. Fatica, I. Bozzoni, Long non-coding RNAs: New players in cell differentiation and development. Nat Rev Genet 15, 7–21 (2014). https://doi.org/10.1038/nrg3606
P. Shahi, E.M. Slorach, C.Y. Wang, J. Chou, A. Lu, A. Ruderisch, Z. Werb, The transcriptional repressor ZNF503/Zeppo2 promotes mammary epithelial cell proliferation and enhances cell invasion. J Biol Chem 290, 3803–3813 (2015). https://doi.org/10.1074/jbc.M114.611202
G.R. Monteith, F.M. Davis, S.J. Roberts-Thomson, Calcium channels and pumps in cancer: Changes and consequences. J Biol Chem 287, 31666–31673 (2012). https://doi.org/10.1074/jbc.R112.343061
C. Ibarra, M. Karlsson, S. Codeluppi, M. Varas-Godoy, S. Zhang, L. Louhivuori, S. Mangsbo, A. Hosseini, N. Soltani, R. Kaba, T. Kalle Lundgren, A. Hosseini, N. Tanaka, M. Oya, P. Wiklund, A. Miyakawa, P. Uhlen, BCG-induced cytokine release in bladder cancer cells is regulated by Ca(2+) signaling. Mol Oncol 13, 202–211 (2019). https://doi.org/10.1002/1878-0261.12397
Q. Lu, T. Liu, H. Feng, R. Yang, X. Zhao, W. Chen, B. Jiang, H. Qin, X. Guo, M. Liu, L. Li, H. Guo, Circular RNA circSLC8A1 acts as a sponge of miR-130b/miR-494 in suppressing bladder cancer progression via regulating PTEN. Mol Cancer 18, 111 (2019). https://doi.org/10.1186/s12943-019-1040-0
Y. Zhong, Z. Wang, B. Fu, F. Pan, S. Yachida, M. Dhara, E. Albesiano, L. Li, Y. Naito, F. Vilardell, C. Cummings, P. Martinelli, A. Li, R. Yonescu, Q. Ma, C.A. Griffin, F.X. Real, C.A. Iacobuzio-Donahue, GATA6 activates Wnt signaling in pancreatic cancer by negatively regulating the Wnt antagonist Dickkopf-1. PLoS One 6, e22129 (2011). https://doi.org/10.1371/journal.pone.0022129
W. Shen, N. Niu, B. Lawson, L. Qi, J. Zhang, T. Li, H. Zhang, J. Liu, GATA6: A new predictor for prognosis in ovarian cancer. Hum Pathol 86, 163–169 (2019). https://doi.org/10.1016/j.humpath.2019.01.001
Acknowledgments
We would like show sincere appreciation to the reviewers for critical comments on this article.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
None declared.
Ethical statement
The study was conducted under approval of the Ethics Committee of The Second Xiangya Hospital, Central South University. All participating patients signed informed consent documentation. Nude mice were used for in vivo studies and were cared for in accordance with the principles of the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health with efforts made to ensure minimal suffering of the animals used in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Fig. S1
Overexpressed ZNF503-AS1 impedes proliferation, migration, invasion and resistance to apoptosis of bladder cancer J82 cells. A: qRT-PCR and Western blot analysis for the expression of ZNF503-AS1 in J82 cells after sh-ZNF503-AS1 or oe-ZNF503-AS1 transfection. B: EdU assay for the effect of ZNF503-AS1 overexpression on the proliferation of J82 cells (× 200, scale bar = 50 μm), C: quantitative flow cytometric detection for the effect of ZNF503-AS1 overexpression on the apoptosis of J82 cells; D: Transwell assay for the effect of ZNF503-AS1 overexpression on migration and invasion ability of J82 cells (× 200, scale bar = 50 μm); E: qRT-PCR and Western blot analysis for the effect of ZNF503-AS1 overexpression on the expression of MMP-2, MMP-9, Ki67 and Bax in J82 cells. Measurement data were summarized as mean ± standard deviation. Data were analyzed by unpaired t test between two groups from three independent experiments. * p < 0.05 vs. the oe-NC group. (EPS 3481 kb)
Fig. S2
ZNF503-AS1 recruits GATA6 to up-regulate SLC8A1 expression in bladder cancer J82 cells. A: qRT-PCR and Western blot analysis for the SLC8A1 expression in J82 cells overexpressing or silencing SLC8A1. B: qRT-PCR and Western blot analysis for the SLC8A1 expression in J82 cells overexpressing or silencing ZNF503-AS1. * p < 0.05 vs. the sh-NC group, # p < 0.05 vs. the oe-NC group; C: RIP assay to identify the binding of ZNF503-AS1 to GATA6. * p < 0.05 vs. the IgG; C: qRT-PCR and Western blot analysis for determining the SLC8A1 expression in J82 cells, * p < 0.05 vs. the oe-NC + sh-NC group, # p < 0.05 vs. the oe-ZNF503-AS1 + sh-NC group. Measurement data were summarized as mean ± standard deviation. Data were analyzed by unpaired t test between two groups and compared by one-way ANOVA with Tukey post hoc test among multiple groups from three independent experiments. (EPS 1553 kb)
Fig. S3
SLC8A1 inhibits the proliferation and migration of bladder cancer J82 cells by promoting Ca2+ influx. A: EdU assay to detect the effect of SLC8A1 overexpression on J82 cell proliferation; B: quantitative flow cytometric detection of SLC8A1 overexpression on J82 cell apoptosis; C: Transwell assay to detect the effect of SLC8A1 overexpression on J82 cell migration ability; D: Transwell assay for the effect of SLC8A1 overexpression on the invasive ability of J82 cells. Measurement data were summarized as mean ± standard deviation and analyzed by unpaired t test between two groups from three independent experiments. * p < 0.05 vs. the oe-NC group. (EPS 598 kb)
Fig. S4
ZNF503-AS1 promotes Ca2+ and inhibits bladder cancer J82 cell proliferation and migration by regulating the SLC8A1 expression. A, qRT-PCR and Western blot analysis for the expression of SLC8A1 in J82 cells. B: EdU assay to detect the proliferation of J82 cells; C: flow cytometric detection of J82 cell apoptosis; D: Transwell assay to detect J82 cell migration ability (× 400, scale bar = 25 μm); E: Transwell assay to detect J82 cell invasive ability. Measurement data were summarized as mean ± standard deviation. Comparisons among multiple groups were performed using one-way ANOVA or repeated measures ANOVA with Tukey’s post hoc test. The experiment was repeated three times. * p < 0.05 vs. the oe-NC + sh-NC group, # p < 0.05 vs. the oe-ZNF503-AS1 + sh-NC group. (EPS 1032 kb)
Rights and permissions
About this article
Cite this article
He, H., Wu, S., Ai, K. et al. LncRNA ZNF503-AS1 acts as a tumor suppressor in bladder cancer by up-regulating Ca2+ concentration via transcription factor GATA6. Cell Oncol. 44, 219–233 (2021). https://doi.org/10.1007/s13402-020-00563-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-020-00563-z